External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39970 439 / 3e-156
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT3G48420 172 / 2e-51
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT1G56500 82 / 4e-17
haloacid dehalogenase-like hydrolase family protein (.1)
AT5G45170 63 / 5e-11
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT2G38740 61 / 1e-10
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT4G25840 55 / 2e-08
GPP1
glycerol-3-phosphatase 1 (.1)
AT4G21470 54 / 4e-08
ATFMN/FHY
riboflavin kinase/FMN hydrolase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G088500
169 / 2e-50
AT3G48420 434 / 5e-154
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.013G007800
73 / 5e-14
AT1G56500 1489 / 0.0
haloacid dehalogenase-like hydrolase family protein (.1)
Potri.003G086900
67 / 1e-12
AT2G38740 359 / 6e-127
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.001G147300
63 / 3e-11
AT2G38740 360 / 5e-127
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.014G043500
59 / 6e-10
AT2G38740 297 / 1e-101
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.002G077200
56 / 1e-08
AT4G21470 561 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Potri.001G147400
52 / 1e-07
AT2G38740 350 / 2e-123
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.005G183400
51 / 3e-07
AT4G21470 575 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Potri.001G104366
47 / 1e-05
AT4G11570 494 / 9e-176
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000633
456 / 3e-163
AT4G39970 427 / 1e-151
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10023149
453 / 5e-162
AT4G39970 423 / 4e-150
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10013663
173 / 1e-51
AT3G48420 427 / 4e-151
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10033034
171 / 4e-51
AT3G48420 424 / 4e-150
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10020699
72 / 1e-13
AT1G56500 1467 / 0.0
haloacid dehalogenase-like hydrolase family protein (.1)
Lus10029840
71 / 3e-13
AT1G56500 1528 / 0.0
haloacid dehalogenase-like hydrolase family protein (.1)
Lus10003018
61 / 3e-10
AT2G38740 355 / 1e-124
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10018435
59 / 1e-09
AT4G21470 553 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Lus10018906
57 / 7e-09
AT4G21470 556 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Lus10016820
50 / 4e-08
AT3G48420 105 / 4e-29
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF13419
HAD_2
Haloacid dehalogenase-like hydrolase
Representative CDS sequence
>Potri.007G095200.11 pacid=42765743 polypeptide=Potri.007G095200.11.p locus=Potri.007G095200 ID=Potri.007G095200.11.v4.1 annot-version=v4.1
ATGGCTTCTTCTCTGCTAATCTCTCACTCCAACTTCCTCTCTTTCTCTCAATCCCCTACTCTCTTCCCTTTCCACCTCAAAACTACCGCTATAAAACAAC
GACAGCGACAACATTACTCTCCTCCTTCTCGTCACCTCAACATCGTCTCTTCTTCTGCCGCTTCAGCTTCTCTTGAAGCTCTAATATTTGACTGCGATGG
AGTTATTTTAGAATCTGAACACCTCCATCGCCAAGCTTATAATGACGCTTTTGCTCACTTCAATGTTATCTGCTCTTCTTCTCTCCCTCTCAATTGGTCT
CCCGATTTCTATGATGTGCTCCAGAATCGTATCGGTGGTGGCAAACCTAAGATGCGATGGTATTTTAAGGAGCATGGATGGCCGTCTTCCAACCTATTTG
AGAAGCCACCGGAGGATGATGAGAGCCGTGCTAAGCTGATTGATACTCTTCAGGATTGGAAAACTGAAAGGTACAAAGAAATAATCAAATCTGGAACTGT
GGAGCCCAGGCCTGGGGTTTTGAGGTTGATGGATGAGGCTAAGGCTGCTGGAAAAAAGCTGGCTGTTTGCTCTGCAGCAACTAAAAGTTCAGTTATACTC
TGTCTTGAGAACCTTATAGGAATGGAGCGGTTTCAAGGCCTTGATTGCTTTCTTGCAGGTGATGATGTCAAGGAAAAGAAGCCAGATCCATCAATTTATG
TGACAGCTTCCAAGATGCTTGGTGTGTCGGAGAGGGATTGTTTGGTGGTGGAGGATAGTGTCATTGGTCTACAGGCGGCAACAACAGCTGGAATGTCATG
TGTTATCACTTACACCCCTTCAACAGCTGATCAGGATTTCAAAGATGCTATAGCAATTTACCCTGATTTGAGTAATGTGAGATTAAAAGACTTAGAGTTG
TTGCTTCAAAATGTTGTTGCTGCCAGCTAG
AA sequence
>Potri.007G095200.11 pacid=42765743 polypeptide=Potri.007G095200.11.p locus=Potri.007G095200 ID=Potri.007G095200.11.v4.1 annot-version=v4.1
MASSLLISHSNFLSFSQSPTLFPFHLKTTAIKQRQRQHYSPPSRHLNIVSSSAASASLEALIFDCDGVILESEHLHRQAYNDAFAHFNVICSSSLPLNWS
PDFYDVLQNRIGGGKPKMRWYFKEHGWPSSNLFEKPPEDDESRAKLIDTLQDWKTERYKEIIKSGTVEPRPGVLRLMDEAKAAGKKLAVCSAATKSSVIL
CLENLIGMERFQGLDCFLAGDDVKEKKPDPSIYVTASKMLGVSERDCLVVEDSVIGLQAATTAGMSCVITYTPSTADQDFKDAIAIYPDLSNVRLKDLEL
LLQNVVAAS
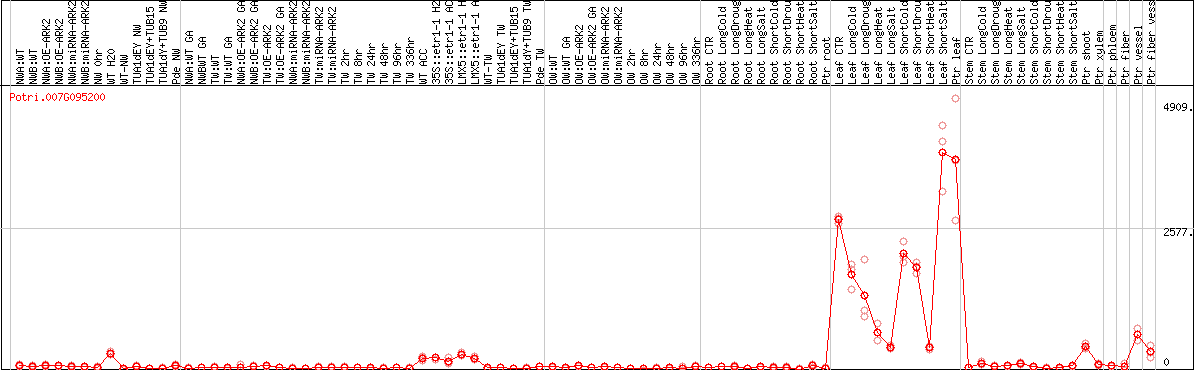
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G095200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.