Potri.008G009100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G009100.1 pacid=42806115 polypeptide=Potri.008G009100.1.p locus=Potri.008G009100 ID=Potri.008G009100.1.v4.1 annot-version=v4.1
ATGGCTTCAGCGAAGACGAGTAGTCGTTCGCGAGGGACGCCGGTGAAGGAGAACGGCACGAAGCTCGAGGAAGGACTCAACGTCTTCAAGTCTGATAGAT
TCAACGCCGATTCTTACGTTCAGTCCAAATGCTCTCTAAACGAGAAGGAAATAAAGCAGTTATGCTCGTATCTTTTGGATTTAAAGAGGGCTTCAGCTGA
CGAAATGCGGAAAAGTGTTTACGCTAATTATGCTGCGTTTATAAGGACGTCGAAGGAGATATCAGATTTGGAAGGGGAGTTGTTGTCTATAAGAAATTTG
TTATCTACTCAAGCAACTTTGATTCATGGTTTAGTAGAGGGAGTTAACATTGATTCTTTATCTTTGAAAGCATCTGAAGGGTCTTTGGTAAATGGGTTAG
AGAATGTTGAAGATAGAGAGCCGACTGATTTGGAGAGATGGTTGGCTGAGTTCCCTGATATGCTTGATGTTTTGTTAGCTGAGAGGAGAGTGGATGAAGC
TCTGGCGGTGATTGACGAGGGAGAACGTATAGCTGCTGAAATGAAAAAAACAGAATTGTCAAGTCCTGGTATACTTAGGTCTTTAGAGATAGCGATTACA
GAGCGCGGGCAAAAGTTGGCTGATCAGCTTGCTGAAGCTGCTTGCCAACCTTCTACTCGCAGTAGTGAACTTCGTGCAGCCATATCAGCTCTTAAAAAGC
TCGGGGATGGCCCTCGAGCTCATAGCTTGCTCCTCAATGCACACCTCCAAAGATATCAGTATAATATGCAAAGCCTCTGCCCGTCTAGCACATCATATGG
AGGAGCTTATACGGCTGCACTGTCACAGATAGTGTTTTCTGCAATCGTTCAAGCTTCTAGTGATTCTTTGGCTATTTTCGGTAAGGAACGGGAGTATAGG
TCTGAGCTTGTGATGTGGGCCACCAAGCAAACAGAGGCCTTTGCGGGTCTTGTTAAAAGACATGCAATAGCGTCATCAGCAGCTGCTGGAGGTTTAAGAG
CCGCTGCAGAGTGTGTTCAAATAGCTTTAGGTCATTGCTCTTTGTTGGAAGCTCGTGGTTTGGCACTCTGTCCTGTGCTATTAAAACTCTTTAGGCCTAG
TGTTGAACAAGCCTTAAATGCCAATCTAAAACGAATTGAAGAGAGCACTGCTGCTTTGGCTGCTGCTGATGATTGGGTACTTACTTATCCTCCAACCAGT
ACTCGGCAGTCTGGCAGGTCTTCTGTTACATCTCTTGGTAATGCAGCAGCATTTCAACATAAACTTACAAGTAGTGCCCATCGCTTCAATTTAATGGTCC
AGGACTTCTTTGAGGATGTAGGACCACTCTTAAGCATGCAAATGGGAGGCCAAACACTGGAAGGTTTGTTCCAAGTATTTAACTCGTACGTGAACATGCT
CATAAAAGCTTTACCAGGTTCGATGGAAGAAGAGGCAAACTTTGAAGGTTGTGGAAATAAAATTGTGCAAATGGCTGAGACTGAAGCTCAACAAATTGCA
TTACTGGCTAATGCTTCATTATTAGCAGATGAACTACTCCCACGTGCAGCCATGAAACTTGCACCTCCGAATCAGGCTAATTACAAGGATGATTCCCGTA
