External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G75170 409 / 9e-145
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT4G36640 396 / 7e-140
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT1G22180 271 / 3e-90
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT4G08690 249 / 6e-82
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT1G01630 93 / 3e-22
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT3G22410 82 / 2e-17
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT1G05370 78 / 3e-16
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT1G19650 76 / 4e-15
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT5G63060 72 / 2e-14
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT1G14820 70 / 7e-14
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G123200
572 / 0
AT1G75170 414 / 9e-147
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.007G025900
471 / 3e-169
AT1G75170 369 / 3e-129
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.002G261000
418 / 2e-148
AT1G75170 427 / 3e-152
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.013G068700
281 / 9e-94
AT1G22180 351 / 6e-121
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.002G093400
278 / 6e-92
AT4G08690 375 / 5e-130
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Potri.005G168100
107 / 9e-28
AT4G08690 134 / 4e-38
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Potri.017G063966
103 / 1e-25
AT1G01630 340 / 2e-118
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.003G162100
96 / 4e-23
AT1G01630 356 / 2e-125
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.001G068000
92 / 1e-21
AT1G01630 361 / 3e-127
Sec14p-like phosphatidylinositol transfer family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014312
457 / 5e-164
AT1G75170 397 / 4e-140
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10026020
459 / 2e-163
AT1G75170 397 / 4e-139
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10010804
414 / 6e-147
AT1G75170 393 / 7e-139
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10016312
407 / 2e-144
AT1G75170 387 / 2e-136
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10025128
374 / 3e-131
AT1G75170 380 / 3e-134
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10020911
263 / 2e-86
AT4G08690 376 / 1e-130
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10033464
261 / 3e-86
AT4G08690 378 / 4e-132
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10023956
164 / 6e-50
AT4G36640 166 / 1e-50
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
Lus10016733
111 / 7e-29
AT1G01630 325 / 2e-113
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10004684
107 / 3e-27
AT1G01630 326 / 2e-113
Sec14p-like phosphatidylinositol transfer family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0512
CRAL_TRIO
PF00650
CRAL_TRIO
CRAL/TRIO domain
CL0214
UBA
PF03765
CRAL_TRIO_N
CRAL/TRIO, N-terminal domain
Representative CDS sequence
>Potri.007G025800.4 pacid=42765298 polypeptide=Potri.007G025800.4.p locus=Potri.007G025800 ID=Potri.007G025800.4.v4.1 annot-version=v4.1
ATGTTTCGAAGAAAGCACTCTCAGAATCATCAGGAGAATGGCCTTGCAGAACAAGAAGCAAAGCAGGTCAGTGAACTCAGGGCTGCTCTAGGACCTCTTT
CTGATCGAAGTGTAAAGTACTGCACTGATGCATGCCTGAGGAGATATTTAATAGCGCGGAACTGGAATGTCGACAAGGCAAAGAAAATGTTGGAAGAGAC
ACTCAAGTGGAGAGCAACTTATAAGCCTGAAGAAATCCGCTGGCATGAAGTAGCTCATGAAGGCGAGACTGGTAAAGTCTCCAGAGCAGATTTTCATGAT
CGATCTGGGAGGACTGTGCTCATAATGAGACCAGGAATGCAGAACACAACATGTGCAGAAGACAATGTCCGCCATTTGGTTTATCTTATAGAGAACGGTA
TTCTTAACCTTGGTGAGGGTCAAGAACAAATGTCATGGTTGATAGACTTCACTGGATGGGGTTTGAGCGTCAAGGTCCCCATCAAAACTGCCCGTGAATG
CATTAACATCTTACAGAACCATTACCCTGAGAGACTTGCTGTTGCATTTCTTTATAATCCGCCTAGAATATTTGAAGCATTTTGGAAGGTTGTCAAGTTC
TTCCTGGATCCAATTACTATCCAGAAGGTGAAGTTTGTTTACCCAAAAAAAGAAAATAGTTTCGAGCTCATGAAGTCTTTCTTTGATGTTGACAACCTCC
CAAACGAGTTCGGGGGAAAAGCCACACTGACGTACGACCATGAAGAGTTTTCACGACTAATGGCCCAGGATGATGTAAAAACAGCCAAGTTCTGGGGATT
AGATGAAAAACCAAGCCACATTGCTAACAGTCACTCAGGGGCACAGGTGGCGCCAGAGCCAACACCTCTTGCACTGCCGGCTAGCTAA
AA sequence
>Potri.007G025800.4 pacid=42765298 polypeptide=Potri.007G025800.4.p locus=Potri.007G025800 ID=Potri.007G025800.4.v4.1 annot-version=v4.1
MFRRKHSQNHQENGLAEQEAKQVSELRAALGPLSDRSVKYCTDACLRRYLIARNWNVDKAKKMLEETLKWRATYKPEEIRWHEVAHEGETGKVSRADFHD
RSGRTVLIMRPGMQNTTCAEDNVRHLVYLIENGILNLGEGQEQMSWLIDFTGWGLSVKVPIKTARECINILQNHYPERLAVAFLYNPPRIFEAFWKVVKF
FLDPITIQKVKFVYPKKENSFELMKSFFDVDNLPNEFGGKATLTYDHEEFSRLMAQDDVKTAKFWGLDEKPSHIANSHSGAQVAPEPTPLALPAS
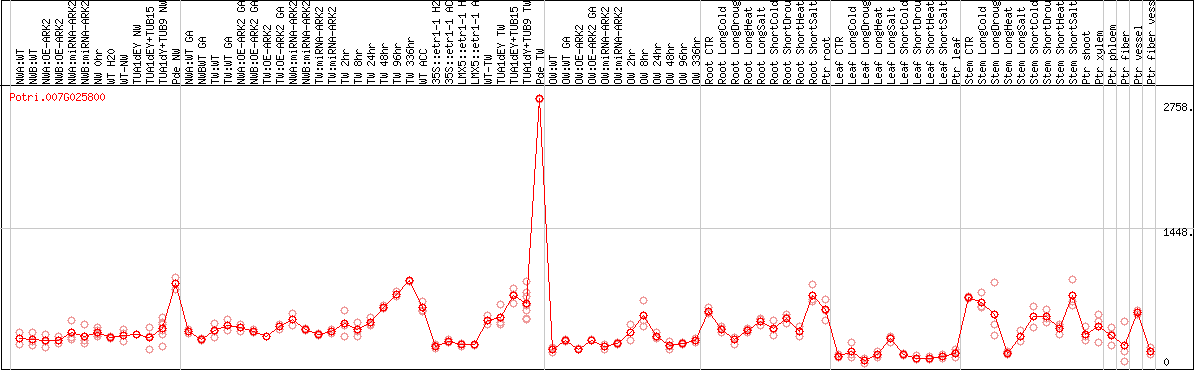
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G025800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.