Potri.007G010700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G010700.1 pacid=42766356 polypeptide=Potri.007G010700.1.p locus=Potri.007G010700 ID=Potri.007G010700.1.v4.1 annot-version=v4.1
ATGGCGGAGCAGAGAAATATGTGGAACTGGGAGGTGGCTGGCTTCGAGCCAAGGCCGGTGGAGGTGGAGCAGCCGATCGTACGGCGGTACTCGATCTCGA
CGACGCGAGAGAATTCGGAGTTCTCGAAGCAGGCTCTCGCCTCCAAAGTTCACCGATTGAAAGATAAAATCAAGCTTGCAAAAGAGGATTACTTGGAGTT
GAGACAAGAAGCAAGTGATTTACAGGAATATTCTAATGCGAAACTTGATCGAGTTACTCGTTATCTTGGAGTCCTTGCTGAAAAAACTCGTAAGCTAGAT
CAAGTTGCCCTTGAAACCGAGGCTAGAATATCTCCACTCATCAATGAGAAGAAAAGATTGTTCAATGACTTGTTAACAGCCAAAGGAAGCATAAAGGTGT
TTTGTCGTGTACGACCATTATTTGAAGATGAAAGTCCTTCTGTTGTTGAATTTCCTGATGATTGCACAATCCGCGTCAATACTGGCTCTGATACCATCTC
CAATCCCAAAAAAGATTTTGAATTTGACAGAGTTTATGGACCTCATGTGGGACAAGCTGAGTTGTTCACAGATGTTCAACCATTTGTGCAGTCTGCTTTG
GATGGATATAATGTTTCTATGTTTGCTTATGGACAAACTCACTCTGGAAAGACACACACTATGGAAGGATCTAGCTATGATCGTGGTTTATATGCTCGTT
GTTTTGAAGAGTTGTTTGATTTGGCTAACTCAGATTCAACTTCTACTTCTCAGTTCAATTTCTCTGTTACGGTTTTTGAGCTATACAATGAACAGATAAC
TGATTTGCTGTCAGAGTCCGAGAGTACTTTGCAGAAGATCTGCATGGGATCACTGGAATCTTTTATAGAGCTTCAACAAGAAAAAGTCGATAACCCATTG
GACTTTTCTAGAATCTTGAAAGCTGCATTTCAGAGACGGGAAAATAACATATCAAAGTTGAATGTTTCTCACTTGATTGTAACAGTGCACATATATTATA
ACAATGTGATCAGTGGAGAGAACTTGTATAGCAAGCTTTCTCTTGTTGACCTGGCTGGAAGTGAAGGTCTAATAGCAGAAGATGATAGCAGTGAGCGTGT
GACAGACATGCTACATGTGATGAAATCACTCTCAGCGTTGGGGGATGTCTTGTCTTCTTTAACTTCAAGGAAGGATGTTGTTCCTTATGAAAATTCAATG
CTGACAAAAGTTCTTGCTGACTCCTTAGGGAGGGATTCAAAAACTTTGATGATACTTAATGTGTGTCCAAATATTGCAAATTTGTCTGAGACATTATCGT
CTCTCAGCTTCTGTTCTAGGGCTAGAAATGCTACACTAAGCCTCGGAAATCGAGATACAATTAAGAAGTGGAGAGATGTTGCTAATGATGCACGCAAGGA
GTTGTATGAAAAGGAAAAGGAAATTCAAGATCTGAAACAAGAGGTTTTGGAATTGACGCAAGCACTCAAAGATGCAAATGACCAGTGTGTCTTGCTCTTC
AATGAAGTTCAGAAGGCGTGGAAAGTTTCTTTCACCCTGCAATCGGATTTAAAGTCAGAGAATATTATGATTGCTGACAAGCATAAGGTGGAAAAGGAAC
AAAATGCTCAGCTTAGAAATCAAGTTGCTCAACTGCTACACACGGAACAGGATCAAAAAATGATAATGCAACAAAAAGATTCCACAATTCAGACACTCCA
