Potri.008G020600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G020600.1 pacid=42807104 polypeptide=Potri.008G020600.1.p locus=Potri.008G020600 ID=Potri.008G020600.1.v4.1 annot-version=v4.1
ATGGCTGGAAATGTGAGGTACGATTTGAGTTCTGCTAGTCCAGAGGAATTAGGTTTTACAGGTAGCTTTTCTAATGGGCAGAGGGGGAGTTATCCTAATG
CCAGTTTTGATAGGTCTGGAAGCTTCCGGGAGAGCAGTGAGAGCAGAATGTTCAGTTCTGGGGCAAGTACACCCAGGGCTTCTGCTTCACCAGCAAGGAG
CATGGGCCCACTTACTCAACATCTGTCATTGGATCCGGTTACTATGGGGGACCCAAAATATACTCGAACAGGCGAGTTAAAGAGGGCTTTTGGGATATCT
CTTGGGAGTGCTACAGAAGATAATTCTTTTGGAGCTGCTCATTCAAAGCCACCCCCTGCAGTAGATGTAGAGGAACTGAAGCGGATCAGAGCAGGTGTAT
TAGATGACTATCGGAAGTCTAGGAATAGAGCAAAAATGTGGAATGAAAACTTGCTAAGACTTCAAAAATTCCCAGAAGATTTAAATTCAAAGAATCAACA
ACGAAGTGAGATGCTAATGAATGAAAGATCTGGTGGTTCAAATTTTTTGAAGATGGGAACGCAGATTCATCGAAACCCTTCAGACCTTGGGACTCAAAGA
TTAGAGGACAGGACTAAGACCATTGTTTTGAATAAGCGTGTTCGCTCTTCTGTAGCAGAATCACGGGTTGATGGCCGAAGCAATACAGTTCTGAGGCAGC
CTTTGGTGACAGGAAAAGACAGAGATATACACAGGGATGGTGAAGTTTCTAATCTTACTGAAGAAAAGGTCCGTAGGCTACCTGCTGGAGGAGAAGGGTG
GGACAAAAAGATGAAAAAGAAACGTTCTGTTGGCACTGTTTTTACCAGGACAATAGACAGTGACGGGGAAGTTAAACGAATGATGAATCATAAGTTTAAT
AATGAACACAGTCTGCAGTCTTATGATGCCCAAGGCTTCAGGTCAGGATCTTTTAATGGGAGTAGTGGCATGAACAAGGTAGATGGAATTTCATCATCTG
CTAATTCAAATACCCGTGCCATTCCCAAGGAATCGGAAAAAGTTTCTTTGACAAGGGATTATGCTGCTGGCATGAATAAGGAGCGACTTGTAGTGAAAGC
AAACAATAAGGTAAATATCACAGAGGACAATAATCACACAGTGAGTCCAAGTCCATTGACAAAAGGAAAGGCTTCCAGGACACCTCGAACAAGCTCATTA
ATGGCAGCAAGTACATCTACTAATACTCCCCTCTCACCTGGGGGTTTTGATGGTTGGGAACAACCACCAGCCATAACCAAGGTCAATTCAGTTGGGGGGC
CTAATAATCGCAAGCGTCCCATGCCTACAGGGTCATCATCTCCTCCAATGGCTAAATGGGTTGGTCAGAGACCACAAAAAATATCTCGTACCAGAAGGGT
GAATGTAGTGTCTCCCGTCTCTAATCATGATGAAGGGCAGATGTCATCAGAAAGAGGACATGTTTCTGATTTTGCCACTCGAGTGACTTCTGGGATTGAT
GGGCCTCCTCTTGCCAAGGATGTGCTTAATGGAACCACACAAGTCAGAGTGAAACATGAAAATGTTTCATCTCCATCAAGATTATCTGAAAGTGAAGAAT
CTGGTGCTGGTGAAAATCGTGAGGGTAAGCCCAAGGACAAGAGAACAGGTAGTGGTGGGGTAGAGGAGAGATCGCTGAATCAGAATGCTGTTCCTTCTCT
