Potri.006G257900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.006G257900.3 pacid=42768449 polypeptide=Potri.006G257900.3.p locus=Potri.006G257900 ID=Potri.006G257900.3.v4.1 annot-version=v4.1
ATGACGTCTTCGCTTAGTAGAGACCTCATCTTTTTGATTTTGCAGTTTCTTGATGAAGAAAAGTTTAAAGAAACTGTTCACAAGCTCGAGCAAGAATCGG
GGTTGTTCTTCAATGCAAAGTACTTTGAGGAGCTTGTGCTGGGTGGCAACTGGGATGAGGTTGAGAAGTATCTATCTGGGTTTACCAAAGTGGATGATAA
TCGATATTCGATGAAGATATTTTTTGAAATTCGCAAGCAAAAGTACCTCGAGGCGTTGGATAAGCTTGATCGGACCAAGGCTGTAGATATCCTCATGAAG
GATCTGAAAGTTTTTGCCTCTTTTAATGAAGACTTGTTTAAAGAAATTACCCAGCTGCTCACATTGGATAATTTTAGGGAAAATGACCAGCTCTCCAATT
ATAGAGATACAAAGTCAGCAAGAGCAATAATGTTGATTGAGCTCAAGAAGCTTATTGAGTCAAATCCCCTTTTCCGTGACAAATTGCAGTATCCTAACAT
AAAAAATTCAAGATTACGAATGCTTATCAACCAAAGCTTGAATTGGCAGCATTCGCTTTGTGGTAATCCAGGGCAAAACCCTGATATAAGAACCCTCTTT
TACGATCACAGTTGTAGAAATGCTAATCATGCGTATCCCCAACTAGCAGCAAACAATCACCTCATAGCTTCTGCACCAAAGTCTGAAGGTTTTCTCCCGA
TAGTTGCAAATGGGTCATTCCAACCAGCACCAGCTCCGGCACCGGCACCGGCACCGGCACCAGTTCAAACACCCTTAACAGCATGGATATCCAATCCCAC
AACTGTAACTCATCCAGTAGTTCCTGGAGTTGGACTAAATTTCATCAGCCCAAATCCAGGTATAGCTGCAATGCCAAAAGGTCCTGGCGATTCTGATGTG
TCAAGGCCAAGATTTTCTGGGGCTTTGGACAGGCAGGTAATGTTGACAGGAAATAATTCTGGTCAAAATCATCATGGCCTGGCCTTTAACATAACCGAAG
AGTTGCCCAGGAATGTGGCATGTACATTAAATCAAGGCTCAGCTCCCACAAGCATGGACTTCCATCCTCTTCGACAGACTCTACTTCTGGTTGGGACTGC
TATTGGAGACATTAGTTTGTGGGACGTTAGTTCTAGGGAGAAGTTGGCCTCTAAAAGTTTTCAGGTATGGGATATTGGAGCAAGTTCAATGGTATTGAAG
GCATCCATAGTCAAAGACCCATCTGTCTCAGTTAAACGTGTATTATGGAGTCCTGATGGTTCTTTATTTGGGGTTGCATATTCTAAGCATATGGTGCAGT
TATACACTTGTTATGGTGGACATGATATTCGGCATCATATAGAGATCGATGCTCATGTTGGAAGTGTAAATGATCTAGCATTTTGCAACCCCAACAAGCA
