Potri.008G023101 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G023101.1 pacid=42806555 polypeptide=Potri.008G023101.1.p locus=Potri.008G023101 ID=Potri.008G023101.1.v4.1 annot-version=v4.1
ATGACGACAACAAACATTTCCCCATCTTCAATTGTTGTTGAAGTCCTAAGAGGTGATAACTATGATGACTGGAGTGCCTGCATGAAAAGCTACATGTTAG
CTCAAGACCTTTGGGATTTTATTGAGCCTCCTGGTCTTCAAGAAGGTGATCAAGAAGTTGACTCTAAGCCTATTGATCATCAAGAAGGTGACTCTAAGGC
GTTGAGAAAAAAGAATGCTGCAGCTTTACATGCAATTCAAATTTCTTGCGCGCCATACGTGCTTTCAAAGATTAGGAGCATCACTTCAGCCAAAGTTGCC
TGGGATACTCTAGCCAATTTGCAACAACAACATTCACCTAGCCACAAAGAGCAATCTGATGAAGCTGAGCCATCAGAAGATGATGAGTCACAAGGTTCAG
AGCAATCTGTTGAATCCGAGCCAGCAGAAGATGATGAGTCACAAGGAGCGGTGAGCAATGAAATGAATGGTCCCTTGCTAACCCTGTACAAGTATGCCCA
TAATGGAGACTGGGATGCCATAAAAACCTATCTTAGCCGATACCCAAATGCAAAAAAGGCCAAGATTAAACCCTATGGTAGAACTGCCCTTCATGTAGCA
GCTTCTTCTGGAAATCTGAAAGTTGTGGAGGAGTTGGTGACGTTGATGTCAGTAAATGAATTGGCAATCAAAGATAATGAAGGCAACACAGCTCTTTCTA
TTGCTGCCATTGTTGGAATAAGAAAGATGGCAGAATGTTTGGTGAGCAAGAACGAAAACTTGGTTACCTTCGCAAATAGATATCCAAAGATCCCACTAGT
TGAGGCATGTGTAGGAAGTCAAATGGATATGGTTCGTTATCTATACTCTGTTACTCCAATTGAATTTCTTTGTCGAGGCAATGTCGACCAAGGATCTCGC
TTCCTAAAAAATGCTATTGGTGCTCAAATGCTTGATATTGCTCTTGATTTTCTCCATCGCTGCCCACGTTCTGCTACTACAATGGATGAAGTTCTGAAGT
CGAACGCTTTGTTTAATTTGTCTAAGATGCCTCAAATATTCCCAAGTGCAAGTCGACTTGCATTTTGGCAGCAATGGATCTATTCATGTATTCCCATGCA
ATCAATTGCTACAACTGATGACAATGTTCGTATTAATATGCCTGATCAAAGCCTAAGTGAGTCGAAGAACATTATCCTCCAAGTGTCAAGCAAATTGCGT
GGGTTTGCAATAAATCTCCTCGCATTCTTAGGAATCAAGCAAATATACGATTTGAAGAAGATCCACATTTATTCAGATAAAATTCTACGGTGTATGTGTG
AGCACATATCAACTCTGGATTATGAGGAATACCTCAAGGCAGATGTAGATGGAGCATTCCACAGTGCTGTAGAGAATGGCATGGTTGAGTTTATTATTGA
GGTGGTAAAAGCTTGTCCTCATGCAATGATTAGTGTAGATGGCAACGGAAGGAACCTATTTATGTCCTCGATTGCAAACAGGCAAGAAAAAGTGTTCAGC
CTTTTCTATGGACTGGAAGCAGGCGGAGCAGAATTTGTTTCCATCGTATATGGATCTGGAAATACAATGTTACATTTGGCAGCAAAACTATCACCTCCTT