ATTGGTTACAAAGAAGAATAAAACACTCGGTAGGGAAGACACTGGTGATGGTGTGCGAAGACAAGGAAGGACTGCTCGGGGTCCATCTTCTAGGACAAAC
ATTTCACCAATGAGGGAGAAGTTGGAGAATCCAGCCTCGACCAAACCACTAAGAAATACGAGGCCCATTTCAGACAAGAGCGGAAGCAAGACAGGTCGTC
CTCCTCTTAAAAAAATATCAGATCGCAAGGCCTTCACTCGACTTGGGCAAATACCAATAAGCGGTTCCCCAGATTTCTCAGGTGAATCAGATGACGATCG
CGAAGAACTCTTAGCTGCTGCAAATTTTGCTTGCAATGCCAGCTATCTTTCCTGTTCTGGTTCTTTTTGGAAGAAAATGGAGCCAGTTTTTGCTCCCATC
TGCTCTGGGGATTCATCCTACTTGAAGCAACAGTTGAAGTCTGTGGAGGATCTCCACAAGAGATTATATGAGATGTTTGACTGCAGCAATAATTCGGGTG
ATTTTGTGCTTGAAGAAGATATTCCATCCCAACTTATTCATGAAGAAAGTGAGAGAAACTTGCAGGACCAGGATCCACCAAAAAAGTTGGTGAGGACTTC
AGATTTGGTAGATCCAAAACAGGACAATAGTGCTGTATGTGGAGGCTCAAGAACAAGAAACAAAGCTACTCCACTGTACCAAAGAGTGTTGTCTGCTCTG
ATTGTAGAAGATGGGTCAGAAAAATTTGCAGAAAACAGTGGAGGGAGAAATATTTCTTTTCAATGTACTGGAGATAGTTCTCCAGGTGATGATTGTCTCT
CAGTGGATTTTGAGCCCGGGAGCACAAATGGCATTGATTTTAATTATGAATCTATGCTGGGTTTTCAACATCAGAAGCAATCTTCCGTTGATGGTTTTTC
TTGTAATGGAAATAGTACTGTTAATAGGATCGGTGGTTTTCACAATAATTCCTATATTGATCATTTGGTGCAAGGAGGTAATGGATTTATGCACTCAAAA
ACCGGAATGTTCCCTGGGAGTTTTGAGAATAATGATGAGAAATCAACCATTCATTCAAATGCCATCAGCATGTCTGCATATGATTGCCAATATGAGCAAC
TAGGCCTGGAGGACAAACTCTTGATGGAGCTGCAGAGTGTTGGTTTATATCCAGAAACAGTGCCTGATCTAGCAGATGGAGAGGATGAAGCAATTAATGA
AGATATTATTGAGCTCCAGAATAAACTTCAACAGGTTGGTAAAAAGGAACACTTGGACAATTTAACTAGAGCTGTTGAGGAGGGAAGGGAGTTGCAAGAA
TGGCCCCTTGAGCAGGTTGCAATGGATAGGCTTGTTGAATTGGCTCACAGAAAGCAGTTGGCCACCAGAGGAAATAATGCTTCAAAATTTGGGGTTCCAA
AGGTTTCGAAACAAGTTGCCTTGGCTTTTACAAGGAGGACTCTTGCTAAATGTAGAAAATTTGAAGATACAGGCAAGAGTTGTTTTTGTGAGCCCCCCCT
TCGAGATGTCATTTTCGCTGCACCTCGTGCAATTGTTGTGGAGTCCACAAGTTGTATTCAGGATCCTGGGGCTTCAGGCTCTTTCACTGGTAGAGCTGAT
AGGCATGATCTTCACAACGATAAATTTGGCAGGGGTGTATCATTAGACCATGATTTTGCCAGAACTGGACCATTATTGAACAGAGGGAGGAAGAAGGAAC
TACTGCTTGACGATGTTGGTGGTAATGCTTTGTTCAAAACCACATCATCTGTTGGTAATACTCAGCTGGGTGGAGCCAAGGGAAAGAGAAGCGAGAGAGA
AAGGGACAAGGATGTATTAGCTAGAAATTCTGTCACCAGAGCTGGTCGTGCTTCTCAGAGCAATATCAAGGGTGACCGTAAAACAAAATCAAAACCCAAG
