Potri.007G041000 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.007G041000.1 pacid=42765055 polypeptide=Potri.007G041000.1.p locus=Potri.007G041000 ID=Potri.007G041000.1.v4.1 annot-version=v4.1
ATGGAGTGGACAACTGTGCAGCATCTGGATCTACGACATGTCGCTCGTGGTTTTCACAGACCATTACAACCTCATGCTGCTGCTTTTCACCCTACTCAAA
CTCTCATTGCCGCCGCGATCGGAACTTACATCATCGAATTTGATGCAGTAACAGGAAGCAAATTATCTTCCATTGACATTGGTGCATCCGTTCTTCGCAT
GGCTTACAGTCCTAATACTAGCCATGCTGTTATTGTTATGGTTGAGGATGGTACGATTCGGTCGTGTGATTTTGATACGGAGCAAAGTTGGGTTTTGCAT
TCGCCTGAAAAGAAAATGGAACCCTTATCTTTTGATACGGAAGTTCATATGGCTTTGACCCCATTGCAACCTGTTGTTTTTTTCGGCTTCCACCGCAGGA
TGAGCGTAACTGTGGTTGGAACGGTTGATGGAGGAAGAGCACCCACAAAAATAAAGACTGATCTGAAGAAGCCTATTGTAAATCTTGCTTGCCATACGCG
CCATCCTGTCCTGTATGTTGCTTATGCAGATGGTTTGATTCGAGCTTACAATATTCACTCATATGCTGTTCATTACACATTACAACTTGATAATTCCATT
AAGCTAATTGGTGCTGGTGCATTTGCTTTTCATCCAACGTTGGAATGGATTTTTGTTGGTGATAGACGTGGTACCCTTCTGGCATGGGATGTCTCAACAG
AGAGACCTAGTATGATTGGAATAACACAAGTGGGCTCCCAACCAATCACATCAATTGCTTGGCTTCCAGCGTTGCGGTTACTTGTTACTGTTTCCAAGGA
TGGGACTTTACAAACGTGGAAAACACGAGTGATACTGAATCCCAATAGACCTCCAATGCAAGCAAATTTTTTCGAGCCTGCTGGAATTGAATCAATTGAC
ATCCCGCGTATTTTATCTCAACAGGGTGGAGAAGCAATTTACCCTTTACCGAAAATTAAAGCCTTAGAAGCTCACCCCAAACTAAATTTAGCAGCTCTTT
TATTTGCAAATATGACTGGTGTTGACAATGTAAAAAGTAGGACTGCTTACACTAGGGATGGAAGGAAACAGCTTTTTGCTGTTCTGCAAAGTGCAAGGGG
ATCATCAGCTTCTGTTTTGAAAGAAAAGCTCTCATCATTGGGCTCATCAGGAATTCTGGCAGACCATCAACTTCAAGCACAACTGCAAGAGCATCACCTC
AAGGGCCAAAGTCAGCTGACAATTTCAGACATTGCAAGGAAGGCATTCCTTTACAGTCATTTCATGGAAGGCCATGCCAAAAGTGCTCCCATATCTCGGC
TGCCTCTCATCACTATTTTGGATACCAAACATCATCTGAGGGACATTCCTGTTTGCCAGCCTATTCATTTGGAGCTAAATTTCTTTAACAAAGAGAACAG
AGTACTTCATTATCCTGTCAGGGCGTTTTATCTTGATGGCTTAAACCTCATGGCATATAATTTCTGCTCTGGTGTGGATAATATATACAAGAAACTCTAC
