Potri.007G063200 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G063200.4 pacid=42765861 polypeptide=Potri.007G063200.4.p locus=Potri.007G063200 ID=Potri.007G063200.4.v4.1 annot-version=v4.1
ATGACAAACTCGAATAATACGGATCTATTCGATTCGTTTTTCAGAAGAGCAGATTTAGACGGTGATGGTCAGATCAGTGGTGCTGAAGCCGTCGGTTTCT
TTCAAGGTTCCGGTTTGCCTAAGCACGTCCTTGCTCAGGTGTGGATGCATGCTGACCAGAGGAAAGCAGGCTACCTCGGTCGACAAGAGTTCTATAATGC
ACTTAAACTTGTTACTGTGGCACAGAGTAAGCGAGAGTTGACCCCAGAAATAGTGAAGGCAGCATTATATGGTCCTGCTTCAGCTAAAATTCCTGCTCCT
CAAGTTAACCTAGCAGCAACTCCTGCTCCTAAAGCATCAGCTCCTGCTCCCCAGCTGGCTGGCACTATGTCAGCAGCATCAACAAATGTTGACATCAGAC
CACCACAAGTGCCTGGAAATGCAGTCACAAACCAGCAGTATTTTCCATCCCAGCAAGGCCAATTTATGAGGCAACCCGGACCCCAACCCCAAGCTATGCC
TCCTATTTCTGCTTCCCATCCACAACAAATTCTTGTAAGTCAAGGTATGCCCAGGGGAGGTACCATGGCTGCTCCTCGGCCTCTGAATTCAAATATCTCA
ACAGATTGGCTTGGTGGAAGTGCTGTTGGCCTGACTTCACAAGCTCCTAGTAGAGGGACTAGTCCTACTACAACTCAGGATGGATTTGGGCTCAGTGCAC
CTGGTTTTACGCCCTCTGTTCAACCTAGACCACAGGTTTCTGCTGGGCAGATGGCAGCCCCTACTTGTAAACCACTAGAAGCAGCCATCACATCCAACCA
ACCTGCAACTAAGGATTTCAAATCAGTGGTTGTTTCTGGGAATGGTTTTGCTTCAGACTCGCATTTTGGCGATGTATTCTCAGCAATTCCTGCTCAAGCA
AAGCAGAGTTCCTTATCAGCTGCACCTTCTACAAGCAGCATACCTGTCTCGTCAGCTATTGTTCCATCTTCTGTCGGATCCCAACATTCACTTAATTCAA
GTTCCCTGGATTCTTTTCAGAGTACTTTTTCTCAGCTGCTTGTGGGTGGTCAATCAACAGCAAGACCAAACCAGCAGGTTCCACCTCAAAGTGTCACCTC
TGCTCCTTCAACTGGTTTTCCTTCTGGATCTAGCAATGCAGCTCTCAGTCAATCCCAGCCTCCATGGCCGAGGATGACTCAATCTGATATTCAAAAGTAC
ACAAAAGTGTTTGTGCAAGTTGACACAGATAGAGATGGGAAGCTTACTGGGGAACAGGCACGCAACCTTTTCTTAAGCTGGAGGCTACCAAGAGAGGTGT
TAAAAAAAGTTTGGGATTTATCCGACCAAGATAATGACAGCATGCTTTCTTTGAGGGAGTTCTGCACTGCTCTCTATTTGATGGAGCGTTATAGAGAAAA
TCGTCCTCTTCCTTCAACACTTCCAACCACTATCATGTCCGATGAGACTCTGTTGTCTGCCACAAGTCACCCTGCAACTTCATATGGCAGTGGAACTTGG
GGACCTGCTTCTGGTTTGCAACAACAACAAGTTGTGACAGTTGCACGGCCATCACCTGCTGCTGCTAGGCCACCAAGGCCACCTGCTGCTCCTCATGCTG
