External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G77980 98 / 7e-26
MADS
AGL66
AGAMOUS-like 66 (.1)
AT1G22130 94 / 2e-24
MADS
AGL104
AGAMOUS-like 104 (.1)
AT1G77950 90 / 2e-23
MADS
AGL67
AGAMOUS-like 67 (.1.2)
AT2G14210 83 / 6e-21
MADS
AGL44, ANR1
ARABIDOPSIS NITRATE REGULATED 1, AGAMOUS-like 44 (.1)
AT1G71692 82 / 1e-20
MADS
XAL1, AGL12
XAANTAL1, AGAMOUS-like 12 (.1)
AT2G45660 82 / 2e-20
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT5G10140 80 / 3e-20
MADS
FLF, AGL25, FLC
FLOWERING LOCUS F, FLOWERING LOCUS C, AGAMOUS-like 25, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
AT4G24540 81 / 4e-20
MADS
AGL24
AGAMOUS-like 24 (.1)
AT3G57230 81 / 5e-20
MADS
AGL16
AGAMOUS-like 16 (.1.2)
AT4G11880 80 / 7e-20
MADS
AGL14
AGAMOUS-like 14 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G087500
94 / 3e-24
AT1G22130 229 / 9e-73
AGAMOUS-like 104 (.1)
Potri.002G092300
94 / 3e-24
AT1G22130 313 / 1e-105
AGAMOUS-like 104 (.1)
Potri.001G058200
89 / 3e-23
AT2G03710 156 / 3e-48
SEPALLATA 4, AGAMOUS-like 3, K-box region and MADS-box transcription factor family protein (.1.2.3)
Potri.012G100200
86 / 5e-22
AT1G26310 149 / 2e-44
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
Potri.019G077200
81 / 3e-20
AT4G09960 274 / 5e-94
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Potri.002G079000
80 / 6e-20
AT5G20240 238 / 2e-80
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Potri.005G155300
81 / 7e-20
AT2G22540 234 / 8e-78
SHORT VEGETATIVE PHASE, AGAMOUS-like 22, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.005G182200
80 / 8e-20
AT5G20240 244 / 2e-82
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Potri.013G102600
80 / 8e-20
AT1G71692 250 / 4e-85
XAANTAL1, AGAMOUS-like 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Potri.007G120002.1 pacid=42764864 polypeptide=Potri.007G120002.1.p locus=Potri.007G120002 ID=Potri.007G120002.1.v4.1 annot-version=v4.1
ATGGGTCGTAACAAGCTTCCGATGAAGAAAATTGATAATCCCTCCCGTCGTAAAATAACATACTCAAAACGTAGAGATGGGATTATCAAGAAAGCCACTG
AATTGTCTGTGCTATGTGATACAGATGTTGGTCTTTTGATGTTTTCTCCTAATGGTAGATTGACTACCTTTGCTAGCAATGGAAGGATTGAAGACGTCTT
TCTTCGCTATTTTGACCAGTCCAATGTACTAGAAGCAGGGCGCTTCAGGAGCAACTCGATGAAATCAGCCAGAAACTACAAGAGAGACAAGAAAAGATGA
AA sequence
>Potri.007G120002.1 pacid=42764864 polypeptide=Potri.007G120002.1.p locus=Potri.007G120002 ID=Potri.007G120002.1.v4.1 annot-version=v4.1
MGRNKLPMKKIDNPSRRKITYSKRRDGIIKKATELSVLCDTDVGLLMFSPNGRLTTFASNGRIEDVFLRYFDQSNVLEAGRFRSNSMKSARNYKRDKKR
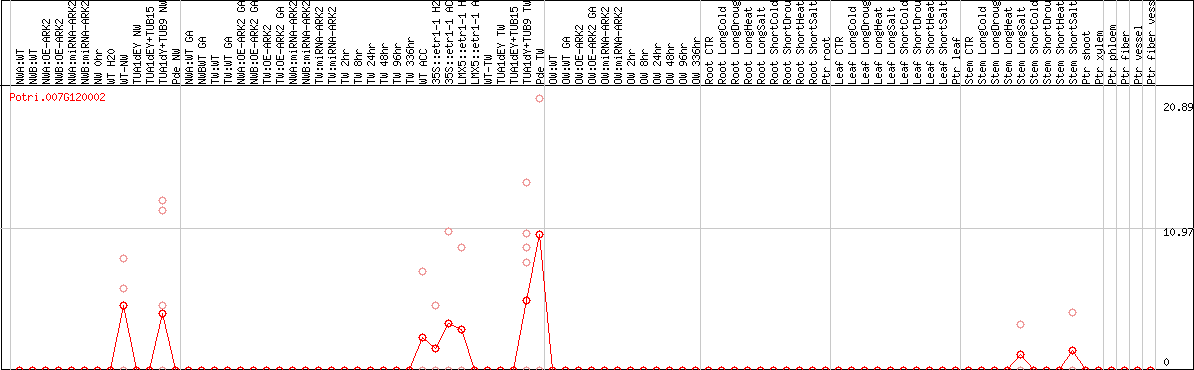
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G120002 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.