HAC901,Pt-HAC1.4 (Potri.007G003400) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | HAC901,Pt-HAC1.4 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G003400.2 pacid=42765714 polypeptide=Potri.007G003400.2.p locus=Potri.007G003400 ID=Potri.007G003400.2.v4.1 annot-version=v4.1
ATGGGGATGATGGGGAATAATATACCGCTTGCTAATGAGCCTGGTACCTCTGAGGGCTATATGACTTCCACACATTATGTGAATTCGCCTAAACCATTGC
CACAGCAGTTTGATCAACATCAGCGACAACTAATGCAAGGTGATGGATATGGGATGAGTAATGCTGATTCTCTTGGATCTGGAAACATATATGGTGCTGT
AACATCTGTTGGATCAATGATGAATGCCCAGTCTATGTCCAAAACTAACTCTTCTCTTGTAAATAATCAGTCTAATTTGCATGCCCTGCATCAAGCTGGA
CATATAAAGCTTCAATCACTTGATCAGTCTGAAAAGATGAATTTCCAGTCTTCACTTCAACAGCAGCAACTTCAACAACATCCTCACCAACAGCAACAAC
TTCAGCAGCATCCTCATCAATTTCAACAGCAGCAACTTGTTCAACAGCAACGTTTACAGAAGCAGCAAAGTCAGCAGCATCAGCATCTGCTGAACAATGA
TGCTTTTGGTCAGTCTCTGCTCATATCTGATCCTAGCAGTCAAGTAAAGCGTGAGCCTGGAATGGAGCACCATAATGATGTTTTGCACTCACAAACCTCA
GACCACTTCCAAATATCTGAATTGCAAAACCAATTCCAGCAAAATGTGCTTGGAGACCATTCTAGGAATGCGCAGAATCCCCCTCATCCAGACCGACAGC
ATGACATGAGCTCATCATTGACACAAAATTCACAGCAAATGCAACAGATGTTGCATCCACATCAGTTGGTTTCAGAGTCTCAAAATAATTTTAATGGCCT
TTCTGTTGGGACACAATCAGATTCAGCATTGCCAGGTCAATGGTATCCACAATCTCAGGACCGGACTCGCATGCCAGGGAGCAATTCACATGAGCAGCAT
GTCCAAGAGGATTTCCTCCAGAGAATATCCGGGCAAGGTGAAGCTCAGTGTAATAATTTAGCTTCAGAAGGATCCATTGTCAGCCAAACAGTACCTCCTA
GAAGCACATCAGAGCCCCAAAATTCAAATGGTGTCACCTATAGATCTGGAAACGCTAACCGTGACCGGCAATTTAGAAATCAACAGAAGTGGCTTTTGTT
CTTGCGGCATGCCCGGCGATGTCCTGCCCCAGAAGGTCAATGTCCTGACCCCAACTGCACTACTGTCCAAAAGCTGTTGAGGCACATGGACAGATGCAAC
TCCACTCCATGCTCATATCCCCGATGCCAGCATACCAGGATATTGATTCATCACTTTAAGCACTGCAGAGATTCAGGTTGCCCTGTTTGTATTCCTGTTA
GAAATTATCTAGAGGCACAAATAAAGATACAAATGAAGGCCCGCACTCTTCCAGCCTTAGATTCTGGTTTGCCAAGCAAAGGCAGTGATACTGGAGACAA
TGCAGCCCGATTGATTTCAAGAACTCCATCAATTGTTGAAAGCTCTGAAAATCTACAACCTTCTCTAAAACGCATGAAGATTGAGCAATCTTCCCAAACT
CTTAAACCTGAGATTGAAGTTTCTGTCATATCAGCATCTGCAGTCAGTGATGCTCATATAACCCTGGATGTCCAGCACCAAGATCACAAGCATGGTGATA
ATTGTCCGCTGGTTAAGTCTGAGTACATGGAAGTGAAGTTGGAAGTACCTGCAATTTCTAGACAAGGAAGTCCTAGTAATAGTGAGATGAAAAAGGATAA
CGTGGATGATGTCAGCAGCCAAATGCCTGCAGATGAATCCATGGTACATGATGAGCCTGCCAGTTTAGCTAAGCAGGACAACGTTAAAGTTGAGAAAGAA