CAGAAGATTGCTCAGCTATCAGCTTCTGGAGATGGAATTATCAACAAGTTCAAAGAAACAGGTAGCAATAAGAAAAGAGAAGTTGGGGCAACATCCAAGG
GTTCAAATCCTGTAGATTCATCGAAAAAAAGCAGAGCGACAAACATTGCCGAGTTCCAAGATTTAGACTCTATAGAACTACACGAGGGCAATGATTTCAG
TGACACTCAGGATCTGAATAGTTTGTTTGATGGGTTGCCAGAAAACGACTTTGCAGGTGAAATTCTACTTGATGACTTGCCCCTTCAAATTCCTATGGAC
GACCTTTCAATGATTCTATAA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G020600.1 pacid=42807104 polypeptide=Potri.008G020600.1.p locus=Potri.008G020600 ID=Potri.008G020600.1.v4.1 annot-version=v4.1
MAGNVRYDLSSASPEELGFTGSFSNGQRGSYPNASFDRSGSFRESSESRMFSSGASTPRASASPARSMGPLTQHLSLDPVTMGDPKYTRTGELKRAFGIS
LGSATEDNSFGAAHSKPPPAVDVEELKRIRAGVLDDYRKSRNRAKMWNENLLRLQKFPEDLNSKNQQRSEMLMNERSGGSNFLKMGTQIHRNPSDLGTQR
LEDRTKTIVLNKRVRSSVAESRVDGRSNTVLRQPLVTGKDRDIHRDGEVSNLTEEKVRRLPAGGEGWDKKMKKKRSVGTVFTRTIDSDGEVKRMMNHKFN
NEHSLQSYDAQGFRSGSFNGSSGMNKVDGISSSANSNTRAIPKESEKVSLTRDYAAGMNKERLVVKANNKVNITEDNNHTVSPSPLTKGKASRTPRTSSL
MAASTSTNTPLSPGGFDGWEQPPAITKVNSVGGPNNRKRPMPTGSSSPPMAKWVGQRPQKISRTRRVNVVSPVSNHDEGQMSSERGHVSDFATRVTSGID
GPPLAKDVLNGTTQVRVKHENVSSPSRLSESEESGAGENREGKPKDKRTGSGGVEERSLNQNAVPSLLVTKKNKTLGREDTGDGVRRQGRTARGPSSRTN
ISPMREKLENPASTKPLRNTRPISDKSGSKTGRPPLKKISDRKAFTRLGQIPISGSPDFSGESDDDREELLAAANFACNASYLSCSGSFWKKMEPVFAPI
CSGDSSYLKQQLKSVEDLHKRLYEMFDCSNNSGDFVLEEDIPSQLIHEESERNLQDQDPPKKLVRTSDLVDPKQDNSAVCGGSRTRNKATPLYQRVLSAL
IVEDGSEKFAENSGGRNISFQCTGDSSPGDDCLSVDFEPGSTNGIDFNYESMLGFQHQKQSSVDGFSCNGNSTVNRIGGFHNNSYIDHLVQGGNGFMHSK
TGMFPGSFENNDEKSTIHSNAISMSAYDCQYEQLGLEDKLLMELQSVGLYPETVPDLADGEDEAINEDIIELQNKLQQVGKKEHLDNLTRAVEEGRELQE
WPLEQVAMDRLVELAHRKQLATRGNNASKFGVPKVSKQVALAFTRRTLAKCRKFEDTGKSCFCEPPLRDVIFAAPRAIVVESTSCIQDPGASGSFTGRAD
RHDLHNDKFGRGVSLDHDFARTGPLLNRGRKKELLLDDVGGNALFKTTSSVGNTQLGGAKGKRSERERDKDVLARNSVTRAGRASQSNIKGDRKTKSKPK
QKIAQLSASGDGIINKFKETGSNKKREVGATSKGSNPVDSSKKSRATNIAEFQDLDSIELHEGNDFSDTQDLNSLFDGLPENDFAGEILLDDLPLQIPMD
DLSMIL
|
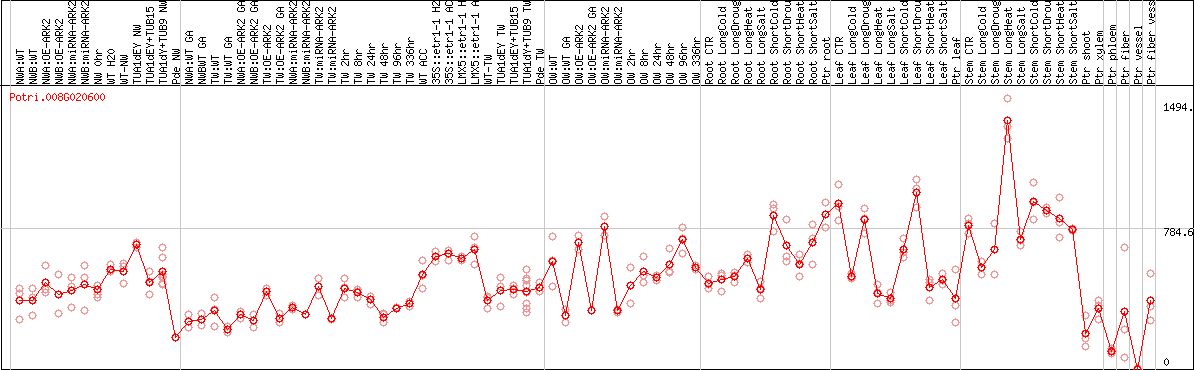
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G020600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
