Potri.007G017400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G017400.3 pacid=42765313 polypeptide=Potri.007G017400.3.p locus=Potri.007G017400 ID=Potri.007G017400.3.v4.1 annot-version=v4.1
ATGGCTAGGCTTCGAAGTAGCAAGCGTAATTCTTCCTCTAGATCATACACATCAACACTAACCACCATAGCCTTCATAGCATTGTGTGCTATAGGTGTTT
GGATGCTAACTTCAAACCCACAGGTTACTCCACAAACCACAACTCACGTCGCCAAACCAGTCATTACCACCACCACTGACATTGCCGCCGACGCGGATGT
TTCAATCTCCAATGAGGTAGAACACACTGAATCCAGGAGCAAAAAGGACACCCATGTTTATGAAGACAATCCAGGAGATCTTCCCGATGATGCTATCAAA
TCCGACGAGCTTAAAAGCAATGATGACAGTGACAATAAAGAAGAAAGTAACTACGGGAAACAGGAAACGGATGGTGGTGATAGTAAGGCTGATCAAGAGA
GCTCATCACAGGATTTAAAGGGAGAAGGATCTGGTGAGGAGCAGCAGAAACAGGAGGAAAGACAGAATCAGATATCTGAAGAAAGTTCACACACTCAAAA
TCGACAAGCTGATCAAACCAGCCAAGAAAGTTCTCAATCTGAAGGTAGCCAGGAAGCAAGTGTCAATCAAGAACAAGAAACAAATGCCAGCCAAGAAGAA
AAAACGAATGACAATCAAGAACAAGAGCAAAGTACAGTATCTGAAACTGATGATAGTAATTCACATGATTCTATAAATAAGAATGAAGAACAAGATCAAG
CACAACAGCAACAGCAACAGCAACAAGAAGATGTCGAAAACAGTAGTAAGGATTTACAGGATTCTCAGAACCAAGAATCAAAAGAAGACCAACAACAAGG
TTCTGGACTCGATGAAAACAATCAAGAATCTGATCACAATGAGAAAACATATGAAGAGCAACAACAAGAGCAAAGAATAGAAGATACTGGAGGTCAGAAT
TCTTCTCAAGAATCCCAAAAAGAGGTATCTGAAGAAGATAAAAAACAAAGAATACAACAGCAGCAGCAGCAACAACAAAAACAAGATGTTGAAGATAGTA
GTAAGGATTTACAGGATTCTCAGAACCAAGAATCGAAAGAAGACGAGCAACAACAAGGTTCTGGACTCAATGAAAACAATCAAGAATCGAATCAAAATGA
AAAACCATTTGAAGACCAACAACAACAAGATAGTACTGTAATCAATGAAAGCAATCAAGAATCTGATCAAAATGAGAAAACTTATGATGAGCAACAACAA
GAGCAAAGACAAGAGGATACTGAGGTTCATGATTCTTCTCAAGAATCCCAAAATGAGGTATCTGAAGAAGATCAAAAACAAAGAATACAACAGCAGCAGC
AGCAGCAACAGCATCAACAAACACATGATCAAGAAACCGAACAAGAATCTCAGGTGGATAGCAACACAAACCAAGAAACCAAGCAGGAATCAAGCTCGGG
CGAGTCAGCATTTCCAGGTGGCGGGAACCCAGGAATACCAAAAGAATCAAAGGAGTCATGGTCAACTCAAGCAGCAGAGTCAGAGAATCAAAAGGAGAGA
AGGAAGGAGGAATCAGACGGTAATGACAGCATGTATGGGTACACTTGGCAGCTCTGTAATGTCACTGCAGGTCCTGATTATATACCTTGTTTAGATAATG
AAAAGGCCTTAAGACAACTACATACAACCGGACACTTTGAACATAGAGAGAGGCATTGTCCTGAGCTTGGACCCACTTGTTTGGTTCCACTTCCTCAAGG
TTACAAAAGACCCATCACATGGCCTCAAAGTAGAGACAAGATATGGTATCACAATGTACCTCATCCAAAATTGGCAGAGGTCAAAGGTCATCAAAATTGG
GTTAAGGTTACTGGTGAGTTCTTGACCTTCCCTGGTGGTGGAACTCAGTTCATACATGGAGCTCTTCACTATATTGATTTTGTTCAACAGGCTGTGCCCA
AAATTAAATGGGGAAAGCACACTAGAGTGATACTGGATGTTGGGTGTGGAGTTGCAAGCTTTGGTGGTTATAATTTTGAAAGGGATGTTCTTACAATGTC
TTTTGCACCCAAAGATGAACATGAAGCTCAAGTCCAATTTGCCCTTGAAAGAGGAATACCTGCAATATCTGCTGTCATGGGTTCCCAGCGTCTCCCGTTC
CCTAGTAGGGTCTTTGATCTCATCCACTGTGCGCGTTGTCGAGTCCCTTGGCATGCAGAAGGTGGCAAGCTACTTTTAGAATTGAATCGCCTTCTCCGGC
