CHR932 (Potri.007G102800) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CHR932 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.007G102800.8 pacid=42765856 polypeptide=Potri.007G102800.8.p locus=Potri.007G102800 ID=Potri.007G102800.8.v4.1 annot-version=v4.1
ATGGCGGCTCTTTCACAAAGCACACACAGAAAACCAATTAGTCTGAACGATCGCCATTATCGCCTCCTTCAAGACGTCTCTGCTCCTCCCAAACAACCAC
CGCCTCCGCCGCCAGTAACGTCCTTCGAGGAAGAAGAAGAGAGCGTATTCAACGTGAAGTTCGACGGACGACGTCGTATTTGCAAAGCAGAACCAGAGGA
TGATAATATCCCTAAATTCTGTGGAATCACTGATTTTGATTCTTCACCTGAAGAAGAAAAGCCTACGAAGGTTAGGATTGAAGGAAGACGGAGACTTTGC
AAAGTATCTTCTGGAGATGATGGGGATGATGCAAGTAGAGAAGAAGTGAAAGATGATTCTAGTTTTGATGATATTGCCGATTTTGATTCACCCATTCCGT
CTAAAAGTGTTGGTGATTGTGGTAATAATAAGGGTGTTAATGAGATAAAAGATATTCTTAGTGATTTGACCTCAAGGCTTGACCTTTTGTCAATTGAGAA
AAAGAGAGTTCCTGAAAATAATAATGTGGTCAAGAAAGTTCATGTGGTTGAATATGCAAGTGCTGAGTCTTTGTTTTCTTTGTCGTCTAGTCCATCTGAT
TCCTCATTGGATGTTATTAAAAATGGTGGAGGTGATGATGAGAGTGCTGTGGATGAGTATGAGGAGGGTGATTTATTGAGTGAGTCATTTGATGATGAGG
TTAGCAGAGGGCTAAAGAAGAACGAATATGGAAGGGTGGATGAAAAGTTAGTCCCTGTGGGAAAACCTTTTGTGTCCAATGTTGTAGAAGATGAAAGTGA
TGTGCAGATTGAGTCTAACCATGATGAATATGTAACTAGGGTGGAGAAAACTAAGAACGTTACTCAAAGAGTGAAGGAAAATGAACCCGATGGGTTCAAT
GAGAGATTGAGGTCTGTAGGACGGTCTTCGGTGCTTAGTCTTCGAGATGAATCAGAGGATGATGAAGATGATTGTGTGGTTTTGACCGGAAAAAAGGTGG
TCAAGAAAGTTGGGAGACCCGGTGCCATTGCCAAGTATAATGTATTATCTGGTGAATCTGAGACTGCTGTGTTGGAAAATCATGCAGAGTCAGAGGATGA
TGGTTCCATTATCTTGCCTGGACTAAAATCTACTTACAAGTTACCTGGTAAAATTGCAAAAATGCTGTATCCACATCAGTGTGAAGGTTTAAGGTGGCTC
TGGTCTTTGCATTGCAAGGGTAAGGGTGGAATCTTAGGAGATGACATGGGCTTGGGAAAAACGATGCAGATTTGTAGCTTTCTAGCAGGTCTCTTCCATT
CGAAATTGATTAAGAGGGTATTGGTTGTGGCCCCTAAAACACTGCTCACTCATTGGATTAAAGAACTATCTGTTGTGGGGCTTTCTGGGAAGACTAGAGA
