Potri.007G081900 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G081900.1 pacid=42765211 polypeptide=Potri.007G081900.1.p locus=Potri.007G081900 ID=Potri.007G081900.1.v4.1 annot-version=v4.1
ATGGAAGAAGAGGGAAAACGGGTTGGAGGGCTTTCTTCTGGATTGGCTGTTTTATTGAAAGGCGAAGATAGAAAGGAGGATTCGTGGAAAACCCGACTTG
TTTCATCGTGTGATGATTTTGGTAATCAGCCCGTGGAGCGAGCTCTTGAATATATCTTTGGTCTCTCAAACAAATCACTTGGTCCATTGACTGGTCCAGT
TGACACTAAATTAGTTCGCTCTATTTTAAAAAATGAGTTTTCCAAGTTTTGTATCAAATCTGGTGATTTGGTTGATAGTAGAGACGGGATTCATATCAGT
AAAGATGGTTGTGAATCTCAGGTGGTCGGGCTTGAAGAAGTGAGCATCTGTGGTGATATTAGAATCATTAAACACCCTTTGCATGTAGAAAGCTTAGCAA
TGTTCAGTAGTGCTAGGTCCAATGCTTGTGTTTGGAAAGGGAAATGGATGTATGAAGTTCTATTGGAAACTTGTGGTGTACAGCAGCTTGGATGGGCAAC
TCGTTCTTGTCCTTTTACTGATCACAAGGGTGTAGGTGATGCAGATGATTCATATGCATTCGATGGGAAAAGGGTTAGTAAGTGGAACAAGGATGCTGAG
CCATATGGTCAGCCATGGGTTGTTGGTGATGTCATTGGATGTTGCATTAATCTGGATCATGATGAGATATTGTTCTATAGAAATGGTGTATCCCTTGGAG
TGGCCTTTCGTGGTATCCGCAAGATGGGACCTGGATCTGGGTATTACCCAGCAATCTCTCTTTCTCAAGGTGAACTCTGTGAATTAAATTTTGGGGCTCG
TCCCTTCAAGTACCCCATTCAGGGGTTCCTTCCCCTTAAAGCACCTCCTTCCGCAAATCTTTTGGCTAAGCAGTTGCTTCAATGCTTATCAAGGCTGTCA
GATGTGCAAGGTGCGGGATGGGCTGAAAGTTCTTTAGTTGGTAAATTGAGAAGATTGAAGAGGTTTGTGTCACTTGACGAAGTATTTTATCCTGTCTGTC
AAGGGATATGTGAGGAATTCTTCTCTGTTCTTGAGGGAGATTCTGGAAGTACTGAGTTTGTGGCATGGGGTCCACTTCTTTCATTCATGATGGAGGTTTT
CAGAGTGCAAGCACCACATGACTGTTCAGGTCTGGATAAGTTTATTGATGTATTTCTAGAGTTTCAAGAATCACGTCTGATGTTTGAGCATATCATCAAT
GCCCTTTCAAGTGGCTGCAAAACAGCATCTTTGGTTTTAACAGAATGCCCGTACTCAGGATCTTATTCTTATCTTGCAATGGTATGCCATATCTTACAAC
GGAAAGAATTGATGGTTCTCTGGTGGAAGTCAGCTGATTTTGAATTATTGTTTGAAGGTTTTCTATCGCAAAAGAGTCCAAACAAGCAGGACCTTCAATG
CATGATGCCTTCTGTTTGGTGGCCTGGTTCAGGTGATGATATCTCCAATGATGGAAGAAGCATGATGCTGACAACTACAGCTTTATCGGAAGCTATCAAA
AAGATTGAGGAGAAGCATAGGGACCTCTGTCTCTTGGTCATGCAATTTGTACCGCCTACAACACCTGCTCAATTACCTGGCTCAGTGTTGAGGACATTTT
TACAAAATATTCTATTGAAGAATAGAGGAGCAGATTGCAATGCGCCACCTCCTGGAGTTTCAAGCAATTCCGTTCTTATTTCTTTGTACTCAGTTATACT
