ATH.2 (Potri.007G076500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ATH.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G076500.1 pacid=42765308 polypeptide=Potri.007G076500.1.p locus=Potri.007G076500 ID=Potri.007G076500.1.v4.1 annot-version=v4.1
ATGAACACCGGGGGTCGACTCATTGCTGGTTCTCACAACAGGAACGAGTTTGTCCTCATAAATGCCGATGAAAATGCAAGAATTAAGTCGGTTCAAGAAT
TGAGCGGACAAGTGTGTCATATATGCGGGGATGAGATTGAGATTACAGTGGATGGAGAGCCATTTGTTGCCTGCAATGAATGTGCTTTCCCTGTGTGCCG
GCCCTGCTATGAATATGAGAGAAGAGAGGGAAATCAAGCTTGCCCTCAATGCAAAACCAGATACAAGCGCCTCAAAGGTAGTCCCAGGGTTGAGGGTGAT
GAGGAAGAAGATGATATTGATGATTTGGAACATGAGTTTGATTATGGAAATTTTGATGGTTTAAGCCCAGAACAAGTTGCAGAGGCAATGCTCTCTTCAC
GCATGAACACTGGCCGTGCTTCTCACTCTAATATATCTGGAATCCCCACACATGGGGAGCTTGATTCCTCTCCCCTTAACTCTAAAATTCCTCTTTTAAC
TTATGGAGAGGAGGATACAGAGATTTCTTCTGACCGACATGCACTTATTGTTCCACCGAGTCATGGAAATAGATTTCATCCGATTTCTTTTCCTGATCCT
TCTATACCTTTAGCTCAACCAAGACCAATGGTTCCAAAGAAAGACATTGCAGTGTATGGGTATGGGAGTGTGGCATGGAAGGACAGAATGGAGGATTGGA
AGAAAAGGCAGAATGACAAACTTCAGGTTGTTAAGCATGAAGGAGGACATGACAATGGAAACTTTGAAGGGGATGAATTGGATGATCCTGATTTGCCCAT
GATGGATGAGGGCAGGCAGCCACTTTCAAGGAAGTTACCCATTCCCTCAAGCAAGATAAACCCGTACAGAATGATAATCATTCTACGTCTTGTAGTTGTT
GGCTTATTTTTTCACTATAGAATTCTCCACCCAGTCAATGATGCGTATGGTTTGTGGTTGACATCAGTAATATGTGAAATATGGTTTGCAGTATCATGGA
TTCTAGACCAGTTTCCAAAATGGTATCCTATAGAACGGGAAACATACCTTGATCGGCTGTCACTGAGGTATGAAAAAGAAGGGAAACCATCTGAGTTAGC
CAGTGTAGATGTTTTCGTAAGTACAGTAGACCCTATGAAAGAACCTCCACTGATCACTGCGAACACTGTCCTCTCTATTCTCGCGGTTGATTATCCTGTG
GATAAAGTTGCATGCTACGTGTCAGATGATGGTGCTGCCATGCTCACATTTGAAGCTCTCTCTGAGACATCTGAATTTGCTAGGAAATGGGTTCCGTTCT
GTAAAAAATTTAATATTGAGCCCCGAGCCCCTGAGTGGTATTTTTCTCAGAAAATGGACTATTTGAAAAATAAAGTTCATCCAGCATTTGTCAGGGAAAG