CTCAATTAGCTCGGATCTCTGGTGCAGCTCTCCAGATGCAGAGAGAACTACAATGGTACAAGGAAGTGGAAAGCATTGTGGATCCTACAGACAATGATTA
TTATAACAAGGACAATCAGACACCCAGGGAATTATTTACGTCTGACCATAAGGACTTGGTAGTGAAAGGAGAGAAATGGATGAAACAAGCAGCAACTTCT
TGTACAGTTGTGGGTGCTCTCATCATTACAATTATGTTTACTGTCGCATTTACTGTTCCCGGCGGGAACGTTCAAGAGACCGGCTATCCGGTTTTTAAAG
ATGAGAAATCTTTTACAGTATTTATAGTAGCGGATGCAATCTCTCTCTTTTCTTCATCAACATCAGTGTTGATGTTCTTAGGAATTCTTACGTCCCGATA
CGCAGAGGAAGATTTCCTTAAATCCTTGCCCACTAAGTTAATCATCGGCCTTTCCATGCTTTTCTTCTCTATTGCAGCCATGATGGTAACCTTCTGTGCT
GCTCTTATAATTATGTTGGATGGGAGATTGCAAGTTATAATTCCAATTGTACTCCTTGCTACTATCCCTGTAACTTTGTTCATGTTGCTGCAATTTCCCC
TGCTAGTGGAGATTGGTGTATCCACATATGGACCAGGAATCTTCAACAGAAAAATGAAACGCTGGTCAAAATCTTCAACTGATAGACTGTTAGAGGTTAA
CAGTTTATTGTCCCAAACTGTTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G023101.1 pacid=42806555 polypeptide=Potri.008G023101.1.p locus=Potri.008G023101 ID=Potri.008G023101.1.v4.1 annot-version=v4.1
MTTTNISPSSIVVEVLRGDNYDDWSACMKSYMLAQDLWDFIEPPGLQEGDQEVDSKPIDHQEGDSKALRKKNAAALHAIQISCAPYVLSKIRSITSAKVA
WDTLANLQQQHSPSHKEQSDEAEPSEDDESQGSEQSVESEPAEDDESQGAVSNEMNGPLLTLYKYAHNGDWDAIKTYLSRYPNAKKAKIKPYGRTALHVA
ASSGNLKVVEELVTLMSVNELAIKDNEGNTALSIAAIVGIRKMAECLVSKNENLVTFANRYPKIPLVEACVGSQMDMVRYLYSVTPIEFLCRGNVDQGSR
FLKNAIGAQMLDIALDFLHRCPRSATTMDEVLKSNALFNLSKMPQIFPSASRLAFWQQWIYSCIPMQSIATTDDNVRINMPDQSLSESKNIILQVSSKLR
GFAINLLAFLGIKQIYDLKKIHIYSDKILRCMCEHISTLDYEEYLKADVDGAFHSAVENGMVEFIIEVVKACPHAMISVDGNGRNLFMSSIANRQEKVFS
LFYGLEAGGAEFVSIVYGSGNTMLHLAAKLSPPSQLARISGAALQMQRELQWYKEVESIVDPTDNDYYNKDNQTPRELFTSDHKDLVVKGEKWMKQAATS
CTVVGALIITIMFTVAFTVPGGNVQETGYPVFKDEKSFTVFIVADAISLFSSSTSVLMFLGILTSRYAEEDFLKSLPTKLIIGLSMLFFSIAAMMVTFCA
ALIIMLDGRLQVIIPIVLLATIPVTLFMLLQFPLLVEIGVSTYGPGIFNRKMKRWSKSSTDRLLEVNSLLSQTV
|
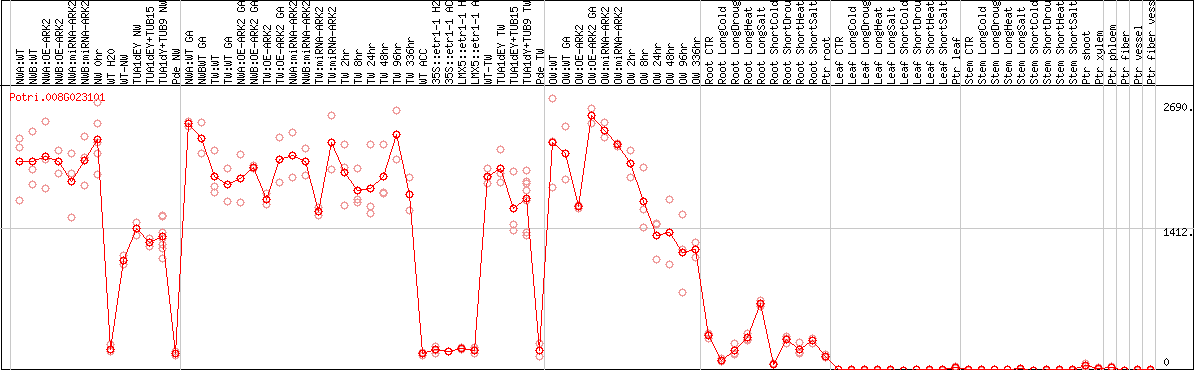
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G023101 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