GGCCCAAATTAAAAGCATGGAGTCACAGCTCAATGAAGCTCTCCGTTTGAGGGAAGCTCAGTCAACATTTGGCTCAGAATCAGGGCCTGTGATTTCATCA
ATCTCTAAGGCAACCGGGGATGGCATGGATTCTTCAGCAGTAACCAAGAAACTTGAAGAAGAGCTAAGAAAGCGTGATGCTTTGATTGAGAGGTTGCATG
AAGAAAATGAGAAACTGTTCGATAGATTAACAGAAAAAGCCTCTTTAGCTGGATCACCCCAGGTGTCTAGTCCATTGTCCAAAGGAACTGTGAATGTTAA
GTCTCAGGAGCTGGGAAGAAATGAGAACAATAAAGGACGTTCAATGGATGTTGCTCCATCACCATTGGGTGCAGATAAGACTGATGGAACCGTTGCCTTG
GTTAAATCAGGATCTGAGAAAGTTAAATCCACTCCGGCAGGAGAATATCTTACAGCTGCACTAAATGACTTTGATCCTGAACAATATGACAGCCTTGCTG
CCATTTCAGATGGTGCAAACAAGCTTTTGATGTTGGTTCTGGCAGCAGTCATTAAAGCAGGTGCTTCTAGAGAGCATGAAATACTTGCTGAAATCAGAGA
TGCTGTTTTCTCTTTTATCCGCAAAATGGAACCAAAGAGGGTAATGGATACCATGCTTGTTTCCCGTGTCAGAATTTTGTACATAAGATCTTTGCTTGCC
AGGTCTCCGGAGCTGCAGTCCATCAAGGTTCCTCCTGTTGAGTGCTTTTTGGAGAGGGCTAATACTGGACGAAGTAGAAGCTCCAGCCGAGCCAATAGTC
CTGGAAGATCCCCTGTGCACTTTGTGGAGGAGCAGATCCAAGGCTTTAAAGTGAACATAAAGCTGGAGAAGAAGTCTAAGCTGTCATCAGTTGTTTTGAG
AATGAGAGGAATTGATCAGGATGCATGGAGGCAGCAGGTAACTGGTGGAAAACTGAGGGAAATTCAAGAGGAAGCAAAAAGTTTTGCAATAGGAAACAAA
GCTCTTGCTGCTCTCTTTGTTCATACTCCAGCTGGCGAGCTGCAGAGGCAAATTAGGTCCTGGCTTGCGGAGAATTTTGAGTTTCTTTCTGTTACTGGAG
ATGATGCATCGGGTGGGATCACTGGTCAATTAGAACTCCTTTCGACTGCAATTATGGATGGCTGGATGGCTGGATTGGGTGCTGCATTGCCTCCCAGTAC
AGATGCTCTTGGCCAGCTTCTATCCGAGTATGCAAAACGGGTTTTTACTTCTCAATTGCAACACTTAAAGGATATTGCTGGGACCTTAGCATCAGAAGAG
GCAGAAGATGCAGCACAAGTAGCGAAACTGCGTTCAGCTCTAGAGTCTGTTGATCACAAAAGAAGAAAGATTTTGCAACAAATGAGAAGTGATGCAGCCT
TATTAACATTGGAAGATGGTGGTTTGCCTGTTCAAAATCCTTCCACTGCTGCTGAAGATGCACGATTAGCATCTCTGATTTCTCTTGATGGCATATTGAA
GCAAGTCAAGGATATATTGAGACAATCCTCAGTGAACACCTTGAGTAAATCTAAGAAGAAAACGCTGCTTGTTTCTCTGGATGAACTTGGGGAAAGGATG
CCTTCCCTCCTCAATATTGATCACCCATGTGCCCAAAGGCAAATTGCAGAAGCTCGTCGCATGGTTGAGTCAATTCCTGAACAAGATGACCCCCTTCATG
AGTTGGCACATGCTCGAAAATCTACTGCTGATTTGGGGTCTGGTACAGAAACTGATGTGGCACAGTGGAACGTTTTGCAATTCAATACAGGCTCTACTAC
CCCGTTTATCATCAAGTGTGGAGCAAACTCAAACTCAGAACTGGTGATTAAGGCTGATGGGCGTGTTCAGGAACCTAAAGGTGGTGAAATCATGAGGGTC