ACATCGATTCCTGGAAACGTGGAATACCAAGCGAAGCACATGGTGTACAGTATAAAACAACATTTATTTCTTGTTGTTTATGAATTCAGTGGCTCTGCAA
ATGAAGTAGTACTATATTGGGAAAATACTAATGCTCAGCCAGCCAACAACAAAGGAAGCACAATCAAAGGTCGAGATGCAGCATTTATTGGTCCAAGTGA
AAGTCAATTTGCAATTCTTGATGAAGACAAGACTGGAGTGGCTCTGTATATTCTGCCAGGAGGAGCTTCAAAAGAAGCTGGTGAAAAAAACTTATTACTC
GAAGAAAACCATTTTGCTGAAACAAATGGTGCCTCTCTTCGGGGTCCTATGCAGTTCCTGTTTGAAAGCGAAGTTGATCGTATCTTTACCACGCCACTAG
AGTCAACTTTGATGTTTGCTTCTACCGGGAGCCATATTGGCTTTGCAAAGATGGTGCAAGGATACCGGCTCTCAACTTCAGATGGTAATTATATATCAAC
AAAAACTGAAGGGAAAAAGTCAATCAAGTTGAAAGTGAATGAGATTGTCCTCCAGGTGCATTGGCAAGAAACTCTTCGAGGCTATGTTGCAGGGATATTA
ACCACACACAGGGTGCTTATGGTTTCAGCGGATCTTGATATACTGGCAAGCAGTTCCACAAAATTTGATAAAGGACTTCCTTCATTTAGATCCCTTTTGT
GGCTTGGACCTGCTCTTCTTTTTTCTACTGCCACTGCGATTAGTGTGCTTGGTTGGGATGGAATTGTGAGGACAATTCTTTCTGTTAGCTTGCCCTATGC
AGTTTTGGTTGGTGCTCTGAACGATAGATTGGTGCTTGCTAACCCAACAGATGTTAATCCCAGACAAAAGAAAGGGGTTGAGATTAAGAGCTGTCTTGTT
GGACTCCTTGAACCTCTTCTTATCGGATTTGCCACAATGCAACACACTTTTGAGCAGAAGCTTGACCTGTCAGAAATACTGTACCAAATAACATCAAGGT
TTGACAGCTTGCGTATTACACCAAGATCTCTTGATATTCTTGCTAGAGGTCCTCCTGTTTGTGGAGATCTTGCAGTTTCATTATCCCAAGCTGGTCCACA
GTTCACTCAGGTGTTGCGGGGTGTTTATGCTATCGAAGCTCTCCGTTTTTCTACTGCTTTAGATGTTTTGAAGGATGAGTTCTTGCGTTCTAGAGATTAT
CCAAAATGTCCTCCAACATCACACTTATTCCACCGATTTCGACAGTTGGGATATGCCTGCATCAAGTATGGCCAGTTCGATAGTGCAAAAGAAACTTTTG
AAGTCATTGCAGATTACGAAGGCATGCTTGATTTGTTTATATGCCACCTTAACCCCAGTGCCATGCGGCGCCTTGCTCAGAAACTGGAAGAAGAAGGTTT
GGATTCACAATTGAGACGGTACTGTGAGAGGATATTAAGAGTTCGCTCTACTGGGTGGACACAAGGCATTTTTGCCAACTTTGCTGCTGAGAGCATGGTT
CCCAAAGGTCCTGAATGGGGTGGTGGGAACTGGGAAATTAAAACTCCTACTAATTTGAAGAGTATACCTCAGTGGGAGCTGGCTGGAGAAGTGATGCCAT
ACATGAAGACTGATGATGGTACCATCCCGGCCATCATTACAGATCATATTGGTGTTTACCTGGGTTCAATTAAAGGAAGAGGGAATGTTGTTGAAGTAAG
GGAAGACAGTTTGGTCAAAGCATTTATCCCTGCTGGTGACAATAAGCCAAATGGGCTTCCAAATGCTTTAGCCAAATCCATATCCAATAAGTCAAATGGG
