External link
Symbol
PCBER5
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39230 407 / 9e-144
NmrA-like negative transcriptional regulator family protein (.1)
AT1G75280 368 / 3e-128
NmrA-like negative transcriptional regulator family protein (.1)
AT1G75290 360 / 6e-125
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G75300 357 / 8e-124
NmrA-like negative transcriptional regulator family protein (.1)
AT1G19540 317 / 3e-108
NmrA-like negative transcriptional regulator family protein (.1)
AT4G34540 254 / 3e-83
NmrA-like negative transcriptional regulator family protein (.1)
AT1G32100 214 / 8e-68
ATPRR1
pinoresinol reductase 1 (.1)
AT4G13660 214 / 2e-67
ATPRR2
pinoresinol reductase 2 (.1)
AT5G18660 47 / 8e-06
PCB2, DVR
PALE-GREEN AND CHLOROPHYLL B REDUCED 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G33360 42 / 0.0003
FLDH
farnesol dehydrogenase, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2), NAD(P)-binding Rossmann-fold superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G168400
465 / 5e-166
AT4G39230 448 / 2e-159
NmrA-like negative transcriptional regulator family protein (.1)
Potri.009G118100
451 / 5e-161
AT4G39230 501 / 0.0
NmrA-like negative transcriptional regulator family protein (.1)
Potri.009G118300
423 / 7e-150
AT4G39230 488 / 8e-176
NmrA-like negative transcriptional regulator family protein (.1)
Potri.005G228700
384 / 2e-134
AT1G75280 451 / 3e-161
NmrA-like negative transcriptional regulator family protein (.1)
Potri.002G034400
379 / 1e-132
AT1G75280 467 / 2e-167
NmrA-like negative transcriptional regulator family protein (.1)
Potri.004G156650
293 / 7e-100
AT4G39230 322 / 1e-111
NmrA-like negative transcriptional regulator family protein (.1)
Potri.009G118000
278 / 1e-92
AT4G34540 432 / 7e-154
NmrA-like negative transcriptional regulator family protein (.1)
Potri.007G123700
253 / 1e-82
AT1G75280 284 / 4e-95
NmrA-like negative transcriptional regulator family protein (.1)
Potri.013G103701
244 / 2e-79
AT1G75280 271 / 4e-90
NmrA-like negative transcriptional regulator family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026350
407 / 2e-143
AT4G39230 454 / 3e-162
NmrA-like negative transcriptional regulator family protein (.1)
Lus10042311
397 / 2e-139
AT4G39230 489 / 4e-176
NmrA-like negative transcriptional regulator family protein (.1)
Lus10026351
395 / 2e-138
AT4G39230 481 / 4e-173
NmrA-like negative transcriptional regulator family protein (.1)
Lus10026348
387 / 8e-136
AT4G39230 442 / 2e-157
NmrA-like negative transcriptional regulator family protein (.1)