GCGTGCCATGAAGAGAGAATATGAAGAATTCAAAGTGAAGATAAATGGGCTGGTTGCCACCGCGCAAAAGGTTCCAGAAGATGGTTGGACAATGCAAGAT
GGAACTCCATGGCCTGGAAACAATGTACGAGACCATCCTGGCATGATTCAGGTGTTCCTTGGACAAAGTGGTGTTCGTGACGTGGAAGGAAACGAGTTAC
CTCGTTTGGTCTATGTTTCTCGTGAGAAGAGACCAGGATTTGAACACCATAAAAAGGCTGGGGCCATGAATGCACTGATGCGGGTCACTGCAGTTCTGTC
AAATGCCCCCTATCTTCTGAATGTTGACTGTGACCATTATATTAACAATAGCAGAGCACTTAGAGAAGCAATGTGTTTCTTGATGGATCCAACATCAGGA
AAAAAAGTCTGTTATGTGCAGTTTCCTCAACGATTTGATGGCATTGATCGTCATGATAGATACTCGAATCGGAATGTTGTATTCTTTGATATCAATATGA
AAGGATTAGATGGCTTACAAGGACCAATATATGTTGGAACTGGGTGTGTTTTCCGGAGGCAAGCTCTTTATGGTTATGATGCACCTGTAAAGAAGAGGCC
CCCTGGCAAGACCTGCAACTGCTGGCCTAAATGGTGTTGTCTATTCTGTGGGTCCAGAAAAAACAAGAAATCGAAGCAAAAGAAGGAGAAGAAGAAGTCA
AAGAACAGGGAAGCATCAAAGCAGATACATGCTCTTGAAAATATCGAAGAAGGAATTGAAGAATCAACTTCTGAAAAATCTTCCGAAACATCCCAAATGA
AGTTGGAAAAGAAGTTTGGGCAATCTCCAGTGTTTGTAGCTTCCACTCTTCTGGAGAATGGTGGAGTTCCTCGAGATGCGAGTCCTGCATCACTGTTGAG
AGAGGCCATTCAAGTTATCAGCTGTGGTTATGAAGATAAGACAGAATGGGGCAAGGAAGTTGGCTGGATATATGGTTCTGTAACAGAAGATATCCTGACA
GGATTCAAGATGCACTGCCATGGGTGGCGGTCTGTGTACTGCATACCTAAGCGGCCTGCATTTAAGGGGTCGGCTCCTATTAACCTCTCAGATCGTCTAC
ACCAGGTTCTTCGATGGGCCCTTGGTTCTGTCGAGATTTTCTTTAGTAGACACTGTCCAATTTGGTATGGTTATGGGGGTGGATTGAAGTGGCTGGAACG
ATTTTCTTACATAAACTCTGTTGTGTATCCTTGGACATCCATCCCCTTGCTTGTTTACTGTACACTACCAGCTATATGCCTTCTCACTGGGAAATTCATT
GTTCCTGAGATCAGTAATTATGCCAGTATTGTGTTCATGGCCCTCTTCATATCTATAGCTGCAACTGGTATCCTCGAGATGCAGTGGGGTGGTGTTGGAA
TAGATGACTGGTGGAGAAATGAGCAATTTTGGGTGATTGGAGGTGCTTCGGCACACCTTTTTGCTCTCTTCCAGGGTTTGCTCAAGGTTTTGGCAGGTGT
CAGCACAAACTTCACTGTGACATCAAAAGCAGCGGATGACGGAGAATTCTCAGAGCTTTACCTATTCAAGTGGACATCACTGTTGATCCCTCCTACAACG
TTGTTGATCATGAACATAGTGGGTGTTGTGGTTGGGGTCTCAGACGCCATCAATAATGGGTATGATTCATGGGGTCCATTGTTTGGGAGGCTGTTTTTCG