CTGCCTGATGGTCACATGAAGCTTGATTCTTTGTTGGGTCTGGAAACTCTGACCAAGCAAAACGCAGGTACATCTGCTGCTGATGAACAGGCAAAAGCAG
AAGAGGAATTCAAGAAAACTATGTATGGAACTGCTAATGATGGTAGCAGCAGTGACGAGGAAGGAGTGTCAAAAACAAAGAAGCTGCAAATTAGAATTCG
GGATAAGCCAGTTTCATCTACTACCGTGGATGTCAATAAGATCAAAGAAGCCACAAGACAGTTTAAACTTGGTGATGGTTTAGGCCCACCCATGAGGACA
AAGTCATTAACTGGTTCCCAGGACCTTGGTCAAATTTTATCCCAACCTCCTGCAACCACTGCCCCAGTTTCTGCCTCTGCTGATATGTTTGTTACTGATT
CATTAATGCAACCTGCGCCAGTATCACAACCAGGTCCTATGGTTATGGGTGGAGGAGTTACAGCTAGACCCATCCCAGAGGACTTCTTCCAGAACACAAT
ACCCTCCCTGCAAGTTGCAGCATCTCTGCCACCTCCAGGAACTTACCTTGCAAAATTGGATCAAGTTTCTCAAGGGGTTGGAAGCAACAATGCAGGTGGT
ATTCCCAACCCAGGTGCTGCTTCTGTGAGTGATATTGGTCTTCCTGATGGTGGTATTCCACCACAAGCAACTCAACTAGCTGCCCCACTTGCGTCCATTG
GACTTGCTGATGGTGGTGTCCCACCTCAAGCTTCCATTCAGGCTGGTATTCCACCTCAGCCACAAGTTCAAGCGCCTCAGGTTCCCCTTTCCACACAACC
TCTTGATCTTAGTGTTCTTGGAGTTACAGATTCAGGAAAAACTCCTGCCCCTGCATCTCTGCCATCATCTGTGAGACCTGGACAGGTTCCCCGAGGAGCA
GCTGCACCAGTATGTTTCAAGACTGGACTTGCACACCTTGAACAGAATCAACTTCCTGATGCCTTGTCATGTTTTGATGAAGCTTTCCTGGCACTAGCTA
AGGATAACTCTCGTGGTGCTGATATTAAAGCTCAAGCAACCATCTGTGCTCAGTACAAGATAGCAGTTACTCTTCTCAAGGAAATAGCACGGCTGCAGAA
AGTACAAGGGCCAAGTGCACTCAGTGCAAAAGATGAGATGGCAAGATTGTCACGGCATCTGGGTTCTTTGCCTCTTCTGGCAAAGCACCGAATAAATTGC
ATCCGAACTGCCATTAAGCGAAACATGGAAGTGCAGAACTTTGCTTATGGCAAGCAGATGCTTGAGCTCCTCATTTCCAAAGCTCCATCAAGTAAGCAGG
ATGAGTTGAGAAGCCTGATTGACATGTGTGTTCAGAGGGGTTCATCCAACAAGTCTATTGATCCTTTGGAAGATCCTTCCCATTTCTGTGCTGCCACACT
CAGCCGTCTTTCAACAATCGGATATGATGTTTGCGATCTCTGTGGAGCCAAATTTTCAGCTCTCTCTGCACCTGGATGCATCATCTGTGGAATGGGAAGT
ATTAAGAGATCAGATGCCCTCGCAGGGCCTGTTCCTTCACCATTTGGCTGA
|
|||||||||||||||
|
AA sequence
|
>Potri.007G041000.1 pacid=42765055 polypeptide=Potri.007G041000.1.p locus=Potri.007G041000 ID=Potri.007G041000.1.v4.1 annot-version=v4.1