TCATTTTTTATCCGAAGGATTTGCTATGAGGGACATTTGTGGCTGGTTAAAGAGGTGTGAACCCAATGGTCTTGATGTTGGATTTCTTCACAGAGGTGGT
GAACAAAGTTTCCCTGTAGATATATTTCTCAAAAATGATCCTCATCGAACTGACATTTCTAGACTTGGGGGATCATTTAGTCATATCTCAAAATCCCATC
CTGCACATGATCAAGAAGCAGAAGTAATCCAGTGGGAGGAAGGTTGTATGGATGATGAAGAAACAAGAGTAACACATAAAACCACACCGAAACCATGTTG
TTGTTCAAGTTATGAGATTGAATTGTCAAAAATCTCCAAGCATCAAATTAGATACAACACTAAAGATTCTCGTGTCCATTGTAGTGGTATTCCAGACAGG
TCTGCTTATGTGGCTGCAGAATGCAGTGAAGGAAGTTTGAATGATGAGATAGCCGACAAACCTAGTACAAGTGATCAATCTGAATCTGATTTTGGTTATT
GTCCAGTGCGAGATATTAGGATTGTGCACAGGGAGAGTGATATGTCTTCAGCAACTCTGAGAGAAGAAGAACTTCTAGATACATTGCTATTGCTGTATCA
CATAGGTGTTGCACCAAAGTTTAAGCAGGCATCGTACTACATGTCTCATCAGGCACAGTCAATATCACTCCTGGAAGAAACTGACAAACAAATAAGAGAA
CGTGCTTGTTGTGAGAAATTAAGACGCCTGAAAGAAGCTCGTAATGAATACCGTGAAGAAGTAATGGATTGTGTAAGACATTGTGCATGGTATCGCATCT
CTCTTTTCTCGCAGTGGAAGCAACGAGGAATGTATGCAACATGCATGTGGATAGTCCAACTACTTCTTGTTCTTAGCAGAGTGGATTCCCTATTTATTTA
TATTCCTGAGTTTTATCTTGAAACCCTGGTTGATTGCTTCCATGTGCTGCGCAAGAGTGACCCCCCATTTGTACCCCCTGCAATATTTATTAAGCAAGGA
CTTGCTTCATTTGTAACTTTTGTTGTTTCCCACTTCAATGATCCGAGGATATTGAGTGCTGATCTCAAGGATCTTCTCCTCCAATCTATTTCAGTCCTGG
TACAGTATAAGGAGTATTTGACAGTGTTTGAGAGCAATGAAGCGGCCACACAAAGAATGCCAAAAGCACTATTGTCGGCATTTGACAACAGATCTTGGAT
CTCAGTGACTAACATTCTTTTGCGGTTGTGCAAGGGTTCTCGTTTCAGTTCTTCAAAGCATGGGGAGTCATCGTCATCATCGTCATTTGTTTTCCAGAAC
TTGTTGCGGGAAGCTTGCATCAATGATGAAGAGTTGTTCTCAGCCTTTCTGAATCGACTATTTAATACACTCAGCTGGACGATGACTGAATTCTCTGTAT
CCATTCGGGAAATGCAAGAAAAATACCAGGTTTTGGAGTTTCAGCAAAGAAAATGTGGTGTCATATTTGATCTCTCATGCAACCTTGCAAAGGTTTTGGA
GTTCTATACTCGTGAAATCCCTCAAGCATTCCTATCTGGAACTGAGACGAACCTTCGGAGGCTAACTGAACTGATAGTCTTTATTCTCAACCACGTTACC
TCAACTGCGGATGCCGAGTTTTTTGATCTGTCACTTAGACGACATGGTCATTCTCCAGAAAAAGTAAATCGAGGCATGATATTAGCACCTCTTGTAGGGA
TCATCTTGAATTTGTTGGATGCAAGGGTGGGGACGGAATGTGGACAGCAGAATGATGTTGTGGGTGTTTTTGCAAGCATGGACTGCCCTGATGCTGTGCA
CTGTGGATTCCAGTATCTCTTAGAGTACAACTGGACTAGGTCCGCAAGAGGGGATGCTTACAGTGGAAAACTTCAGCAGCTAGAGAGTTTCTTGAGCCTC
