External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G53970 225 / 9e-73
TAT7
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
AT5G36160 224 / 3e-72
Tyrosine transaminase family protein (.1)
AT4G28410 206 / 3e-65
Tyrosine transaminase family protein (.1)
AT2G20610 206 / 5e-65
RTY1, RTY, HLS3, ALF1, SUR1
SUPERROOT 1, ROOTY 1, ROOTY, HOOKLESS 3, ABERRANT LATERAL ROOT FORMATION 1, Tyrosine transaminase family protein (.1.2)
AT4G28420 201 / 7e-63
Tyrosine transaminase family protein (.1.2)
AT4G23600 187 / 5e-59
JR2, CORI3, TAT1
JASMONIC ACID RESPONSIVE 2, CORONATINE INDUCED 1, Tyrosine transaminase family protein (.1.2.3)
AT2G24850 180 / 5e-55
TAT3
tyrosine aminotransferase 3 (.1)
AT4G23590 178 / 1e-54
Tyrosine transaminase family protein (.1)
AT1G80360 42 / 0.0001
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033659
251 / 1e-82
AT5G36160 481 / 3e-169
Tyrosine transaminase family protein (.1)
Lus10017703
250 / 3e-82
AT2G20610 486 / 1e-170
SUPERROOT 1, ROOTY 1, ROOTY, HOOKLESS 3, ABERRANT LATERAL ROOT FORMATION 1, Tyrosine transaminase family protein (.1.2)
Lus10018626
247 / 3e-81
AT2G20610 517 / 0.0
SUPERROOT 1, ROOTY 1, ROOTY, HOOKLESS 3, ABERRANT LATERAL ROOT FORMATION 1, Tyrosine transaminase family protein (.1.2)
Lus10017934
246 / 4e-81
AT5G53970 619 / 0.0
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
Lus10039861
246 / 1e-80
AT2G20610 509 / 1e-179
SUPERROOT 1, ROOTY 1, ROOTY, HOOKLESS 3, ABERRANT LATERAL ROOT FORMATION 1, Tyrosine transaminase family protein (.1.2)
Lus10017705
243 / 2e-79
AT5G53970 507 / 1e-179
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
Lus10033661
239 / 6e-78
AT5G36160 499 / 1e-176
Tyrosine transaminase family protein (.1)
Lus10017704
235 / 1e-76
AT5G53970 508 / 4e-180
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
Lus10013676
234 / 3e-73
AT5G53970 617 / 0.0
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
Lus10033660
194 / 9e-62
AT5G53970 408 / 3e-142
tyrosine aminotransferase 7, Tyrosine transaminase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0061
PLP_aminotran
PF00155
Aminotran_1_2
Aminotransferase class I and II
Representative CDS sequence
>Potri.007G138002.1 pacid=42766465 polypeptide=Potri.007G138002.1.p locus=Potri.007G138002 ID=Potri.007G138002.1.v4.1 annot-version=v4.1
ATGGGTGTTTTTGGATCAACTGTGCCTGTTATTACCCTTGGATCCATATCGAAAAGATGGATGGTTCCCGGTTGGCGACTTGGATGGCTAGTAACAAGTG
ATCCTACTGGTTTACTTCAAAAATGTGGGATCGCTGATTCCATTAAGAATGTTCTCAATCCTGCCCCTTTCTCGCCGACCTTTATTCAGGCCGCTGTTCC
TGAAATCCTTGAGAAAACAACAGAGGAGTTCTTTTCAAAAACTATAAACATTCTTAGAGCAGCTTCAGCCTTCTGTTACGATAAACTGAAGGAGATACCG
TGCATTACTTGCCCACAAAGGGCAGAAGGAGCCATGTTTGTTTTGGTGAAGCTTAACCTTTCACTTCTTGAAGACATCAAAGATGACATGGAGTTCTGTC
TGAAGCTAGCTAAAGAGGAATCTCTCGTCATTTTACCAGGGGTGACTGTGGGACTAAAGAATTGGCTCCGCATAACATTTTCAGTTGAACAATCATCTCT
TGAAGATGGTCTTGGAAGGTTAAGGTACTTCTGTGGGAGACATGCGAAGAAACCATAG
AA sequence
>Potri.007G138002.1 pacid=42766465 polypeptide=Potri.007G138002.1.p locus=Potri.007G138002 ID=Potri.007G138002.1.v4.1 annot-version=v4.1
MGVFGSTVPVITLGSISKRWMVPGWRLGWLVTSDPTGLLQKCGIADSIKNVLNPAPFSPTFIQAAVPEILEKTTEEFFSKTINILRAASAFCYDKLKEIP
CITCPQRAEGAMFVLVKLNLSLLEDIKDDMEFCLKLAKEESLVILPGVTVGLKNWLRITFSVEQSSLEDGLGRLRYFCGRHAKKP
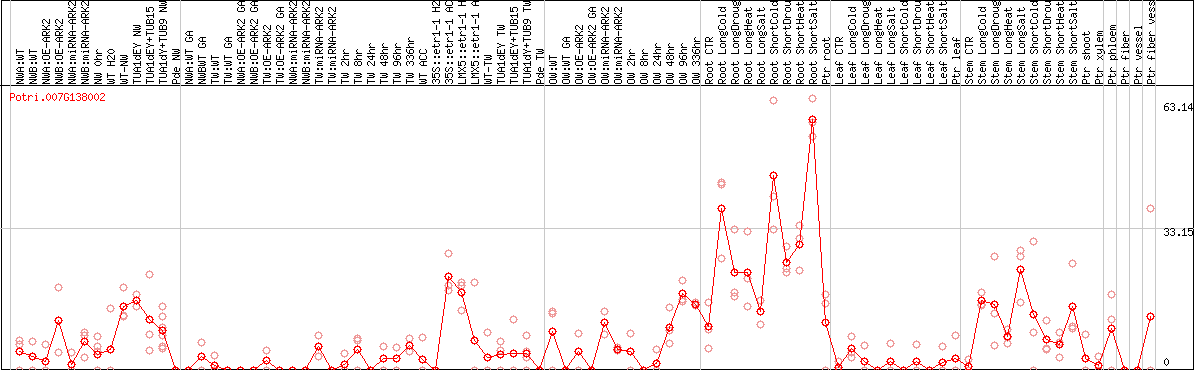
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.007G138002 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.