Potri.007G024600 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.007G024600.1 pacid=42766805 polypeptide=Potri.007G024600.1.p locus=Potri.007G024600 ID=Potri.007G024600.1.v4.1 annot-version=v4.1
ATGGATTCCGATACTCTAACAGAACCGATGGACTTTAAAGTATCAGAATTAATAAACGAAGTTCAAATCGAACATTCTCCCTCTTTCACCAAGCTCGTAA
ACGACACCGTTTCATCTATCCAAAATTCCATCGACAAAATCCCTAACAACCTCGTGGTTACAGGCGAGGAAGCGGCGGGGTTCGTTAGAGACGTCGGAGC
GGATAAAGTGGAGTTTAAGTTTAAGAAGCCGAAATCGATTGCAATTGGTGGCAGTTATTCGATAAAGTGTGTCGTGAAGCCGGATGTTAGTGTTGACCTT
TTCATTCAATTGCCTAAGGAGTGTTTTCATGAGAAAGATTATTTAAATCATCGCTATCATGCGAAGAGGTTTGTTTATTTATGTGTGATTAACAAGTTCT
TGAAGTCGGATTCATCGTTTGAAAAGGTTGAATGGTCTACGCTTCAAAACGAGGCCCGGAAACCTGTGCTGCTTGTCTATCCAGCTGACAAGCTTGCCGA
AATTCCTGGGTTCTTTGTGAGGATAATTCCTACAGCGAAATCTTTGTTTAATACTGCAAAGTTGGACTTGAAACGGAACAATGTCCGTGTTTTGAATCAA
GGTGGGACTGCTTTGCCTACACCAAGATATAACAGTAGTATTTTGGAAGACATGTGTTTAGAGGACAATACAGAATTTCTCAAGAAAACTTTTCTTGGAC
AGAAGGCATTGGGAGAGGCTTTAGTTTTGCTGAAGGTTTGGGCTCGACAGAGAGATTCTATACATTCTCACGATAGCCTAAATGGATATTTAATAGCTAT
CATATTGTCATATCTTGTGGCTTATGAGAAGGTTAATAGTTCCATGAGGCCACTGCAAATATTTCGAGTCACCTTGGACTTCATAGCCAATTCAAAATTA
TGGACTCGGGGGCTTTTCTTACAAAAGCAAGGTGAGGTCAAGATATTGAAGGAGGATAGGATGCTGTATAAAGAATCATTTCCAGTTGTCATATTCGATT
CAACCACTCACATCAATTTAACTTTTCGGATAAAAGATAGTGGTTTCTCAGAGCTCCAAGACGAGGCTGCTCAGACACTTCAATGCTTTGGGAAATCTGG
AGATAGTGCATTTGAGGATATCTTCATGACCAAGATTGACTTTCCTGCCAGATATGATTATTGTGTAAGATTGAGCTTAAAAGGAAATAGTGAGTTCTAT
TCTTCTGGATATTGCTTGGACGAAGAATGTTGGAGATTGTATGAGAAAAAGGTGCAGAGTCTTCTATCTCAAGGACTGAGTGATCGAGCCAAGTCAATTC
GTGTTATTTGGAGAAATATCCCTTCAGGTTGCAGTCTGGAAAATGGTTTGTCAACACTTGATGGAGAACCATTGCTAGCTGGAATTTCTCTAAGCTCTTT
