External link
Symbol
SVP.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G22540 330 / 1e-115
MADS
AGL22, SVP
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
AT4G24540 231 / 6e-77
MADS
AGL24
AGAMOUS-like 24 (.1)
AT3G57230 135 / 5e-39
MADS
AGL16
AGAMOUS-like 16 (.1.2)
AT4G37940 133 / 2e-38
MADS
AGL21
AGAMOUS-like 21 (.1)
AT2G14210 129 / 5e-37
MADS
AGL44, ANR1
ARABIDOPSIS NITRATE REGULATED 1, AGAMOUS-like 44 (.1)
AT2G22630 129 / 6e-37
MADS
AGL17
AGAMOUS-like 17 (.1)
AT5G13790 126 / 2e-35
MADS
AGL15
AGAMOUS-like 15 (.1)
AT2G45660 124 / 5e-35
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT2G45650 123 / 2e-34
MADS
AGL6
AGAMOUS-like 6 (.1)
AT1G69120 123 / 2e-34
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G105600
301 / 2e-104
AT2G22540 288 / 3e-99
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.005G155300
290 / 3e-100
AT2G22540 234 / 8e-78
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.017G044200
242 / 4e-81
AT2G22540 206 / 9e-67
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.007G115200
242 / 4e-81
AT2G22540 189 / 3e-60
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.017G044500
238 / 7e-80
AT2G22540 201 / 4e-65
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.007G115100
228 / 2e-75
AT2G22540 186 / 1e-58
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.007G115000
211 / 5e-69
AT2G22540 181 / 9e-57
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.009G079000
151 / 3e-45
AT4G37940 293 / 3e-101
AGAMOUS-like 21 (.1)
Potri.002G109601
146 / 1e-43
AT3G57230 299 / 1e-103
AGAMOUS-like 16 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035942
318 / 2e-109
AT2G22540 307 / 1e-104
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10025719
285 / 7e-99
AT2G22540 286 / 4e-99
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10005325
154 / 5e-46
AT5G13790 249 / 7e-83
AGAMOUS-like 15 (.1)
Lus10039580
150 / 1e-44
AT5G13790 246 / 2e-81
AGAMOUS-like 15 (.1)
Lus10007100
130 / 2e-37
AT3G57230 226 / 4e-75
AGAMOUS-like 16 (.1.2)
Lus10034662
127 / 9e-36
AT5G60910 284 / 3e-97
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10005080
126 / 2e-35
AT3G02310 372 / 1e-131
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10006715
124 / 9e-35
AT4G22950 223 / 1e-73
AGAMOUS-like 19 (.1)
Lus10034371
124 / 1e-34
AT3G02310 364 / 3e-128
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10017871
121 / 1e-34
AT1G69120 264 / 2e-90
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
PF01486
K-box
K-box region
Representative CDS sequence
>Potri.007G010800.8 pacid=42765915 polypeptide=Potri.007G010800.8.p locus=Potri.007G010800 ID=Potri.007G010800.8.v4.1 annot-version=v4.1
ATGGCAAGAGAGAGGATTCAGATAAAAAAGATCGATAACGCCACCGCTAGGCAAGTCACGTTTTCAAAACGAAGAAGAGGGCTTTTCAAGAAAGCTGAGG
AGCTTTCAGTTCTCTGTGATGCTGATGTTGCTCTCATCATCTTCTCCTCCACTGGCAAGCTCTTTGAGTTCTCCAGCTCAAGCATGAAGGAAATACTGGA
AAGGCATAATTTGCACTCGAAGAATCTTGAGAAGCTGGAGCAACCATCTCTTGAGTTGCAGCTGGTAGAAGACAGCACCTGCTCCAGGTTGAGTAAGGAA
GTTGCGGAGAAAAGCCATCAGCTGAGGCAAATGAGAGGGGAAGATCTGCGAGGATTGGATATAGACGAATTGCTGCAGCTAGAGAAGTCTCTTGAGGCTG
GATTGAGCTGTGTGATAGAGAAGAAGGGTGAGAAGATTATGAACGAGATCACTGATCTTCAAAGAAAGGGAATGCAATTGATGGAAGAGAATGAGAGACT
CAAACAGCAAGTGGTGGAGATATCTAATGGCCGAAAGCATGTTACAGCTGATTCAGAGAATGTTGGTTACGAGGAAGGGCAGTCATCAGAGTCCGTAACC
AATGTCTGCAACTCAAATGGCCCTCTACATGATTATGAAAGCTCTGATACATCCCTCAAGTTGGGGTTGCCATTTTCAAACTGA
AA sequence
>Potri.007G010800.8 pacid=42765915 polypeptide=Potri.007G010800.8.p locus=Potri.007G010800 ID=Potri.007G010800.8.v4.1 annot-version=v4.1
MARERIQIKKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFSSSSMKEILERHNLHSKNLEKLEQPSLELQLVEDSTCSRLSKE
VAEKSHQLRQMRGEDLRGLDIDELLQLEKSLEAGLSCVIEKKGEKIMNEITDLQRKGMQLMEENERLKQQVVEISNGRKHVTADSENVGYEEGQSSESVT
NVCNSNGPLHDYESSDTSLKLGLPFSN
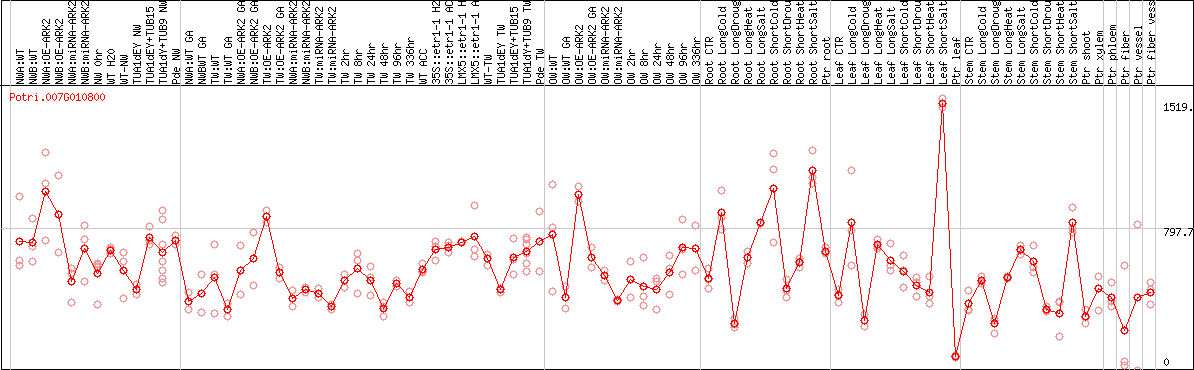
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G010800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.