CCTTGTGGGTCATCATCCACCTCTACCCATTCCTCAAGGGGCTTCTTGGGAAACAAGATCGCATGCCCACCATCATTTTGGTATGGTCAATTCTGCTGGC
TTCAATCCTAACTCTTTTGTGGGTTCGCATAAACCCATTTGTGTCCAAAGGTGGTCCTGTTCTGGAACTGTGCGGATTGAATTGTGATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G076500.1 pacid=42765308 polypeptide=Potri.007G076500.1.p locus=Potri.007G076500 ID=Potri.007G076500.1.v4.1 annot-version=v4.1
MNTGGRLIAGSHNRNEFVLINADENARIKSVQELSGQVCHICGDEIEITVDGEPFVACNECAFPVCRPCYEYERREGNQACPQCKTRYKRLKGSPRVEGD
EEEDDIDDLEHEFDYGNFDGLSPEQVAEAMLSSRMNTGRASHSNISGIPTHGELDSSPLNSKIPLLTYGEEDTEISSDRHALIVPPSHGNRFHPISFPDP
SIPLAQPRPMVPKKDIAVYGYGSVAWKDRMEDWKKRQNDKLQVVKHEGGHDNGNFEGDELDDPDLPMMDEGRQPLSRKLPIPSSKINPYRMIIILRLVVV
GLFFHYRILHPVNDAYGLWLTSVICEIWFAVSWILDQFPKWYPIERETYLDRLSLRYEKEGKPSELASVDVFVSTVDPMKEPPLITANTVLSILAVDYPV
DKVACYVSDDGAAMLTFEALSETSEFARKWVPFCKKFNIEPRAPEWYFSQKMDYLKNKVHPAFVRERRAMKREYEEFKVKINGLVATAQKVPEDGWTMQD
GTPWPGNNVRDHPGMIQVFLGQSGVRDVEGNELPRLVYVSREKRPGFEHHKKAGAMNALMRVTAVLSNAPYLLNVDCDHYINNSRALREAMCFLMDPTSG
KKVCYVQFPQRFDGIDRHDRYSNRNVVFFDINMKGLDGLQGPIYVGTGCVFRRQALYGYDAPVKKRPPGKTCNCWPKWCCLFCGSRKNKKSKQKKEKKKS
KNREASKQIHALENIEEGIEESTSEKSSETSQMKLEKKFGQSPVFVASTLLENGGVPRDASPASLLREAIQVISCGYEDKTEWGKEVGWIYGSVTEDILT
GFKMHCHGWRSVYCIPKRPAFKGSAPINLSDRLHQVLRWALGSVEIFFSRHCPIWYGYGGGLKWLERFSYINSVVYPWTSIPLLVYCTLPAICLLTGKFI
VPEISNYASIVFMALFISIAATGILEMQWGGVGIDDWWRNEQFWVIGGASAHLFALFQGLLKVLAGVSTNFTVTSKAADDGEFSELYLFKWTSLLIPPTT
LLIMNIVGVVVGVSDAINNGYDSWGPLFGRLFFALWVIIHLYPFLKGLLGKQDRMPTIILVWSILLASILTLLWVRINPFVSKGGPVLELCGLNCD
|
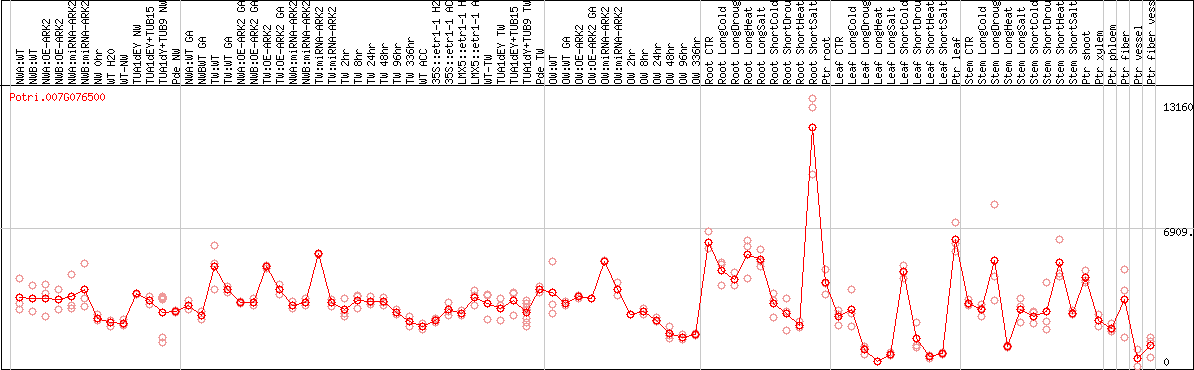
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G076500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