GAAGACCTTTAGATAGGCAGAACCGTCATCCTGAGCAACGAGAATGGAGGAAGCGGCTTGCAGGTTCGGTTGATAGATTAAAAGATGCTTTCTGTCGACA
ACATGCACTGGATCTCATTTTTACAGAAGATGGTGATAGCTATCTTACTGCAGAAATGTATACAAACATGGTTGGAAGTGCAGATGAGGTAGATCGGTTT
CCATCCCCAATATTCCAGGAACTTTTTGTAAAACTGAACCGTATGGCTAGTATAGCAGCAGAGATGTTTGTAGGAAGGGAAAGATTTGCAACATTACTAT
TGATGAGACTTACAGAAACAGTCATTTTATGGCTTTCAGAAGATCAAAATTTTTGGGATGATATCGAGGAAGGTCCAAGGCCTTTAGGTCCTCTTGGTAT
ACAACAGTTCTATTTGGACATGAAGTTTGTCATGTGCTTTGCTTCCCAAGGACGCTACTTATCACGAAATTTGCATCGCGTTGTCAATGAAATAATTGCA
AAAGCTTTGGCAGTATTCTCTGCAACAGGGATGGATCCAGACAGGGAGCTGCCAGAAGATGATTGGTTTAATGACATTTGCCAAGAAGCAATGGAAAGAT
TAAGCGGAAAACCCAAAGCCATCGATGGGGATAATGAACTTGGTAGCCCAACAGCCTCTGTCTCAGCACAATCAATTTCATCAGTCAGATCTCATGGAAG
TTCATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G009100.1 pacid=42806115 polypeptide=Potri.008G009100.1.p locus=Potri.008G009100 ID=Potri.008G009100.1.v4.1 annot-version=v4.1
MASAKTSSRSRGTPVKENGTKLEEGLNVFKSDRFNADSYVQSKCSLNEKEIKQLCSYLLDLKRASADEMRKSVYANYAAFIRTSKEISDLEGELLSIRNL
LSTQATLIHGLVEGVNIDSLSLKASEGSLVNGLENVEDREPTDLERWLAEFPDMLDVLLAERRVDEALAVIDEGERIAAEMKKTELSSPGILRSLEIAIT
ERGQKLADQLAEAACQPSTRSSELRAAISALKKLGDGPRAHSLLLNAHLQRYQYNMQSLCPSSTSYGGAYTAALSQIVFSAIVQASSDSLAIFGKEREYR
SELVMWATKQTEAFAGLVKRHAIASSAAAGGLRAAAECVQIALGHCSLLEARGLALCPVLLKLFRPSVEQALNANLKRIEESTAALAAADDWVLTYPPTS
TRQSGRSSVTSLGNAAAFQHKLTSSAHRFNLMVQDFFEDVGPLLSMQMGGQTLEGLFQVFNSYVNMLIKALPGSMEEEANFEGCGNKIVQMAETEAQQIA
LLANASLLADELLPRAAMKLAPPNQANYKDDSRRRPLDRQNRHPEQREWRKRLAGSVDRLKDAFCRQHALDLIFTEDGDSYLTAEMYTNMVGSADEVDRF
PSPIFQELFVKLNRMASIAAEMFVGRERFATLLLMRLTETVILWLSEDQNFWDDIEEGPRPLGPLGIQQFYLDMKFVMCFASQGRYLSRNLHRVVNEIIA
KALAVFSATGMDPDRELPEDDWFNDICQEAMERLSGKPKAIDGDNELGSPTASVSAQSISSVRSHGSS
|
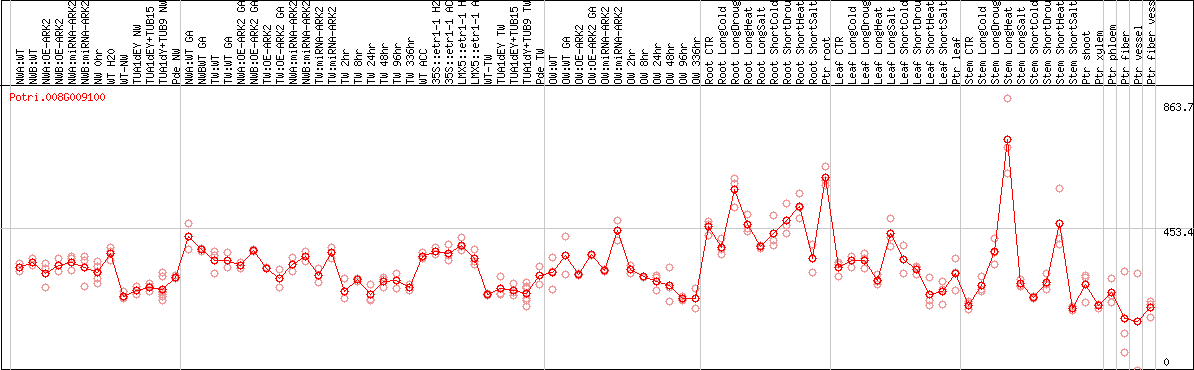
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G009100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