AGATAAAGCATTTCGAGTTGTTGATATTGGTCCGGATGCTGAAAATAAAGAAGAGGCTGCCAGGTTTCGGAAGTTTTGGGGGGAGAAAGCAGAGCTCAGG
AGATTTAAGGATGGCAAAATTGCTGAAAGCACAGTATGGGAAAGTGAGCAGTGGAAGAAACATCTCATTTTAAAAAGAATAGTAGAATATATACTTCTTC
GTCATCTTTCTATATCAAAAACCAGTATTGAGCAAACTGTGGATCAACTTGACTTTTCTCTTCTTCATGGTGTTGAAGATCCTATGTCATTTTCTGCAAG
TTTGCTTGGGGCATTTGATATCCTATCAAAGCGCTTGCGTCTTATAGAAGACATTCCCTTGAAGGTGTCCAGCGTGCAGCCTTTAGACCCAGCTTTCAGG
TTCACATCTGTCTTCCCTCCTGAACCACACCCAATAGCAAGTGAAAAGGGTAATGTTCCAAGACCACATAAGCTCACTTCATCCTGCATCCAGCCATTGG
AAGTTATGATTCAGTTGGAAGGTTCTGGGAACTGGCCAATGGATGATGTGGCAATTGAGAAAACCAAATCTGCTTTTCTTCTCAAAATTGGAGAGAGTCT
CGAGAATAGTTGGGGGATGACATGTACTGCGACTGAAGATGACGTGGATGTTTTTCTGTCTGGATATGCATTTCGTCTAAAAATTTTGCACGAGAGAGGC
TTGAGCTTGGTGAAAAGAGAAACTGGAAGTGACCAGGGAAAACAGGTCTCTTCTGCAGACCAGAAACTCTTTGTTCGCAGTCAGCATTCTAGCATGATTA
ATGGGTTGCAAGGTGTTTTCCCAATATATGGGCCAGTTGTTAGGCTTGCTAAAAGATGGGTTGCGTCACATATGTTCTCGGCATGCTTGTCAGAGGAGGC
AATTGAGTTATTGGTTGCGCATCTCTTTGTGAAGCCTTTGCCTTTCACTGCTCCTTGTTCACGCATCACTGGATTTCTAAGGTTCTTGCGATTGCTAGCA
GAGTATGATTGGACCTTTTCTCCTCTTATTGTTGACATAAATAGTGATTTCAACCCAAGTGATAAGAAAGAGATTTATGACAAGTTTATGCTGACTAGAA
AAGGTTATGAAGAAAGTAGCCAAAACATAAGTCCAGCAATGTTCTTGGCGACATCCTATGACAAGGCATCTGAAGCTTGGACCAGACTCTCACCAAACGT
ACTGGAACTTAAAAGGTTGGTGGCTTATGCTCGTAGCAGTGCAAACCTGCTTACCAGACTCGTATTTCAGGATCAGACAGAATCTTACAGATGGGAGTGC
CTTTTCTGCACTCCTCTTACCAACTATGATGCTGTAATCCTTCTCCATGGGGACAGATTACCATATCCTCAACGTCTTCTCTTCCCTTCAAAATTAAACC
ATGGGAGGCTTGTGGCTCGTGGGAATGCTAGCAAAGCTTTCCAGCCCTTTATGTTACCTGGAGATTTACGAGGGAGCTTAGACAAGTTGAAAAATAAGCT
ATTGGTGGACTTTGATCCACTTAGGTGCTACATTGCAGATCTGGAGAAAGAATGCAACACATTGAAGATGTGGTATGATTCTCTGGGAGGCGACGCCATT