CTTGTTAGCAGGATTGAGTTGCAACAAATTGAAAGAACGAAACATGAGGAAGAGACAGAAGCTGATGATAATACTTGCTGCATTTGTTATTCTTGTAAAG
CAGATGCAAGGTTTGCCCCTTGTTCACATAGGTCCTGCCATGGCTGCATAACCAGGCATCTTTTAAATTGTCATAGATGTTTCTTTTGCAATGCAACAGT
CTTGGAGGTTATCAAGATTGATGAAAGCAGAGCTTAA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G081900.1 pacid=42765211 polypeptide=Potri.007G081900.1.p locus=Potri.007G081900 ID=Potri.007G081900.1.v4.1 annot-version=v4.1
MEEEGKRVGGLSSGLAVLLKGEDRKEDSWKTRLVSSCDDFGNQPVERALEYIFGLSNKSLGPLTGPVDTKLVRSILKNEFSKFCIKSGDLVDSRDGIHIS
KDGCESQVVGLEEVSICGDIRIIKHPLHVESLAMFSSARSNACVWKGKWMYEVLLETCGVQQLGWATRSCPFTDHKGVGDADDSYAFDGKRVSKWNKDAE
PYGQPWVVGDVIGCCINLDHDEILFYRNGVSLGVAFRGIRKMGPGSGYYPAISLSQGELCELNFGARPFKYPIQGFLPLKAPPSANLLAKQLLQCLSRLS
DVQGAGWAESSLVGKLRRLKRFVSLDEVFYPVCQGICEEFFSVLEGDSGSTEFVAWGPLLSFMMEVFRVQAPHDCSGLDKFIDVFLEFQESRLMFEHIIN
ALSSGCKTASLVLTECPYSGSYSYLAMVCHILQRKELMVLWWKSADFELLFEGFLSQKSPNKQDLQCMMPSVWWPGSGDDISNDGRSMMLTTTALSEAIK
KIEEKHRDLCLLVMQFVPPTTPAQLPGSVLRTFLQNILLKNRGADCNAPPPGVSSNSVLISLYSVILHFLSEGFAMRDICGWLKRCEPNGLDVGFLHRGG
EQSFPVDIFLKNDPHRTDISRLGGSFSHISKSHPAHDQEAEVIQWEEGCMDDEETRVTHKTTPKPCCCSSYEIELSKISKHQIRYNTKDSRVHCSGIPDR
SAYVAAECSEGSLNDEIADKPSTSDQSESDFGYCPVRDIRIVHRESDMSSATLREEELLDTLLLLYHIGVAPKFKQASYYMSHQAQSISLLEETDKQIRE
RACCEKLRRLKEARNEYREEVMDCVRHCAWYRISLFSQWKQRGMYATCMWIVQLLLVLSRVDSLFIYIPEFYLETLVDCFHVLRKSDPPFVPPAIFIKQG
LASFVTFVVSHFNDPRILSADLKDLLLQSISVLVQYKEYLTVFESNEAATQRMPKALLSAFDNRSWISVTNILLRLCKGSRFSSSKHGESSSSSSFVFQN
LLREACINDEELFSAFLNRLFNTLSWTMTEFSVSIREMQEKYQVLEFQQRKCGVIFDLSCNLAKVLEFYTREIPQAFLSGTETNLRRLTELIVFILNHVT
STADAEFFDLSLRRHGHSPEKVNRGMILAPLVGIILNLLDARVGTECGQQNDVVGVFASMDCPDAVHCGFQYLLEYNWTRSARGDAYSGKLQQLESFLSL
LVSRIELQQIERTKHEEETEADDNTCCICYSCKADARFAPCSHRSCHGCITRHLLNCHRCFFCNATVLEVIKIDESRA
|
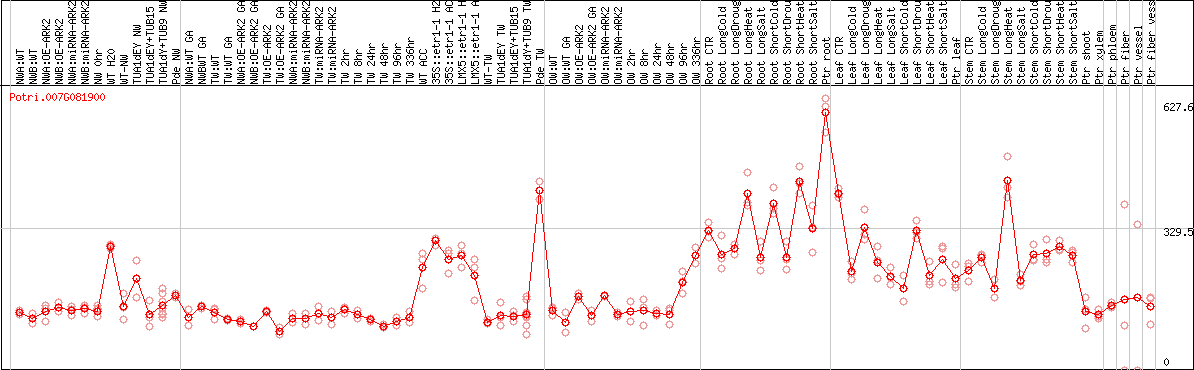
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G081900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
