External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G42240 334 / 6e-115
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT3G13700 177 / 9e-54
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT3G21215 136 / 2e-37
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G30260 51 / 3e-07
U2B''
U2 small nuclear ribonucleoprotein B (.1)
AT5G19350 50 / 2e-06
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G02840 48 / 4e-06
ATSRP34, SR1, SR34, At-SR34
Serine/Arginine-Rich Protein Splicing Factor 34, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT1G76940 47 / 4e-06
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G47580 47 / 5e-06
U1A
spliceosomal protein U1A (.1)
AT1G06960 42 / 0.0004
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G035000
184 / 2e-56
AT3G13700 348 / 3e-121
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.008G009400
143 / 5e-40
AT3G21215 219 / 4e-69
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.010G249700
135 / 5e-37
AT3G21215 376 / 9e-131
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.006G224900
52 / 1e-07
AT1G21320 122 / 2e-33
nucleotide binding;nucleic acid binding (.1.2)
Potri.012G019300
51 / 2e-07
AT1G76940 167 / 2e-51
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.015G009000
49 / 1e-06
AT1G76940 170 / 8e-53
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.014G127800
47 / 4e-06
AT2G47580 347 / 6e-122
spliceosomal protein U1A (.1)
Potri.002G070200
47 / 9e-06
AT1G76940 215 / 4e-70
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.013G153700
45 / 4e-05
AT2G30260 297 / 8e-103
U2 small nuclear ribonucleoprotein B (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034844
119 / 4e-31
AT3G21215 370 / 1e-128
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10033388
109 / 3e-27
AT3G21215 337 / 2e-115
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10028598
69 / 6e-13
AT1G76940 135 / 9e-38
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10028599
52 / 1e-07
AT1G21320 206 / 4e-66
nucleotide binding;nucleic acid binding (.1.2)
Lus10018903
52 / 2e-07
AT1G21320 182 / 5e-54
nucleotide binding;nucleic acid binding (.1.2)
Lus10020656
45 / 4e-05
AT1G02840 297 / 2e-100
Serine/Arginine-Rich Protein Splicing Factor 34, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Lus10029885
45 / 6e-05
AT1G02840 253 / 2e-81
Serine/Arginine-Rich Protein Splicing Factor 34, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Lus10042240
44 / 7e-05
AT2G30260 352 / 2e-124
U2 small nuclear ribonucleoprotein B (.1)
Lus10026413
42 / 0.0002
AT2G30260 350 / 1e-123
U2 small nuclear ribonucleoprotein B (.1)
Lus10000904
42 / 0.0004
AT5G46840 305 / 9e-102
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.007G046600.6 pacid=42765977 polypeptide=Potri.007G046600.6.p locus=Potri.007G046600 ID=Potri.007G046600.6.v4.1 annot-version=v4.1
ATGGAGGAAAACATGGCCGCATACTACCCGCCACCACCACCACCACCACCGGGTCACTACCCAACCTACTACCAACCGCAACCTCCGCCACCAACGGTGG
CACCTATTCCACCTCCACCTCCTCATCACCTCCCTTACATACCTCATCCACAACCGCCACCGGCACCTTTCGTATCCTACGTCAGTGTTCCTGCTTATGT
CCCACACGATCAGGTCCGTACTCTCTTTGTTGCTGGCCTTCCTGATGACATCAAGCCCCGCGAAATGTACAATTTATTCCGCGAGTTCCCTGGCTACGAA
TCTTCTCATCTCCGTACCCCCTCTCAAAATTCTCAGCCATTTGCCTTTGCTACATTCACTGACCAGCCGTCCGCGGTTGCAGCCATGCATGCTCTGAATG
GGATGGTTTTCGATCTTGAAAAGGGATCTACCCTATATATTGATCTAGCAAAATCTAATTCTAGATCTAAGCGCTCAAGAACAGATGATGAATGGTCCAG
TTTGGATAAAAAGGCTAGGGTTTCCTCTGGATTCTCGATGGGCACTCCTGATTCAGCAGGTTTTGGCAGCGTTCACTTGCCTGGAATGGCTAATTCTGCT
TTCAACACGATTGGTTTTCCATCTGCACAAAGTCCTGGAAGCATCGATGACAGATCTAGAAATGAGTCAAAAGCAGGAAGAATGAATAATTCTTCAGCAC
CTCCATGCCCAACACTTTTTGTGGCTAATCTGGGACAAAATTGTACAGAGGAGGAGCTAATTCAAGTATTCTCAAGATGCCCTGGATTTCTGAAATTGAA
GATGCAGAGCACATATGGGGCTCCAGTCGCATTTGTTGATTTCCAGGATACTGCCAGCTCAACTGGGGCACTAAATCATCTGCAAGGCACCGTTCTCTAT
TCATCAGTTGCTGGCGAGGGCTTGCGGTTGGAATATGCAAAATCACGCATGGGAATGCGGAGGAAACCAAGGTAA
AA sequence
>Potri.007G046600.6 pacid=42765977 polypeptide=Potri.007G046600.6.p locus=Potri.007G046600 ID=Potri.007G046600.6.v4.1 annot-version=v4.1
MEENMAAYYPPPPPPPPGHYPTYYQPQPPPPTVAPIPPPPPHHLPYIPHPQPPPAPFVSYVSVPAYVPHDQVRTLFVAGLPDDIKPREMYNLFREFPGYE
SSHLRTPSQNSQPFAFATFTDQPSAVAAMHALNGMVFDLEKGSTLYIDLAKSNSRSKRSRTDDEWSSLDKKARVSSGFSMGTPDSAGFGSVHLPGMANSA
FNTIGFPSAQSPGSIDDRSRNESKAGRMNNSSAPPCPTLFVANLGQNCTEEELIQVFSRCPGFLKLKMQSTYGAPVAFVDFQDTASSTGALNHLQGTVLY
SSVAGEGLRLEYAKSRMGMRRKPR
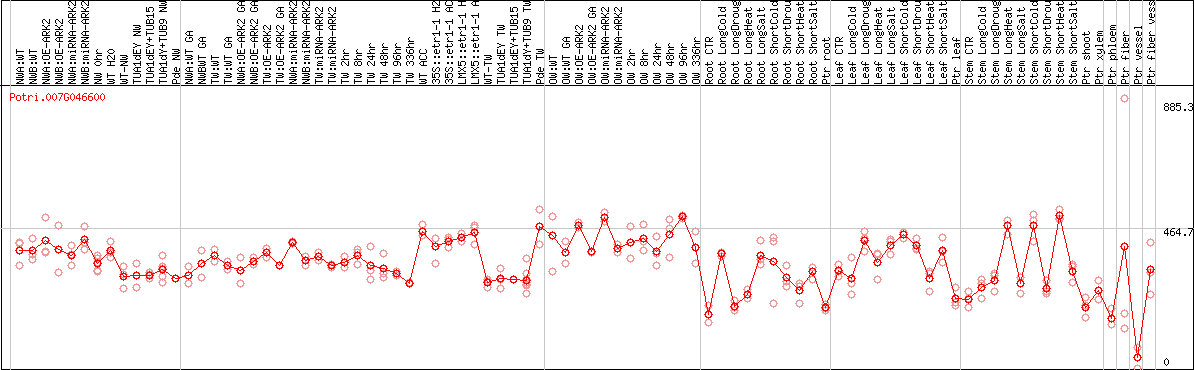
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G046600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.