GGACTAACATGGGAGAGATCTTGTTCAAAGAAGCGGGATCGAGAAGAAGCCAGCAGCGATGAAGACCCAATTGATGTCTTAAAGGCTGTGGGTGAAGCTG
GTAAACGGTTTGTAAAGAGTGTACATTTTCTCAAGGCTCCAAGGCTCATGAATTAA
|
|||||||||||||||
|
AA sequence
|
>Potri.007G024600.1 pacid=42766805 polypeptide=Potri.007G024600.1.p locus=Potri.007G024600 ID=Potri.007G024600.1.v4.1 annot-version=v4.1
MDSDTLTEPMDFKVSELINEVQIEHSPSFTKLVNDTVSSIQNSIDKIPNNLVVTGEEAAGFVRDVGADKVEFKFKKPKSIAIGGSYSIKCVVKPDVSVDL
FIQLPKECFHEKDYLNHRYHAKRFVYLCVINKFLKSDSSFEKVEWSTLQNEARKPVLLVYPADKLAEIPGFFVRIIPTAKSLFNTAKLDLKRNNVRVLNQ
GGTALPTPRYNSSILEDMCLEDNTEFLKKTFLGQKALGEALVLLKVWARQRDSIHSHDSLNGYLIAIILSYLVAYEKVNSSMRPLQIFRVTLDFIANSKL
WTRGLFLQKQGEVKILKEDRMLYKESFPVVIFDSTTHINLTFRIKDSGFSELQDEAAQTLQCFGKSGDSAFEDIFMTKIDFPARYDYCVRLSLKGNSEFY
SSGYCLDEECWRLYEKKVQSLLSQGLSDRAKSIRVIWRNIPSGCSLENGLSTLDGEPLLAGISLSSLDKAFRVVDIGPDAENKEEAARFRKFWGEKAELR
RFKDGKIAESTVWESEQWKKHLILKRIVEYILLRHLSISKTSIEQTVDQLDFSLLHGVEDPMSFSASLLGAFDILSKRLRLIEDIPLKVSSVQPLDPAFR
FTSVFPPEPHPIASEKGNVPRPHKLTSSCIQPLEVMIQLEGSGNWPMDDVAIEKTKSAFLLKIGESLENSWGMTCTATEDDVDVFLSGYAFRLKILHERG
LSLVKRETGSDQGKQVSSADQKLFVRSQHSSMINGLQGVFPIYGPVVRLAKRWVASHMFSACLSEEAIELLVAHLFVKPLPFTAPCSRITGFLRFLRLLA
EYDWTFSPLIVDINSDFNPSDKKEIYDKFMLTRKGYEESSQNISPAMFLATSYDKASEAWTRLSPNVLELKRLVAYARSSANLLTRLVFQDQTESYRWEC
LFCTPLTNYDAVILLHGDRLPYPQRLLFPSKLNHGRLVARGNASKAFQPFMLPGDLRGSLDKLKNKLLVDFDPLRCYIADLEKECNTLKMWYDSLGGDAI
GLTWERSCSKKRDREEASSDEDPIDVLKAVGEAGKRFVKSVHFLKAPRLMN
|
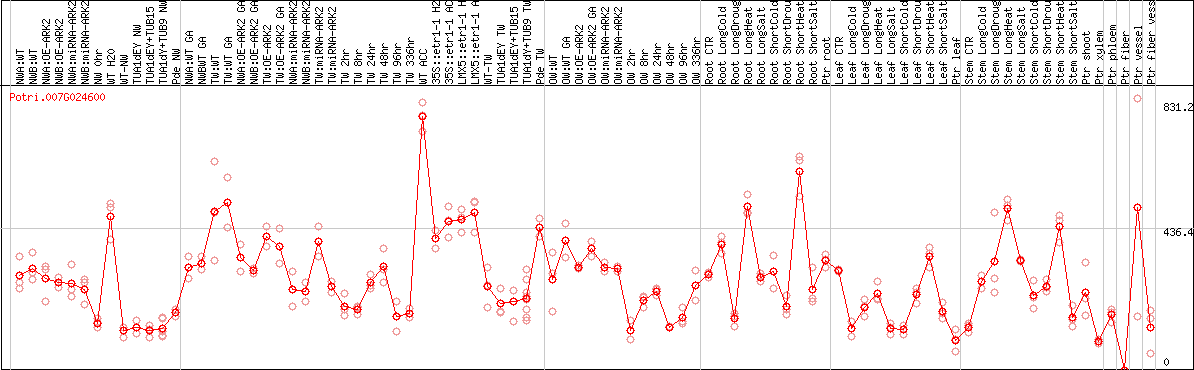
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G024600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