ATCTGTTATCACTTGTGGTGATGACAAGACCATCAAGGTGTGGGAAGTGGCCACTGGTGCCAAATTGTCTACGTTTGAAGGTCATGAAGCTCCTGTTCAT
TCGATCTGTCCCCATTCCAGGGAAACTGTTCATTTTGTCTTTTCAACATCACTGGATGGGAAAATAAAGGCATGGTTGCATGGCGTGATGGGATCTAGGG
TTGATTATGATGCTCCTGGTCGATCATGCTCCACAATGGCATATAGTGCTGATGGGAAAAGGCTATTTTCATGTGGGACCAGTCAAGATGGGGAGTCACA
CATGGTGGAATGGAATGAAAATGAAGGTACTATTAAGAGAACCTACCAAGGATTTCAAAAGCGTTCTTTGGGTGTTGTTCAATTCGATACTACCAAGAAT
AGGTTCTTGGCTGTTGGTGATGACTACTCAATCAAATTCTGGGATATGGATAACCCTTCTCTTTTGACAACCATTGACGCGGAGGGAGGCCTACCAACAA
GTCCACGTATTCGATTCAATAAAGGGGGCAACCTATTGGCTGTATCTGCAAATGATAATAGAATCAAAATCTTGGCAACAGTTGATGGTCTCTGCTTGAT
GCGCACCTTTGAAGGCCATTCTCTGGTTGCCTCGAGACTTGGAATTGCATCTGAGGCACTGATAAAGAATGGTGACACCAGAAACTCAGAAGGTGTAAAA
CCAAGGGTGCCTGAAGAAGCACACCCACCAAAGATATGGAAGCTCACTGAGATTAATGATCCTTCCAAGCTTCATTCGCTGAGACTCTCAGCTCGTGTTA
AAACAGACAAGATCGCGAGGTTGGTCTACACAAATTCAGGCACTGCCATCTTGGCACTTGCATTAAATGCAATTCATCTACTTTGGAAGTGGCCTCGAAA
TGACCTTAATTCGAGTGGGAAGGCAACAACCAAGGCTACCCCTCAATTGGTGCAACCAGCATCTGGCATATTAATGACCAATGATCTCATGGATGCTAGA
CCAGAAGAGGCTGTGCCATGTTTTGCTCTGTCGAAGAATGATTCCTATATCATGTCTGCCTCTGGGGGGAAAATTTCCTTGTTCAACACAATGACATTCA
AGATTATGACAGCATTCATGCCTCCGCCACCTGCAGCGACGTATCTTGCGTTCCATCCTCAAGATAACAATATTGTTGCAGTGGGAATGGACGATTCCAC
AGTACACATATATAATGTTCGTGTGGATGAGGTCAAGAGCAAGCTTAAGGGTCACTCTAAGAGAATCACTGGTCTTGCCTTCTCTAGTGTGCTGAATACA
CTTGTTTCCTCTGGCGCAGATTCTCAGGTTATCGTGTGGAGCATTGATAGGTGGGAAAGGAAGAAAAATTGTGTCTTGCAGGTTCCAGCTGGAAGGACAC
CTGCAGCAATGTCAGACACCCAAGTTCAGTTTCACCAAGATCAAGTCCATTTGCTTGTTGCACATGACACTCAGCTTGGCATTTATGAAACAACAAAACT
TGAATGCTTAAAACAGTGGACCATTGGAGAGTTTTCAGCACCAATCTCGCATGCAACATTCTCTTGTGATAGCCAGTTAGTTTATGCTAGCTTCCTGGAT
GGAACTCTGCGTGTATTTAGTGCATCAAATCTTCAAGTGAGATGCCAGATTAATCCCAATTCCTATCTTCCATCAGATGTCAGTTCTACTGTATATCCTC