GCTCATCTACTTAAGCAAGAAAATGCTACGCACCCTGCTGAAAATGCAGCTGGAACCAAGTCTGGGAAGCCAAAAATAAAGGGGGTATCATTGACTGAAC
TGTTCACGCCTGAACAAGTGAGGGAGCATATTATAGGCCTCAGGCAGTGGGTTGGCCAGAGTAAGTCAAAGGCAGAAAAGAATCAAGCAATGGAGCATTC
AATGAGTGAGAACTCTTGTCAATTGTGTGCAGTTGAAAAGCTTACTTTTGAACCACCACCAATATACTGTACACCCTGTGGTGCTCGCATTAAGCGGAAT
GCAATGTTTTATACTATGGGAGCTGGTGATACACGACACTATTTCTGTATTCCATGCTATAATGAGGCTCGTGGGGACACTATTGTTGCTGATGGGAATG
CTATTCCAAAGGCAAGACTCGAGAAGAAGAAAAATGATGAGGAGACTGAAGAATGGTGGGTTCAATGTGACAAATGTGAAGCTTGGCAACACCAAATTTG
TGCTTTATTTAATGGTAGAAGGAATGATGGTGGGCAAGCTGAATATACTTGCCCTAATTGCTACATAACAGAGGTTGAAAGAGGAGAGCGCAAGCCTTTA
CCACAAAGTGCAGTTCTGGGGGCTAAAGATCTACCTAGAACAATACTCAGTGATCATATAGAGCAGAGGTTATTTAGGACACTGAAGCAGGAAAGACAGG
ATAGGGCCAGGGCACAGGGGAAAAGTTTTGATGATGTTCCTGGAGCAGAATCACTTGTAGTCCGAGTTGTTTCATCAGTTGACAAAAAGTTGGAAGTGAA
GCAGCGATTTCTTGAGATATTTCGGGAGGAGAATTATCCAACTGAATTCCCGTATAAGTCTAAGGTAGTTTTGCTGTTTCAGAAGATTGAAGGTGTTGAG
GTATGCCTATTTGGCATGTATGTTCAAGAATTCGGATCAGAAGCTCACTTTCCAAATCAGCGTCGTGTCTATCTTTCATATCTGGATTCTGTGAAGTACT
TCAGACCTGAGATTAAAGCAGTGACTGGAGAGGCTCTCCGTACATTTGTTTACCATGAAATTTTGATTGGATACCTTGAATACTGCAAGAAAAGGGGTTT
TACAAGCTGCTACATATGGGCTTGCCCTCCGTTAAAGGGTGAGGATTATATATTGTATTGCCATCCAGAAATCCAGAAAACACCAAAATCTGACAAGCTC
CGGGAATGGTATTTAGTGATGTTACGAAAAGCAGCCAAGGAAAATGTTGTTGTTGACCTCACTAATTTGTATGACCATTTCTTCATATCTACTGGTGAAT
GTAAGGCAAAGGTCACAGCAGCTAGGCTACCCTACTTTGATGGGGATTATTGGCCTGGTGCTGCAGAGGATTTAATCTATCAGCTTAATCAAGATGAAGA
TGGGAGAAAACAAAATAAGAAGGGAAGTACCAAAAAGACCATTACAAAAAGGGCTCTAAAAGCATCTGGCCAGGCAGATCTTTCTGGTAATGCATCGAAG
GATCTGCTACTGATGCATAAACTTGGTGAGACAATTTGCCCTATGAAGGAAGACTTTATCATGGTTCACTTGCAGCCTTGCTGTTCTCATTGTTGTATTC
TTATGGTATTGGGCACCCATTGGGTTTGCAACCAGTGCAAAAACTTTCAGATTTGTGACAAGTGTTATGAAGTAGAACAGAAACGTGAAGAAAGAGAGCG
ACATCCCATCAACCAAAGGGAAAAACATGCATTCTATCATGTTGAAATCACTGATGTACCTGCTGACACAAAGGATAAAGATGAAATTCTCGAGAGTGAA
TTTTTTGACACGAGACAGGCATTTTTGAGTCTTTGTCAAGGAAACCATTATCAATATGACACTTTACGCCGGGCTAAACATTCCTCAATGATGGTCCTTT
ACCATCTCCATAATCCAACTGCTCCTGCATTTGTGACAACATGCAATATATGTCATCTTGACATCGAAACGGGTCAAGGTTGGCGTTGTGAGGTTTGCCC
TGATTATGATGTATGCAATTCTTGTTATCAGAAGGATGGGGGTATGGATCATCCTCATAAGTTGACGAATCATCCATCCCTGGCAGAGCGCGATGCACAG