ATGAAAAACATCCCACGCAACAAAAGCCCAATGTTCTTGTGCTTGAAAAACATCTCACGAATCAACTCAACCAAGAGGAGCAGGATGCTCTCAACTCAAA
GTTCCAAGAGGCATCTCAGGCCAATAAAAAGGTAGAAGAACTAGAAAAGGAAATATTAGATTCTAGACAAAAGATTGAGTTTTATCATGTCAAGATGCAG
GAACTTATTTTATATAAAAGTCGATGTGACAATCGACTTAACGAGGTCACAGCAAGGGTATCTACTGATAAGCATGAGGTTGAGACATTAGGAAAGAAGT
ATGAAGAGAAATACAAGCAGACTGGAGATGTTGCCTCCAAACTAACCATTGAAGAGGCTACATTTCATGATATTCAGGAGAAAAAAATGGATTTATATCG
GTCAATTGTTAAAATGGAAGAAGGGGGAGCTGCTGATGGCGTGGTGAAGGAGCATGCTGAGAATATCCAATCGAGTCTTGAGGAACTAGTAAAAACTGTG
AATGAACGCTGTAAACTATATGGATTGCGTTCTAAACCAATCTCACTGGTGGAACTTCCATTTGGCTGGCAACCTGGAATCCAGGAAGCTGCAGCTGATT
GGGATGAAGGTTGGGACAAGTTTGACAACGAAGGATTCACATTTGTCAAAGAGCTCACCTTGGATGTACGAAATGTTGTAGCTTCTCCAAAACAAAAAAC
TTCAGTTCCAAAAGAAACAACTTCCACAGATAAGGATTCAGGTGCAAAGTCAGAAAAGGTTTCAAGGCCAAGCAAATCAAATTCTGAGAAAGATTTGTTG
GATCATCAACATGAGAATGGCACTCTAAAATGTCCTCCTGACAGTCCTGTCAGAAGGAGTACAACAGAGAGCCACCAATCCAGTGAATTCCGAGACTCCC
CATTTAAAGAAAGTGGTGCAGAGAATTCACCCCATGCTAGAGAGATCCAAACTGATGTGGGTGGTACTGAATCTGTGCATTCTGGTGACATAATTGTTGA
AACTGGCTGGGGTACATTTGATGACACACATTATGATACAGAATCAGCCTGGGGCTTCGATTCTGTCAGTGGCAAGGACATGGATTTCAGCATTGGTGAA
TTTGGTTTAAATCCAATAAAAACAGGGTCCTCACATGGAGATAACATGTTCCCAGGGAAGGGCCAGTTTATGTTTGATTCTATTCCAAGCACCCTGGCTC
ATAACCAGGGGAACAGCTCGTATGCTTTTGCAGATTCTGTTCCAAGCACCCCTGCATATAACCCACAGAATGCGTTTGCAGATTCTGTTCCAAGTACCCC
TGCCTATAACACAGGAAAGAGTCCATTTTCATTTGCAGATTCTATCCCAAGCACCCCAGCATACAATTTCGGGAACTCTCCACGAAGGTTTAGTGAAGGT
TCAGAGGACCACCCTTTTGACAGCTTCTCTAGGTTTGATTCATTTAACATGCATGATGGTGGATTATTTCAATCTCCTCGCCATTCTCTTTCAAGATTTG
ATTCCATGCAGAGCACTAAGGACTCGGATCAAAGTTATGGATTCCCATCAAGGTTTGATTCCTTTAGAGAATTTGGAGACTCTAATCGAAGTCATGGGTT
TTCAAGGTTTGACTCATTCAGAGAGTCTGATCAGAATCATGGGTTCTCGAGGTTTGATTCCTTCAAAGAGTCTGATCCTGGTCATGGGTTCTCATCAAGT