MEWTTVQHLDLRHVARGFHRPLQPHAAAFHPTQTLIAAAIGTYIIEFDAVTGSKLSSIDIGASVLRMAYSPNTSHAVIVMVEDGTIRSCDFDTEQSWVLH
SPEKKMEPLSFDTEVHMALTPLQPVVFFGFHRRMSVTVVGTVDGGRAPTKIKTDLKKPIVNLACHTRHPVLYVAYADGLIRAYNIHSYAVHYTLQLDNSI
KLIGAGAFAFHPTLEWIFVGDRRGTLLAWDVSTERPSMIGITQVGSQPITSIAWLPALRLLVTVSKDGTLQTWKTRVILNPNRPPMQANFFEPAGIESID
IPRILSQQGGEAIYPLPKIKALEAHPKLNLAALLFANMTGVDNVKSRTAYTRDGRKQLFAVLQSARGSSASVLKEKLSSLGSSGILADHQLQAQLQEHHL
KGQSQLTISDIARKAFLYSHFMEGHAKSAPISRLPLITILDTKHHLRDIPVCQPIHLELNFFNKENRVLHYPVRAFYLDGLNLMAYNFCSGVDNIYKKLY
TSIPGNVEYQAKHMVYSIKQHLFLVVYEFSGSANEVVLYWENTNAQPANNKGSTIKGRDAAFIGPSESQFAILDEDKTGVALYILPGGASKEAGEKNLLL
EENHFAETNGASLRGPMQFLFESEVDRIFTTPLESTLMFASTGSHIGFAKMVQGYRLSTSDGNYISTKTEGKKSIKLKVNEIVLQVHWQETLRGYVAGIL
TTHRVLMVSADLDILASSSTKFDKGLPSFRSLLWLGPALLFSTATAISVLGWDGIVRTILSVSLPYAVLVGALNDRLVLANPTDVNPRQKKGVEIKSCLV
GLLEPLLIGFATMQHTFEQKLDLSEILYQITSRFDSLRITPRSLDILARGPPVCGDLAVSLSQAGPQFTQVLRGVYAIEALRFSTALDVLKDEFLRSRDY
PKCPPTSHLFHRFRQLGYACIKYGQFDSAKETFEVIADYEGMLDLFICHLNPSAMRRLAQKLEEEGLDSQLRRYCERILRVRSTGWTQGIFANFAAESMV
PKGPEWGGGNWEIKTPTNLKSIPQWELAGEVMPYMKTDDGTIPAIITDHIGVYLGSIKGRGNVVEVREDSLVKAFIPAGDNKPNGLPNALAKSISNKSNG
LPDGHMKLDSLLGLETLTKQNAGTSAADEQAKAEEEFKKTMYGTANDGSSSDEEGVSKTKKLQIRIRDKPVSSTTVDVNKIKEATRQFKLGDGLGPPMRT
KSLTGSQDLGQILSQPPATTAPVSASADMFVTDSLMQPAPVSQPGPMVMGGGVTARPIPEDFFQNTIPSLQVAASLPPPGTYLAKLDQVSQGVGSNNAGG
IPNPGAASVSDIGLPDGGIPPQATQLAAPLASIGLADGGVPPQASIQAGIPPQPQVQAPQVPLSTQPLDLSVLGVTDSGKTPAPASLPSSVRPGQVPRGA
AAPVCFKTGLAHLEQNQLPDALSCFDEAFLALAKDNSRGADIKAQATICAQYKIAVTLLKEIARLQKVQGPSALSAKDEMARLSRHLGSLPLLAKHRINC
IRTAIKRNMEVQNFAYGKQMLELLISKAPSSKQDELRSLIDMCVQRGSSNKSIDPLEDPSHFCAATLSRLSTIGYDVCDLCGAKFSALSAPGCIICGMGS
IKRSDALAGPVPSPFG
|
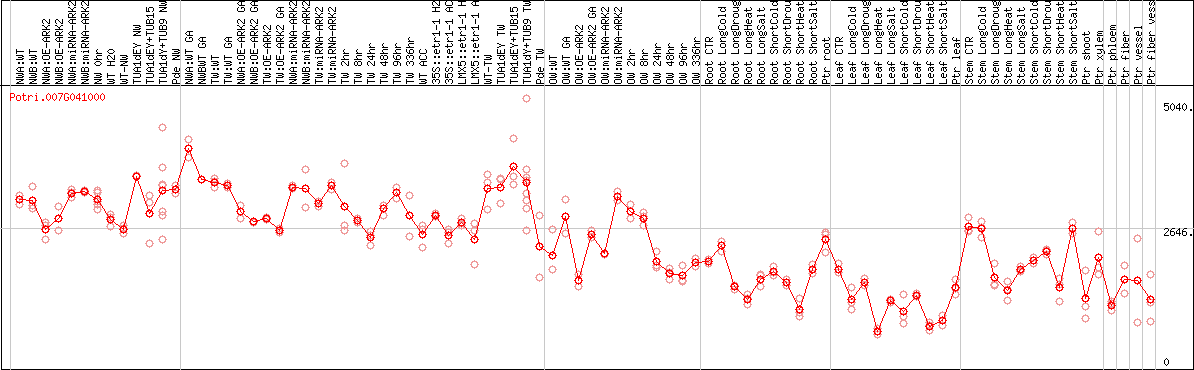
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G041000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
