External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G37200 305 / 5e-105
HCF164
HIGH CHLOROPHYLL FLUORESCENCE 164, Thioredoxin superfamily protein (.1)
AT1G43560 56 / 2e-09
ATY2
thioredoxin Y2 (.1)
AT1G50320 55 / 5e-09
ATHX, ATX
thioredoxin X (.1)
AT1G76760 54 / 5e-09
ATY1, TRX-Y1
thioredoxin Y1 (.1)
AT5G06430 52 / 5e-08
Thioredoxin superfamily protein (.1)
AT3G15360 52 / 7e-08
ATHM4, ATM4, TRX-M4
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
AT2G15570 51 / 1e-07
TRX-M3, GAT1, ATHM3, ATM3
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
AT1G21750 51 / 3e-07
ATPDI5, ATPDIL1-1
ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 5, PDI-like 1-1 (.1.2)
AT2G47470 50 / 5e-07
ATPDI11, ATPDIL2-1, UNE5, MEE30
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
AT1G19730 47 / 1e-06
ATTRX4, ATH4
thioredoxin H-type 4, Thioredoxin superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016780
305 / 7e-105
AT4G37200 300 / 4e-103
HIGH CHLOROPHYLL FLUORESCENCE 164, Thioredoxin superfamily protein (.1)
Lus10022477
303 / 3e-104
AT4G37200 298 / 2e-102
HIGH CHLOROPHYLL FLUORESCENCE 164, Thioredoxin superfamily protein (.1)
Lus10036337
58 / 1e-09
AT2G47470 548 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Lus10010274
58 / 2e-09
AT2G47470 549 / 0.0
UNFERTILIZED EMBRYO SAC 5, MATERNAL EFFECT EMBRYO ARREST 30, PDI-LIKE 2-1, ARABIDOPSIS THALIANA PROTEIN DISULFIDE ISOMERASE 11, thioredoxin family protein (.1.2.3.4)
Lus10014798
56 / 2e-09
AT3G15360 181 / 2e-58
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10040887
56 / 2e-09
AT3G15360 182 / 4e-59
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10018875
56 / 3e-09
AT1G76760 176 / 6e-57
thioredoxin Y1 (.1)
Lus10028569
56 / 3e-09
AT1G76760 174 / 3e-56
thioredoxin Y1 (.1)
Lus10029755
52 / 3e-08
AT3G15360 162 / 3e-51
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
Lus10042784
52 / 6e-08
AT3G15360 165 / 4e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00085
Thioredoxin
Thioredoxin
Representative CDS sequence
>Potri.007G034400.4 pacid=42766315 polypeptide=Potri.007G034400.4.p locus=Potri.007G034400 ID=Potri.007G034400.4.v4.1 annot-version=v4.1
ATGGCTCGCGTGGCTTCAAACCCTCTCTGTATCCATAGATTCCGGCCATATCTTCAAGCCTCTCATTCATCACCATCACTCCTCAAATCCACTGTCCACT
ATCAAACCAAACCTCGCAAATTCCCTGCTGTTTCTTGCCAAACAAACCCCGACTCATCTGAAACCTCCACACCAACGGAAAAAGAGGTCCTTGAACCTGG
TCCAGTTAGTGATGATACCAAAAGAGAGGATACAGGTGCGTCCCAAGATTCTGGGTTACCTGAATTTCCAAATAAAGATGTTAATAAACGAATAGCAGTG
GTTTCTCTTCTTGCTGCTGCCGGACTTTTCTTGTCTTCAAGGTTGGATTTTGGTGTTTCTTTGAAGGACTTATCTGTTGCTGCTTTGCCTTATGAAGAGG
CCCTCTCAAATGGGAAGCCAACTGTTGTGGAGTTCTATGCAGATTGGTGTGAAGTATGTAGAGAGTTAGCTCCAGATGTCTACAAAGTAGAGCAGCAGTA
CAAGGACCGTGTGAATTTTGTTATGCTGAACGTTGATAATACCAAGTGGGAACAAGAACTTGATGAGTTTGGTGTTGAGGGTATTCCGCACTTTGCATTC
CTGGATAGAGAGGGGAATGAAGAGGGTAATGTAGTTGGCAGGCTTCCAAGGAAGTACCTACTTGAGAATGTTGAAGCTCTTGCACGTGGGGAAGGATCCA
TACCTCATGCCCGAGTTGTTGGACAATATTCAAATGCAGAAAACAGGAAGGTGCGCTCTGTTGATCCTAGAAGCCATGGGTAG
AA sequence
>Potri.007G034400.4 pacid=42766315 polypeptide=Potri.007G034400.4.p locus=Potri.007G034400 ID=Potri.007G034400.4.v4.1 annot-version=v4.1
MARVASNPLCIHRFRPYLQASHSSPSLLKSTVHYQTKPRKFPAVSCQTNPDSSETSTPTEKEVLEPGPVSDDTKREDTGASQDSGLPEFPNKDVNKRIAV
VSLLAAAGLFLSSRLDFGVSLKDLSVAALPYEEALSNGKPTVVEFYADWCEVCRELAPDVYKVEQQYKDRVNFVMLNVDNTKWEQELDEFGVEGIPHFAF
LDREGNEEGNVVGRLPRKYLLENVEALARGEGSIPHARVVGQYSNAENRKVRSVDPRSHG
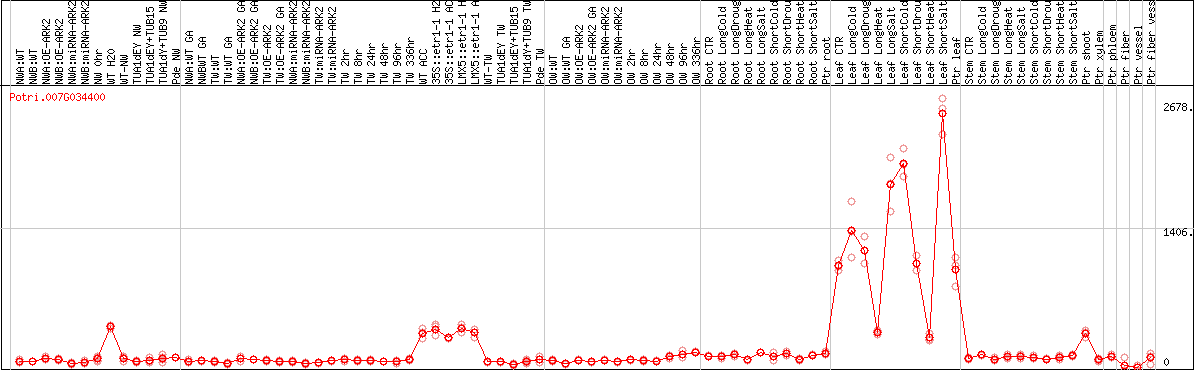
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G034400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.