ATACTTTGGGACCTCTTTAAAAGCTCGGGACTATGAGCTGCAATATATACTTCAGGACAAAGGAATTCTTCTCACAACTTATGATATTGTGCGGAACAAC
TCAAAATCTTTACGAGGGGACCATTACTTTCTTGACGAGGAAAGTGAGGATAGTTATATTTGGGACTATATGATTCTAGATGAGGGACATCTCATAAAAA
ATCCTAGTACTCAGAGAGCGAAAAGTTTGATTGAGATACCAAGTGCTCATTGTATTGTCATTAGTGGCACTCCGATTCAAAACAATCTTAAGGAATTGTG
GGCTTTGTTCAACTTCTGTTGTCCTGACCTTCTTGGTGACAACAAGTGGTTTAAGCAAACATACGAGCATCCCATACTTCGTGGAAATGAAAAAAATGCT
TCTGATAGGGAGAAGCGTATTGGTTCAACAGTTGCAATGGAACTAAGAGAACGCATTCAACCTTACTTTTTGCGCCGCATGAAGAATGAAGTATTCAAGG
AAGATGATGCCACAACTGCTAAACTTTCCCGAAAGAATGAGATCATTGTGTGGCTTAGACTAACTACTTGCCAGCGGCAACTTTATGAAGCCTTTTTGCG
GAGCGAGATTGTTCTCTCAGCTTTTGATGGCTCGCCATTGGCTGCACTAACGATATTGAAGAAAATATGTGATCACCCGCTTCTCTTGACAAAAAGAGCG
GCTGAAGATTTGTTAGAGGGAATGGAGTCCATGCTAAATCCTGAAGATGTTGCTGTGGCTGAAAAATTGGCCATGCATGTTGCTGATGTTGCTGAAAGAA
CTGACTTTCAAGAGAAGCATGACAGCATCTCCTGCAAGATCTCTTTTGTGTTGTCACTACTGGATAATTTGATTCCAGAGGGACATAATGTTCTTATATT
CTCTCAAACTCGTAAGATGCTTAATCTCATCGAGGAATCTCTGGTGTCCAATGGTTATGAGTTCTTACGCATAGATGGTACGACAAAAGTTACTGATAGA
GCAAAAATTGTTGATGACTTCCAAGAAGGAAATGGTGCTCCTATATTTTTGCTAACATCTCAGGTTGGCGGTCTTGGTCTTACACTTACAAAAGCAGATC
GTGTGATTGTGGTTGATCCTGCTTGGAATCCTAGCACGGATAACCAAAGTGTTGATCGTGCATATCGAATTGGGCAAAAGAAGGATGTGGTCGTATACAG
ATTAATGACTTGTGGAACTGTTGAAGAGAAAATTTATAGAAAGCAGATATTCAAGGGGGGGTTATTTAGAACAGCAACTGAAAACAAAGAACAAATTCGA
TACTTTAGCCAACAGGATCTGCGGGAGCTATTCAGTCTCCCTAAGCAGGGGTTCAATATTTCCCTCACACAGCAACAATTACATGAAGAGCATGACAGCC
AGCACAAAATGGATGAGTATCTGGAATCCCACATAAAATTCTTGGAGAGTCAAGGTATAGCAGGAGTCAGTCACCACAGCTTACTCTTCTCAAAGACAGA
AACTGTACAACTTGCACAGGAAGAAGAGGATGAGATAAGGAAGAAAGTATCCACAATGGTGGGAAATTCATCGTCAAGCTATTCACTTGAACGCAATGTT
GACGGGGCTGCTCGTGCTTTCAATCCCAAGGATGTAAATCTGAACAAGAAAACCTCTTCTCCAGATAGTGTGGGCAAACTGACAGAATCTGAAATCTTAG