GTTCCAAGACCATCAGTTTTGGAAAATATGAGCATGGATGAAATGAAACATGTATTCTCTCAACTACCTGAAGCTTTAAGCTTGCTTGCCCTTGCAAGGA
CTGCAGATGGTACCCGAGCTAGGTATTCTAGACTTTATAGAACTTTAGCTATGAAAGTTCCCTCATTGAGGGACCTTGTTGGTGAGCTTGAGAAGGGTGG
AGTGCTGAAAGATGTGAAGTCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G010700.1 pacid=42766356 polypeptide=Potri.007G010700.1.p locus=Potri.007G010700 ID=Potri.007G010700.1.v4.1 annot-version=v4.1
MAEQRNMWNWEVAGFEPRPVEVEQPIVRRYSISTTRENSEFSKQALASKVHRLKDKIKLAKEDYLELRQEASDLQEYSNAKLDRVTRYLGVLAEKTRKLD
QVALETEARISPLINEKKRLFNDLLTAKGSIKVFCRVRPLFEDESPSVVEFPDDCTIRVNTGSDTISNPKKDFEFDRVYGPHVGQAELFTDVQPFVQSAL
DGYNVSMFAYGQTHSGKTHTMEGSSYDRGLYARCFEELFDLANSDSTSTSQFNFSVTVFELYNEQITDLLSESESTLQKICMGSLESFIELQQEKVDNPL
DFSRILKAAFQRRENNISKLNVSHLIVTVHIYYNNVISGENLYSKLSLVDLAGSEGLIAEDDSSERVTDMLHVMKSLSALGDVLSSLTSRKDVVPYENSM
LTKVLADSLGRDSKTLMILNVCPNIANLSETLSSLSFCSRARNATLSLGNRDTIKKWRDVANDARKELYEKEKEIQDLKQEVLELTQALKDANDQCVLLF
NEVQKAWKVSFTLQSDLKSENIMIADKHKVEKEQNAQLRNQVAQLLHTEQDQKMIMQQKDSTIQTLQAQIKSMESQLNEALRLREAQSTFGSESGPVISS
ISKATGDGMDSSAVTKKLEEELRKRDALIERLHEENEKLFDRLTEKASLAGSPQVSSPLSKGTVNVKSQELGRNENNKGRSMDVAPSPLGADKTDGTVAL
VKSGSEKVKSTPAGEYLTAALNDFDPEQYDSLAAISDGANKLLMLVLAAVIKAGASREHEILAEIRDAVFSFIRKMEPKRVMDTMLVSRVRILYIRSLLA
RSPELQSIKVPPVECFLERANTGRSRSSSRANSPGRSPVHFVEEQIQGFKVNIKLEKKSKLSSVVLRMRGIDQDAWRQQVTGGKLREIQEEAKSFAIGNK
ALAALFVHTPAGELQRQIRSWLAENFEFLSVTGDDASGGITGQLELLSTAIMDGWMAGLGAALPPSTDALGQLLSEYAKRVFTSQLQHLKDIAGTLASEE
AEDAAQVAKLRSALESVDHKRRKILQQMRSDAALLTLEDGGLPVQNPSTAAEDARLASLISLDGILKQVKDILRQSSVNTLSKSKKKTLLVSLDELGERM
PSLLNIDHPCAQRQIAEARRMVESIPEQDDPLHELAHARKSTADLGSGTETDVAQWNVLQFNTGSTTPFIIKCGANSNSELVIKADGRVQEPKGGEIMRV
VPRPSVLENMSMDEMKHVFSQLPEALSLLALARTADGTRARYSRLYRTLAMKVPSLRDLVGELEKGGVLKDVKS
|
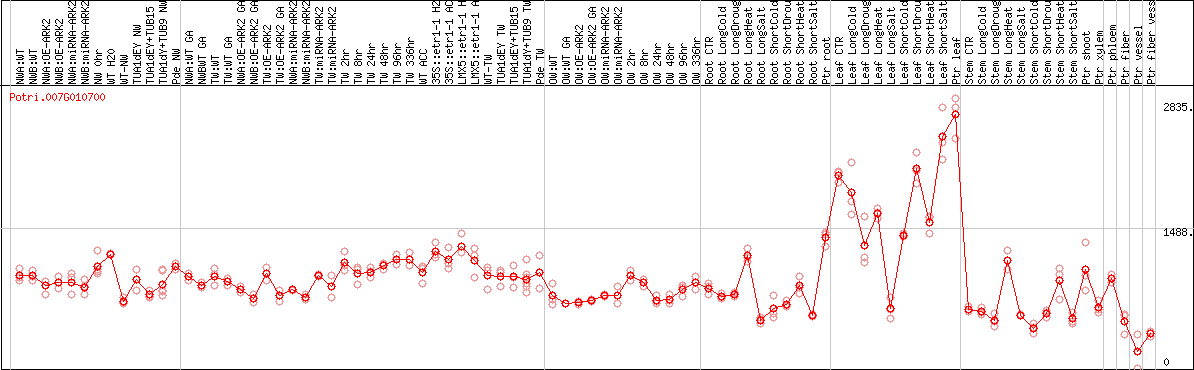
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G010700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