Lus10040442
379 / 1e-132
AT4G39230 462 / 2e-165
NmrA-like negative transcriptional regulator family protein (.1)
Lus10023557
373 / 5e-128
AT4G39230 461 / 2e-162
NmrA-like negative transcriptional regulator family protein (.1)
Lus10042313
315 / 7e-108
AT4G39230 369 / 2e-129
NmrA-like negative transcriptional regulator family protein (.1)
Lus10023558
281 / 8e-94
AT4G34540 405 / 7e-143
NmrA-like negative transcriptional regulator family protein (.1)
Lus10042312
232 / 3e-75
AT4G39230 253 / 1e-83
NmrA-like negative transcriptional regulator family protein (.1)
Lus10040443
212 / 5e-67
AT4G34540 290 / 5e-98
NmrA-like negative transcriptional regulator family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01370
Epimerase
NAD dependent epimerase/dehydratase family
Representative CDS sequence
>Potri.007G036500.1 pacid=42765497 polypeptide=Potri.007G036502.1.p locus=Potri.007G036500 ID=Potri.007G036500.1.v4.1 annot-version=v4.1
ATGGCAGATAAAAGCAAGATTCTACTCATTGGAGGCACAGGATACATAGGAAAATTCATTGTAGAAGCAAGTATCAAGGAAGGCCACCCAACTTTTGTTC
TTGTCAGAGAGTCCACTCTCTCTCTCTCTAGCCCTGCAAAATCCATTGTCATTAACAACTTCAATAATCTTGGTGTCAATTTCCTAATTGGAGACTTGTT
GGATCATGAGAGTTTGGTGAAGGCAATAAAGCAAGTGGATGTTGTGATATCTTCCATTGCTCATGATCAGGTAGATAATCAAGTCAACATCATTGCTGCC
ATTAAAGAATCTGGAAATATCAAGAGATTTATCCCATCAGAGTTTACAAATGATGTGGATAGAGCGCATATTGTTGAACCAGCAACAGGATTGGCTGCTT
CAAAGGCTAAAATTCGACGAGCTATTGGGGCTGAAGGAATTCCTTACACACATGTGTGGCATCTAATTCTTTCGCTGGCTTCAAGCATTAGGAGCACTGC
CGTAACAACAACTTCGAACCAACCTGGAGCTACTGCTTCCCATGGAGATAGACTTGTCATTTTAGGTGATGGAAATGTAAAAGTGGTTTTCAACAAGGAA
GAGGACATTGCCACATACACTATCAAAGCAGTGGATGATCCAAGAACAGTGAACAAAATCCTCTACATTAAACCCCCAGCCAACATTATCTCATCCAATG
AACTTGTTTCTTTGTGGGAGAAGAAGATAGGGAAAACAATTGAAAGGGTCTATGTTCCAGAGGAGCAAATTTTGAAGAATATTCAAGAAACTTCAGACTT
TCTACGCAAACTGACTTTAGCAATTTGCCACTCTTGGTTTGTGAATGGAGATCAGACAAACTTTGAGATTGAGCCATCATTTGGTGTAGAGGCTTCTGAG
CTATACCCTGATGTCAAATACACTACTGTGGATGAATACCTTAACCAGTTTGTTTGA
AA sequence
>Potri.007G036500.1 pacid=42765497 polypeptide=Potri.007G036502.1.p locus=Potri.007G036500 ID=Potri.007G036500.1.v4.1 annot-version=v4.1
MADKSKILLIGGTGYIGKFIVEASIKEGHPTFVLVRESTLSLSSPAKSIVINNFNNLGVNFLIGDLLDHESLVKAIKQVDVVISSIAHDQVDNQVNIIAA
IKESGNIKRFIPSEFTNDVDRAHIVEPATGLAASKAKIRRAIGAEGIPYTHVWHLILSLASSIRSTAVTTTSNQPGATASHGDRLVILGDGNVKVVFNKE
EDIATYTIKAVDDPRTVNKILYIKPPANIISSNELVSLWEKKIGKTIERVYVPEEQILKNIQETSDFLRKLTLAICHSWFVNGDQTNFEIEPSFGVEASE
LYPDVKYTTVDEYLNQFV
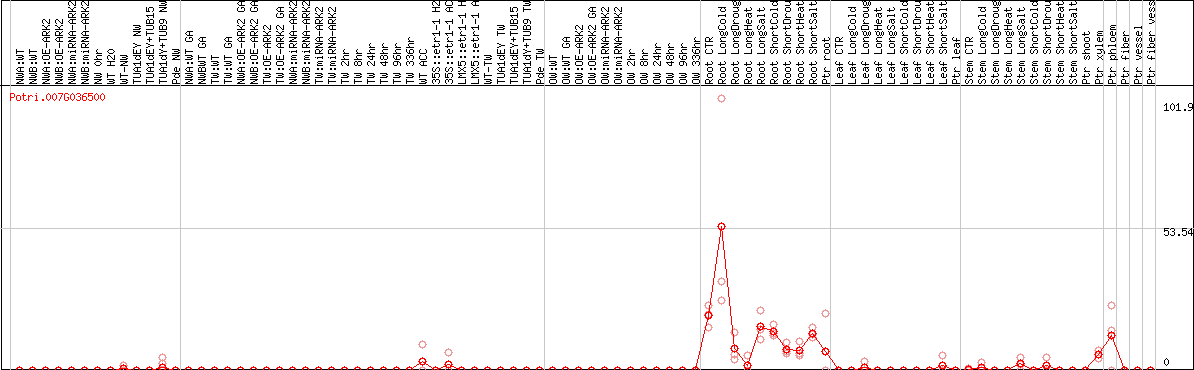
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G036500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.