External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G15248 108 / 1e-31
CO
B-box type zinc finger family protein (.1)
AT3G21890 103 / 2e-29
CO
B-box type zinc finger family protein (.1)
AT3G21150 70 / 2e-15
CO
EIP6, BBX32
EMF1-Interacting Protein 1, B-box domain protein 32, B-box 32 (.1)
AT4G27310 67 / 3e-14
CO
B-box type zinc finger family protein (.1)
AT5G54470 64 / 3e-13
CO
B-box type zinc finger family protein (.1)
AT2G33500 60 / 2e-11
CO
COL14
B-box type zinc finger protein with CCT domain (.1.2)
AT1G28050 57 / 2e-10
CO
COL15
B-box type zinc finger protein with CCT domain (.1)
AT1G73870 56 / 5e-10
CO
COL7
B-box type zinc finger protein with CCT domain (.1)
AT5G15840 55 / 1e-09
CO
FG, CO
CONSTANS, B-box type zinc finger protein with CCT domain (.1.2)
AT3G02380 54 / 3e-09
CO
ATCOL2, COL2
CONSTANS-like 2 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039694
142 / 9e-45
AT4G15248 108 / 1e-31
B-box type zinc finger family protein (.1)
Lus10027151
142 / 1e-44
AT4G15248 109 / 7e-32
B-box type zinc finger family protein (.1)
Lus10015097
121 / 1e-36
AT4G15248 94 / 5e-26
B-box type zinc finger family protein (.1)
Lus10031583
121 / 1e-36
AT3G21890 96 / 1e-26
B-box type zinc finger family protein (.1)
Lus10031593
67 / 2e-14
AT4G27310 119 / 2e-33
B-box type zinc finger family protein (.1)
Lus10033751
66 / 5e-14
AT4G27310 114 / 9e-32
B-box type zinc finger family protein (.1)
Lus10033380
65 / 2e-13
AT3G21150 112 / 3e-30
EMF1-Interacting Protein 1, B-box domain protein 32, B-box 32 (.1)
Lus10034830
64 / 3e-13
AT3G21150 112 / 3e-30
EMF1-Interacting Protein 1, B-box domain protein 32, B-box 32 (.1)
Lus10018513
61 / 8e-12
AT4G27310 117 / 3e-32
B-box type zinc finger family protein (.1)
Lus10039727
60 / 9e-12
AT5G54470 118 / 1e-32
B-box type zinc finger family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00643
zf-B_box
B-box zinc finger
Representative CDS sequence
>Potri.007G121100.1 pacid=42766507 polypeptide=Potri.007G121100.1.p locus=Potri.007G121100 ID=Potri.007G121100.1.v4.1 annot-version=v4.1
ATGTGTAGGGGTATTCATCAGGAGAGAAATAACCAAGGCGGTTCTTGTGGCGAAGAGGTGGTTTCATCTAAGGCAACCTCAAGATTGGTTTGTTGTGAGC
TGTGTGGATCAAGGGCATCATTATATTGTCAAGCAGATGATGCATTTTTATGTCAAAAATGTGACAAATGGGTGCATGGAGCAAATTTCCTAGCTCAAAG
GCATGTTAGATGCATGCTATGCAACACATGCCAAAATCTTACACAAAGATATCTCATAGGGGCTTCAACTGAGTTGTTGCTTCCAACTATTGTGAGTTGG
AGAGAAAGAAGGCAGTGCAATTCTAATCTTGAAAAGAAGAGCTCTGGATCACTTAAAATGCCCTTTTTGTTTCTTTGA
AA sequence
>Potri.007G121100.1 pacid=42766507 polypeptide=Potri.007G121100.1.p locus=Potri.007G121100 ID=Potri.007G121100.1.v4.1 annot-version=v4.1
MCRGIHQERNNQGGSCGEEVVSSKATSRLVCCELCGSRASLYCQADDAFLCQKCDKWVHGANFLAQRHVRCMLCNTCQNLTQRYLIGASTELLLPTIVSW
RERRQCNSNLEKKSSGSLKMPFLFL
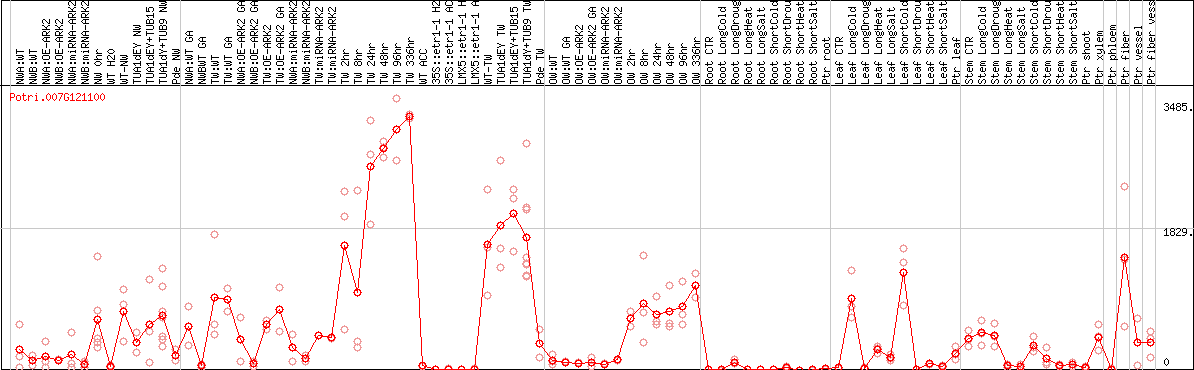
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G121100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.