TTCAGTTCATTTGGAGAATCTAGAGATACTGATCATAGTCATGGATTCTCAAAGATGGATTCATTTAATGCCCATGACAGTGGGTTCTTCCAAGCATCTG
ACAATTCCGTAGCAAGGTTTGATTCTGTTCGTGGCTCTAAAGACTTTGATAATAGCCATGGGTTCCCATCATTTGATGATACGGATCCATTTGGCTCAAG
TGGACCATTTAGGACATCACTAGAGAGCGAAACTCCGAAGGGAAGTTCTGATAACTGGCGAGCGTTTTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G063200.4 pacid=42765861 polypeptide=Potri.007G063200.4.p locus=Potri.007G063200 ID=Potri.007G063200.4.v4.1 annot-version=v4.1
MTNSNNTDLFDSFFRRADLDGDGQISGAEAVGFFQGSGLPKHVLAQVWMHADQRKAGYLGRQEFYNALKLVTVAQSKRELTPEIVKAALYGPASAKIPAP
QVNLAATPAPKASAPAPQLAGTMSAASTNVDIRPPQVPGNAVTNQQYFPSQQGQFMRQPGPQPQAMPPISASHPQQILVSQGMPRGGTMAAPRPLNSNIS
TDWLGGSAVGLTSQAPSRGTSPTTTQDGFGLSAPGFTPSVQPRPQVSAGQMAAPTCKPLEAAITSNQPATKDFKSVVVSGNGFASDSHFGDVFSAIPAQA
KQSSLSAAPSTSSIPVSSAIVPSSVGSQHSLNSSSLDSFQSTFSQLLVGGQSTARPNQQVPPQSVTSAPSTGFPSGSSNAALSQSQPPWPRMTQSDIQKY
TKVFVQVDTDRDGKLTGEQARNLFLSWRLPREVLKKVWDLSDQDNDSMLSLREFCTALYLMERYRENRPLPSTLPTTIMSDETLLSATSHPATSYGSGTW
GPASGLQQQQVVTVARPSPAAARPPRPPAAPHADEKHPTQQKPNVLVLEKHLTNQLNQEEQDALNSKFQEASQANKKVEELEKEILDSRQKIEFYHVKMQ
ELILYKSRCDNRLNEVTARVSTDKHEVETLGKKYEEKYKQTGDVASKLTIEEATFHDIQEKKMDLYRSIVKMEEGGAADGVVKEHAENIQSSLEELVKTV
NERCKLYGLRSKPISLVELPFGWQPGIQEAAADWDEGWDKFDNEGFTFVKELTLDVRNVVASPKQKTSVPKETTSTDKDSGAKSEKVSRPSKSNSEKDLL
DHQHENGTLKCPPDSPVRRSTTESHQSSEFRDSPFKESGAENSPHAREIQTDVGGTESVHSGDIIVETGWGTFDDTHYDTESAWGFDSVSGKDMDFSIGE
FGLNPIKTGSSHGDNMFPGKGQFMFDSIPSTLAHNQGNSSYAFADSVPSTPAYNPQNAFADSVPSTPAYNTGKSPFSFADSIPSTPAYNFGNSPRRFSEG
SEDHPFDSFSRFDSFNMHDGGLFQSPRHSLSRFDSMQSTKDSDQSYGFPSRFDSFREFGDSNRSHGFSRFDSFRESDQNHGFSRFDSFKESDPGHGFSSS
FSSFGESRDTDHSHGFSKMDSFNAHDSGFFQASDNSVARFDSVRGSKDFDNSHGFPSFDDTDPFGSSGPFRTSLESETPKGSSDNWRAF
|
DESeq2's median of ratios [POPLAR]
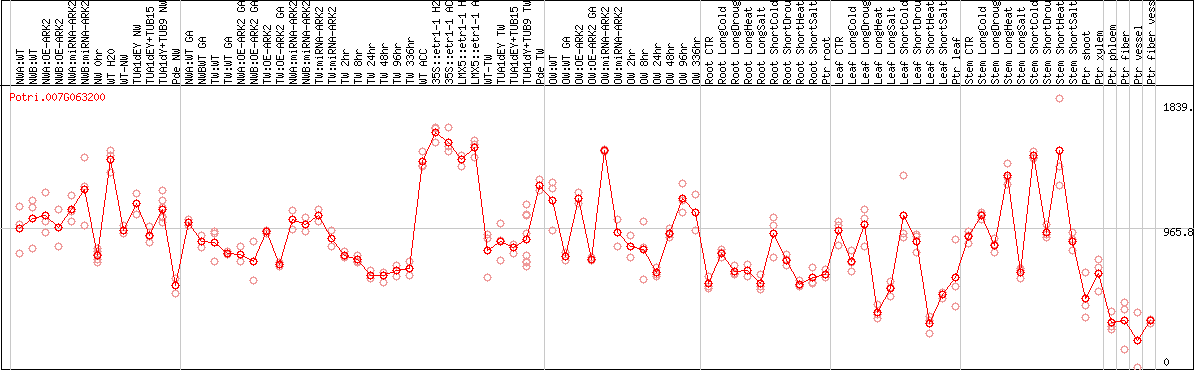
Coexpressed genes
Potri.007G063200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