AGAGAATCAATCGGCTGTCACAATTACTTGGAAATAAGGTTACCGTTTTAAGGTTACCAGACCAAGGAGCGAAACTACAGAAGCAGATTAGTGAATTAAA
TTCAGTGCTCATCGAGCTGAGGATGGAGAAAGCAACAGAGAGAGAAGGAGTCATCAGTCTGGATGATTTAACTGGGGAATTCGAAAGAGGATTGAATGTA
TAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.007G102800.8 pacid=42765856 polypeptide=Potri.007G102800.8.p locus=Potri.007G102800 ID=Potri.007G102800.8.v4.1 annot-version=v4.1
MAALSQSTHRKPISLNDRHYRLLQDVSAPPKQPPPPPPVTSFEEEEESVFNVKFDGRRRICKAEPEDDNIPKFCGITDFDSSPEEEKPTKVRIEGRRRLC
KVSSGDDGDDASREEVKDDSSFDDIADFDSPIPSKSVGDCGNNKGVNEIKDILSDLTSRLDLLSIEKKRVPENNNVVKKVHVVEYASAESLFSLSSSPSD
SSLDVIKNGGGDDESAVDEYEEGDLLSESFDDEVSRGLKKNEYGRVDEKLVPVGKPFVSNVVEDESDVQIESNHDEYVTRVEKTKNVTQRVKENEPDGFN
ERLRSVGRSSVLSLRDESEDDEDDCVVLTGKKVVKKVGRPGAIAKYNVLSGESETAVLENHAESEDDGSIILPGLKSTYKLPGKIAKMLYPHQCEGLRWL
WSLHCKGKGGILGDDMGLGKTMQICSFLAGLFHSKLIKRVLVVAPKTLLTHWIKELSVVGLSGKTREYFGTSLKARDYELQYILQDKGILLTTYDIVRNN
SKSLRGDHYFLDEESEDSYIWDYMILDEGHLIKNPSTQRAKSLIEIPSAHCIVISGTPIQNNLKELWALFNFCCPDLLGDNKWFKQTYEHPILRGNEKNA
SDREKRIGSTVAMELRERIQPYFLRRMKNEVFKEDDATTAKLSRKNEIIVWLRLTTCQRQLYEAFLRSEIVLSAFDGSPLAALTILKKICDHPLLLTKRA
AEDLLEGMESMLNPEDVAVAEKLAMHVADVAERTDFQEKHDSISCKISFVLSLLDNLIPEGHNVLIFSQTRKMLNLIEESLVSNGYEFLRIDGTTKVTDR
AKIVDDFQEGNGAPIFLLTSQVGGLGLTLTKADRVIVVDPAWNPSTDNQSVDRAYRIGQKKDVVVYRLMTCGTVEEKIYRKQIFKGGLFRTATENKEQIR
YFSQQDLRELFSLPKQGFNISLTQQQLHEEHDSQHKMDEYLESHIKFLESQGIAGVSHHSLLFSKTETVQLAQEEEDEIRKKVSTMVGNSSSSYSLERNV
DGAARAFNPKDVNLNKKTSSPDSVGKLTESEILERINRLSQLLGNKVTVLRLPDQGAKLQKQISELNSVLIELRMEKATEREGVISLDDLTGEFERGLNV
|
DESeq2's median of ratios [POPLAR]
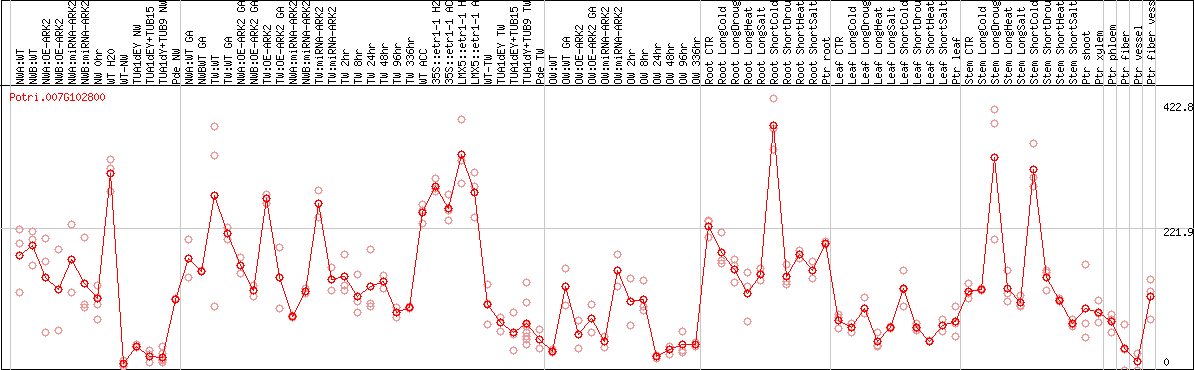
Coexpressed genes
Potri.007G102800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
