External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12910 268 / 2e-89
NAC
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT2G43000 255 / 1e-84
NAC
ANAC042, JUB1, JUNGBRUNNEN1
NAC domain containing protein 42 (.1)
AT1G26870 223 / 3e-70
NAC
FEZ, ANAC009
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G39820 215 / 4e-68
NAC
ANAC094
NAC domain containing protein 94 (.1)
AT2G02450 184 / 4e-55
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT1G61110 177 / 2e-53
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G04070 174 / 3e-52
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G69490 168 / 1e-50
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT5G63790 169 / 2e-50
NAC
ANAC102
NAC domain containing protein 102 (.1)
AT3G15510 168 / 2e-49
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031189
317 / 2e-107
AT3G12910 264 / 9e-87
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10031767
251 / 9e-84
AT2G43000 236 / 3e-78
NAC domain containing protein 42 (.1)
Lus10038332
241 / 6e-79
AT2G43000 231 / 1e-75
NAC domain containing protein 42 (.1)
Lus10030478
239 / 2e-76
AT5G39820 297 / 4e-98
NAC domain containing protein 94 (.1)
Lus10007263
234 / 4e-75
AT5G39820 300 / 3e-100
NAC domain containing protein 94 (.1)
Lus10036194
228 / 6e-74
AT3G12910 224 / 1e-72
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10036749
232 / 4e-73
AT1G26870 306 / 4e-100
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10015367
228 / 7e-73
AT1G26870 298 / 8e-99
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10037178
231 / 1e-72
AT5G39820 306 / 3e-101
NAC domain containing protein 94 (.1)
Lus10023208
179 / 5e-54
AT2G02450 321 / 3e-108
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.007G066300.1 pacid=42765598 polypeptide=Potri.007G066300.1.p locus=Potri.007G066300 ID=Potri.007G066300.1.v4.1 annot-version=v4.1
ATGAACAACTGCCAAGGTCATATTAGCAGTACTAATTATAAAGATGATGGCGTTGAAGAAGCTCAACTTCCAGGGTTTCGATTTCACCCAACAGATGAAG
AGCTTGTTGGGTTCTATCTTCGTCGAAAGGTGGACAAGAAGCCTCTCAATATCGAGCTCATCAAACAAGTTGATATCTACAAATATGATCCATGGGATCT
GCCAAAGCCAAGCAGTGTGGGAGATAATGAAGGATACTTCTTTTGCAAAAGAGGAAGAAAGTATAGGAATAGCATCAGACCTAATAGGGTGACAGGGTCT
GGATTTTGGAAAGCAACTGGCATTGACAAGCCGGTATATTCACTAGGAGGAGAAGGCCGTGACTGCATTGGACTCAAGAAAACACTTGTTTACTATCGCG
GAAGTGCAGGAAAAGGCACCAAAACCGATTGGATGATGCACGAGTTTCGCCTTCCCACCAGTGATAACAACACCAGCTCCACTGCCATTGCCAAAGCTGA
AATATCTCCCCAAGAAGCTGAAGTATGGACACTCTGTAGGATTTTCAAAAGAAATGTGTCATACAGAAAATACACACCGGATTGGAGGCAATTATCAACA
AAATGTCAACAACCACCAATTGACACGAGCTCCAAGATATGCAGTCCAGCCGAGTCTAATTACAAACAAGAAAGCTACGTCAGTTTTGGCGCTCCACTCA
TTCAACATTACGATAATATGCCTCCTGGTAGTAACGCCATCGAAAGGAAACCACTGCATTTGGACCAGTTGAGCCACATAGCTCAGCCACCATCTATGGC
TTCATCCTCAAACATGTCCAGCCCGTATATCCATGAGATGTTTACACATGGGGACTGGGATGAGCTTAGATCAGTCGTGGAGTGTGCTTTTGATCCTTTT
CTTATGTAA
AA sequence
>Potri.007G066300.1 pacid=42765598 polypeptide=Potri.007G066300.1.p locus=Potri.007G066300 ID=Potri.007G066300.1.v4.1 annot-version=v4.1
MNNCQGHISSTNYKDDGVEEAQLPGFRFHPTDEELVGFYLRRKVDKKPLNIELIKQVDIYKYDPWDLPKPSSVGDNEGYFFCKRGRKYRNSIRPNRVTGS
GFWKATGIDKPVYSLGGEGRDCIGLKKTLVYYRGSAGKGTKTDWMMHEFRLPTSDNNTSSTAIAKAEISPQEAEVWTLCRIFKRNVSYRKYTPDWRQLST
KCQQPPIDTSSKICSPAESNYKQESYVSFGAPLIQHYDNMPPGSNAIERKPLHLDQLSHIAQPPSMASSSNMSSPYIHEMFTHGDWDELRSVVECAFDPF
LM
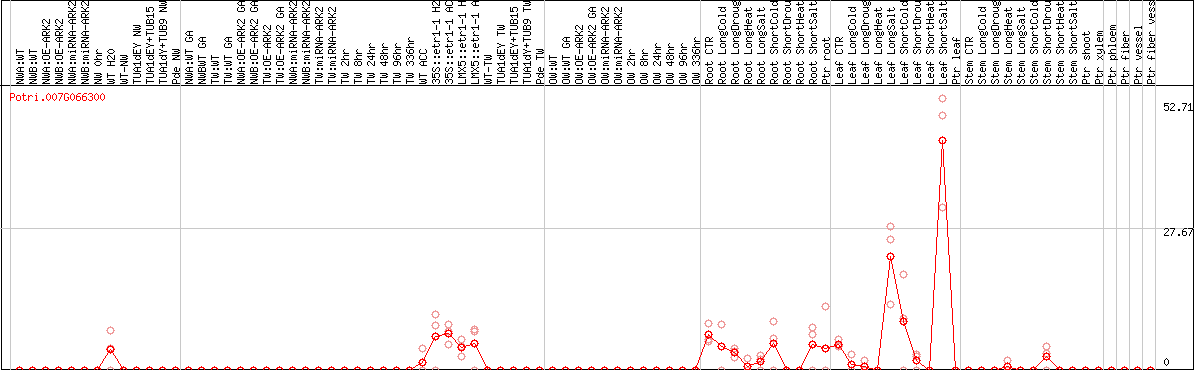
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G066300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.