CTGGAGGTTATTTTGTCTGGTCAGCAACTCCTGTTTACCAAAAGCTTCAGGAAGATGTGGAGATATGGCAAGCTATGTCTGCGTTGACGGTATCTATGTG
TTGGGAGCTTGTGACTATCAAGAAAGATAAACTTAACGGTATCGGTGCTGCCATCTACAGAAAACCTACCACAAATAATTGCTATGATCAAAGAATCAAG
AACAGCCCTCCAATGTGTGATAATGATGACGATGCAAATGCTGCCTGGTATGTGCCTCTGCAGGCATGCATGCACCGGGTACCTCGTTCTAAATCTCAGA
GAGGGGGTAAATGGCCAGAGGATTGGCCTGAAAGACTACAAATACCTCCTTACTGGCTAAAGAGCTCTCAGATGGGGATCTATGGTAAGCCAGCTCCTCA
AGATTTTGAAGCAGATTATGAACATTGGAAACATGTCGTCAGCAACTCATACATGAAGGGATTGGGTATCAGCTGGTCCAACGTCAGGAATATAATGGAT
ATGAGAGCCGTTTATGGGGGGTTTGCAGCAGCTCTCAAGGATCTTAAGGTCTGGGTGTTCAATGTAGTGAACACAGACTCTCCAGATACACTACCGATAA
TTTATGAGAGAGGTCTTTTTGGGATATACCATGATTGGTGCGAATCCTTCAGCACATACCCTCGAACTTACGACCTCCTACATGCTGATCATCTCTTCTC
AAAGCTGAAAAAAAGGTGCCAGCTTGCTCCTGTATTGGCAGAAGTTGATAGAATCGCAAGACCTGGAGGTAAACTGATTGTTCGTGATGAGTCAAGTGCA
ATTGAGGAAGTGGAGAACTTGTTGAAGTCTCTGCATTGGGAGGTTCATCTAATCTTCTCCAAGGACCAAGAAGGTTTACTCAGCGCACAAAAAGGTGAAT
GGCGACCACAGACATATGCAGCTTTCACCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G017400.3 pacid=42765313 polypeptide=Potri.007G017400.3.p locus=Potri.007G017400 ID=Potri.007G017400.3.v4.1 annot-version=v4.1
MARLRSSKRNSSSRSYTSTLTTIAFIALCAIGVWMLTSNPQVTPQTTTHVAKPVITTTTDIAADADVSISNEVEHTESRSKKDTHVYEDNPGDLPDDAIK
SDELKSNDDSDNKEESNYGKQETDGGDSKADQESSSQDLKGEGSGEEQQKQEERQNQISEESSHTQNRQADQTSQESSQSEGSQEASVNQEQETNASQEE
KTNDNQEQEQSTVSETDDSNSHDSINKNEEQDQAQQQQQQQQEDVENSSKDLQDSQNQESKEDQQQGSGLDENNQESDHNEKTYEEQQQEQRIEDTGGQN
SSQESQKEVSEEDKKQRIQQQQQQQQKQDVEDSSKDLQDSQNQESKEDEQQQGSGLNENNQESNQNEKPFEDQQQQDSTVINESNQESDQNEKTYDEQQQ
EQRQEDTEVHDSSQESQNEVSEEDQKQRIQQQQQQQQHQQTHDQETEQESQVDSNTNQETKQESSSGESAFPGGGNPGIPKESKESWSTQAAESENQKER
RKEESDGNDSMYGYTWQLCNVTAGPDYIPCLDNEKALRQLHTTGHFEHRERHCPELGPTCLVPLPQGYKRPITWPQSRDKIWYHNVPHPKLAEVKGHQNW
VKVTGEFLTFPGGGTQFIHGALHYIDFVQQAVPKIKWGKHTRVILDVGCGVASFGGYNFERDVLTMSFAPKDEHEAQVQFALERGIPAISAVMGSQRLPF
PSRVFDLIHCARCRVPWHAEGGKLLLELNRLLRPGGYFVWSATPVYQKLQEDVEIWQAMSALTVSMCWELVTIKKDKLNGIGAAIYRKPTTNNCYDQRIK
NSPPMCDNDDDANAAWYVPLQACMHRVPRSKSQRGGKWPEDWPERLQIPPYWLKSSQMGIYGKPAPQDFEADYEHWKHVVSNSYMKGLGISWSNVRNIMD
MRAVYGGFAAALKDLKVWVFNVVNTDSPDTLPIIYERGLFGIYHDWCESFSTYPRTYDLLHADHLFSKLKKRCQLAPVLAEVDRIARPGGKLIVRDESSA
IEEVENLLKSLHWEVHLIFSKDQEGLLSAQKGEWRPQTYAAFT
|
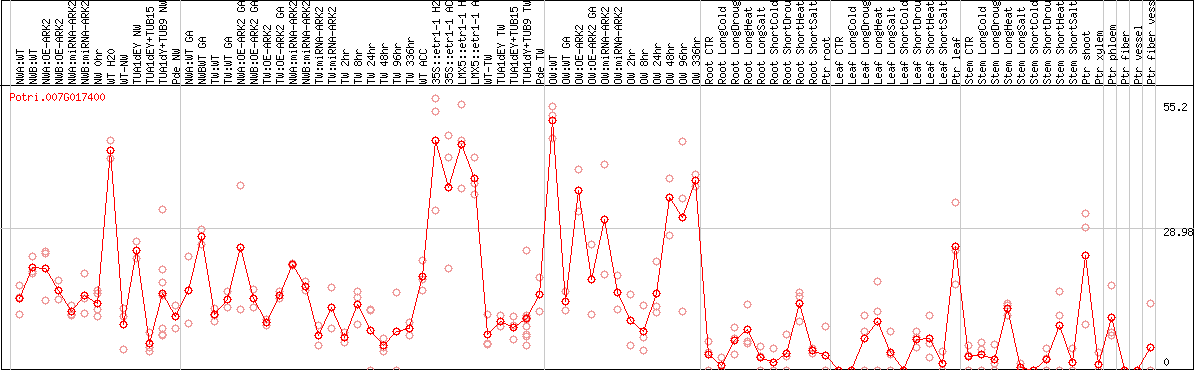
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G017400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