AACAAGGAAGCTAGGCAACAGAGGGTTCTGCAGCTAAGGAAAATGCTTGATCTTCTGGTACATGCATCACAATGCCGTTCTCCACATTGCCAATATCCTA
ATTGTCGCAAGGTGAAAGGTCTTTTCCGGCATGGGATACAGTGCAAAACTCGTGCATCTGGAGGCTGTGTCCTGTGTAAAAAGATGTGGTATCTCCTTCA
ACTTCATGCTCGGGCATGCAAAGAATCAGAATGCCATGTTCCACGCTGCAGGGACTTGAAAGAACATTTGAGGAGACTTCAGCAGCAGTCTGATTCACGA
CGAAGGGCTGCTGTGATGGAGATGATGAGGCAGAGAGCTGCTGAGGTTGCTGGCAATACTGGGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G003400.2 pacid=42765714 polypeptide=Potri.007G003400.2.p locus=Potri.007G003400 ID=Potri.007G003400.2.v4.1 annot-version=v4.1
MGMMGNNIPLANEPGTSEGYMTSTHYVNSPKPLPQQFDQHQRQLMQGDGYGMSNADSLGSGNIYGAVTSVGSMMNAQSMSKTNSSLVNNQSNLHALHQAG
HIKLQSLDQSEKMNFQSSLQQQQLQQHPHQQQQLQQHPHQFQQQQLVQQQRLQKQQSQQHQHLLNNDAFGQSLLISDPSSQVKREPGMEHHNDVLHSQTS
DHFQISELQNQFQQNVLGDHSRNAQNPPHPDRQHDMSSSLTQNSQQMQQMLHPHQLVSESQNNFNGLSVGTQSDSALPGQWYPQSQDRTRMPGSNSHEQH
VQEDFLQRISGQGEAQCNNLASEGSIVSQTVPPRSTSEPQNSNGVTYRSGNANRDRQFRNQQKWLLFLRHARRCPAPEGQCPDPNCTTVQKLLRHMDRCN
STPCSYPRCQHTRILIHHFKHCRDSGCPVCIPVRNYLEAQIKIQMKARTLPALDSGLPSKGSDTGDNAARLISRTPSIVESSENLQPSLKRMKIEQSSQT
LKPEIEVSVISASAVSDAHITLDVQHQDHKHGDNCPLVKSEYMEVKLEVPAISRQGSPSNSEMKKDNVDDVSSQMPADESMVHDEPASLAKQDNVKVEKE
AHLLKQENATHPAENAAGTKSGKPKIKGVSLTELFTPEQVREHIIGLRQWVGQSKSKAEKNQAMEHSMSENSCQLCAVEKLTFEPPPIYCTPCGARIKRN
AMFYTMGAGDTRHYFCIPCYNEARGDTIVADGNAIPKARLEKKKNDEETEEWWVQCDKCEAWQHQICALFNGRRNDGGQAEYTCPNCYITEVERGERKPL
PQSAVLGAKDLPRTILSDHIEQRLFRTLKQERQDRARAQGKSFDDVPGAESLVVRVVSSVDKKLEVKQRFLEIFREENYPTEFPYKSKVVLLFQKIEGVE
VCLFGMYVQEFGSEAHFPNQRRVYLSYLDSVKYFRPEIKAVTGEALRTFVYHEILIGYLEYCKKRGFTSCYIWACPPLKGEDYILYCHPEIQKTPKSDKL
REWYLVMLRKAAKENVVVDLTNLYDHFFISTGECKAKVTAARLPYFDGDYWPGAAEDLIYQLNQDEDGRKQNKKGSTKKTITKRALKASGQADLSGNASK
DLLLMHKLGETICPMKEDFIMVHLQPCCSHCCILMVLGTHWVCNQCKNFQICDKCYEVEQKREERERHPINQREKHAFYHVEITDVPADTKDKDEILESE
FFDTRQAFLSLCQGNHYQYDTLRRAKHSSMMVLYHLHNPTAPAFVTTCNICHLDIETGQGWRCEVCPDYDVCNSCYQKDGGMDHPHKLTNHPSLAERDAQ
NKEARQQRVLQLRKMLDLLVHASQCRSPHCQYPNCRKVKGLFRHGIQCKTRASGGCVLCKKMWYLLQLHARACKESECHVPRCRDLKEHLRRLQQQSDSR
RRAAVMEMMRQRAAEVAGNTG
|
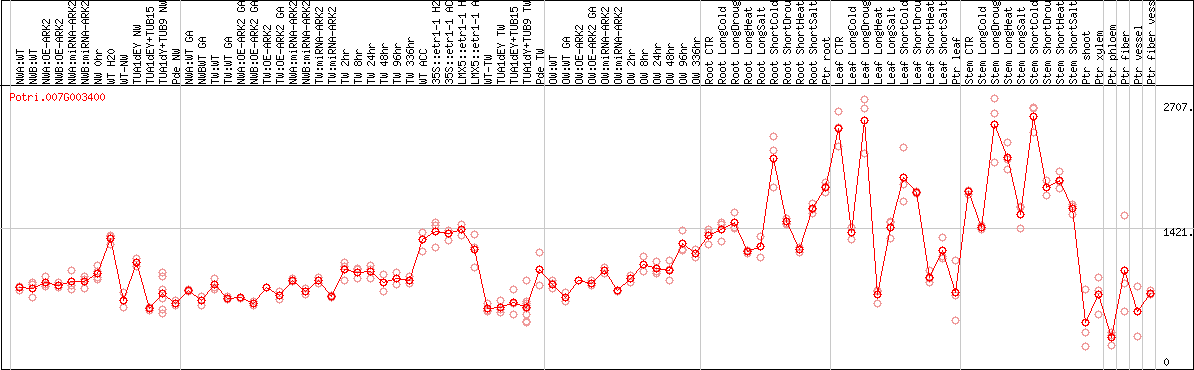
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G003400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