TAGCAATTGCTGCACATCCCCAAGAACCTAATCAGTTTGCCATAGGCCTAACAGATGGCAGTGTTCAAGTTTTCGAGCCTCTTGAGTCTGACGGCAAATG
GGGCGTTCCCCCACCTGCTGAAAATGGGGCATCAGGTAGCATGCCTTCCACTACCGCACCTCCAGCTGTTGCTTTAGATCAACCCCAGAACATGGAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.006G257900.3 pacid=42768449 polypeptide=Potri.006G257900.3.p locus=Potri.006G257900 ID=Potri.006G257900.3.v4.1 annot-version=v4.1
MTSSLSRDLIFLILQFLDEEKFKETVHKLEQESGLFFNAKYFEELVLGGNWDEVEKYLSGFTKVDDNRYSMKIFFEIRKQKYLEALDKLDRTKAVDILMK
DLKVFASFNEDLFKEITQLLTLDNFRENDQLSNYRDTKSARAIMLIELKKLIESNPLFRDKLQYPNIKNSRLRMLINQSLNWQHSLCGNPGQNPDIRTLF
YDHSCRNANHAYPQLAANNHLIASAPKSEGFLPIVANGSFQPAPAPAPAPAPAPVQTPLTAWISNPTTVTHPVVPGVGLNFISPNPGIAAMPKGPGDSDV
SRPRFSGALDRQVMLTGNNSGQNHHGLAFNITEELPRNVACTLNQGSAPTSMDFHPLRQTLLLVGTAIGDISLWDVSSREKLASKSFQVWDIGASSMVLK
ASIVKDPSVSVKRVLWSPDGSLFGVAYSKHMVQLYTCYGGHDIRHHIEIDAHVGSVNDLAFCNPNKQSVITCGDDKTIKVWEVATGAKLSTFEGHEAPVH
SICPHSRETVHFVFSTSLDGKIKAWLHGVMGSRVDYDAPGRSCSTMAYSADGKRLFSCGTSQDGESHMVEWNENEGTIKRTYQGFQKRSLGVVQFDTTKN
RFLAVGDDYSIKFWDMDNPSLLTTIDAEGGLPTSPRIRFNKGGNLLAVSANDNRIKILATVDGLCLMRTFEGHSLVASRLGIASEALIKNGDTRNSEGVK
PRVPEEAHPPKIWKLTEINDPSKLHSLRLSARVKTDKIARLVYTNSGTAILALALNAIHLLWKWPRNDLNSSGKATTKATPQLVQPASGILMTNDLMDAR
PEEAVPCFALSKNDSYIMSASGGKISLFNTMTFKIMTAFMPPPPAATYLAFHPQDNNIVAVGMDDSTVHIYNVRVDEVKSKLKGHSKRITGLAFSSVLNT
LVSSGADSQVIVWSIDRWERKKNCVLQVPAGRTPAAMSDTQVQFHQDQVHLLVAHDTQLGIYETTKLECLKQWTIGEFSAPISHATFSCDSQLVYASFLD
GTLRVFSASNLQVRCQINPNSYLPSDVSSTVYPLAIAAHPQEPNQFAIGLTDGSVQVFEPLESDGKWGVPPPAENGASGSMPSTTAPPAVALDQPQNME
|
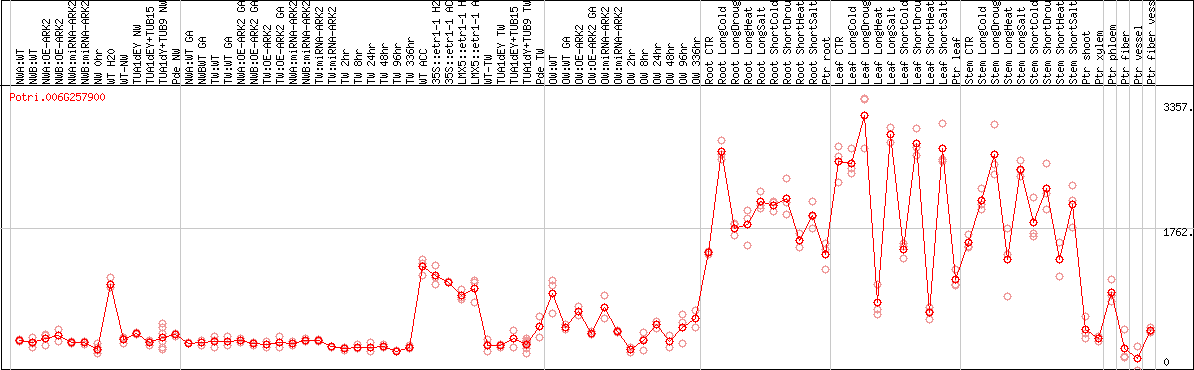
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.006G257900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
