Potri.008G037800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G037800.1 pacid=42808429 polypeptide=Potri.008G037800.1.p locus=Potri.008G037800 ID=Potri.008G037800.1.v4.1 annot-version=v4.1
ATGGGTAATGTGGAAATACCCAAATGGCTAAAAGGGTTGCCTTTGGCACCCGAGTTTAGGCCAACTGATACTGAATTTGCAGACCCAGTTGCATATATAT
CGAAAATTGAGAAGGAAGCTAGTGCTTTTGGGATTTGTAAGATCATACCCCCATTGCCTAAGCCTTCGAAAAGATATGTTTTTAGTAACTTGAATAGGTC
GCTTTCTAAGTGTCCGGAGTTGGGTGATGATGTGGACTTGTCGAATGTTTGTTCCTCGTCAAACGGTGGTTTAAGGGATGGTGGTAATGATGGGGAGAAT
AGGGCGGTGTTTACAACTAGGCAGCAGGAGTTAGGGCAGTGTGTGAAGAAAGCAAAAGGGATGGTTAAGGAGAATCTGCAATCAGGTGTTCATAAGCAAG
TGTGGCAAAGTGGGGAGGCTTACACATTGGAACAGTTTGAGTCAAAGTCTAAGGCTTTTGCAAGGAGTTTGTTAGGTATGCTTAAGGAGGTTAATCCATT
GGTTATAGAGGCATTGTTTTGGAAGGCAGCTTCTGAGAAGCCTATCTACGTGGAATATGCAAATGATGTGCCTGGTTCGGGTTTTGGGGAACCAGAAAGC
CACTCTAGGTACTTTCCTAGACGGAGGAGGAAGAGGGCATCATATCAGTCTTATCGCAGGAGTAGAGAAAGCCCTGTTTGCAGTACAAATGATATGGATG
ATGTAAAGAATTCCCATAATGATGAGGTTAAAGGTGTTTCCATTAAGAATGTGCCAAGCTTGTGTTTGGAGACGACACCCAGGTCATCTATGGCATCTTT
GACTTCATTCGCAGAGGATAATCTGATGTATTCTAAGCAAAAAAGTGTAACTGCAACTAATGATATGGAAGGTACTGCAGGTTGGAAACTTTCAAATAGC
CCTTGGAATCTGCAGGTGATCGCGCGCTCACCTGGCTCTCTTACACGATTTATGCCAGATGATATCCCAGGTGTTACTTCACCTATGGTCTACATTGGTA
TGTTGTTTAGTTGGTTTGCTTGGCATGTTGAGGATCATGAACTTCACAGCATGAATTTTCTACATACCGGTTCTCCAAAGACTTGGTATGCTGTCCCTGG
AGATTATGTATTTTCTTTTGAGGAAGTTATACGCACTGAGGCTTATGGTGGCAATATTGATCGCTTAGCTGCCCTGAGTCTGTTGGGTGAGAAGACAACT
CTTCTGTCTCCTAAAGCAATAATTTCCTCAGGCATTCCTTGTTGTAGGTTAGTACAGAATCCTGGTGAATTTGTTGTGACTTTTCCAAGGGCCTACCATG
TAGGATTCAGCCACGGTTTTAACTGTGGGGAAGCTGCCAATTTTGGAACTCCACAATGGCTTAAAGTAGCCAAGGAAGCTGCGGTCCGTAGAGCTGCCAT
GAACTATCTTCCCATGCTTTCACATCAGCAGCTTCTATACCTGTTGACCATGTCTTTTGTTTCAAGGGTGCCGAGATCCTTACTACCAGGTGCTCGGAGT
TCTCGTCTTAGAGATCGTCAGAGAGAAGAAAGAGAGTTGTCAGTAAAGGAGGCTTTTCTAGAAGACATGTTAAAAGAAAATGACATTTTATCTGCTTTTC
TTGAGAAAAACTCCACTTGTCATGCGGTGATATGGAACCCTGATTTGCTTCCTTGTGCAAGTAAAGAGTCTCACTTACTCAACATAACTTCTACTATTAC
TACCACACCAAAACAAAATGCTTCCCATAATAATTTCGATGTAAATAGGAATTGCAATGAGAATGACCTGTTTAAGGAGATGAGCTTGTATATGGAAACT
CTCGATGATTTATATATGGAAGAAGATGATTTGTCATGTGATTTTCAAGTTGATTCGGGGACATTAGCATGTGTGGCTTGTGGAATCCTTGGTTTTCCTT
TTATGTCAGTGTTGCAACCACATGAGAAAGCATCAATAGAATTGATGCCCGGCGAAGAGCCAAGAGTTACAAGAATTGATAATGTTCAGCCCTCTTTAGA
CTCTGATAGCACCGGCAAAGGCTCTGTTTCAGATGATCATGGTCCTGTCAAAGATTATTCTGTGCCTCTGAAGGATCTTCCAATGCCCACAGGGTGGAAC
ACTTCTCATAAGTTTTTGAGACCTCGTATTTTCTGCCTGGAGCATGGTGTCCAAATTGAGGAGCTGTTACAGTCTAAAGGTGGAGCTAACTTGCTTATAA
TTTGCCATTCAGACTATCAAAAAATAAAAGCACATGCTTATGCCATCGCTGAGGAAATTGAGAGTCCTTTCAATTATAATGAGGTTCCTTTAGAAGCTGC
ATTGAAAGAAGATCTGAACTTGATAAATCTTGCAATTGATGATGAAGATCATCATGAATGTGGAGAAGACTGGACATCAAAACTGGGCATCAACTTACGG
TATTGTGTAAAGATCAGAAAGAACTCTCCATCTAAGAAAGTCCAACATGCATTGGCATTGGGAGGATTATTCTCTGACAGAAGTCTCACAGATTTCTTAA
ACATCAAATGGCAATCTAGAAGATCTCGGTCAAGAATAAAGTTAAATCAGCCTTTCCATCCTAAACCATGCAAGATTATTGAACCAGACAAAGATGAAAT
GTCGGGGAATAAGTCAGATGGTCTCACTGTCAAAAAAGAGGAAAAACTTGTTCAGTACACTAGAAGGAAGTACAAGGTAAAAATAGATTATTCAACAAAT
GGGCTTGAAGGATGCTCTAGAAGGTGCTTTGCAGAAGAGGTTTCAGGAGCTAGTGGTGATGATCCTGATAAATATACTGAACAAACAACTGTAATTTACC
CTTGCAATATTGGGATTACCAGAAGTGGTTCTGCAGGATTTGGTTTTTCTCCCATTGAAGATCCTGAAATGCTGCATGAAGTCAAGGTGCTTGAAGCAGC
TGGTGGTTTGACCTTGAATTCTGCTCCATCACAAGATGCATGTTCAGTTCTCACTGCTACTGTTGCTGTCAAAAGTGTTGGGGGCCAGATTGAGGATCAA
CTGCTGAAGGAATCAAAAAATGCGAGAAATATTTGTAATGTAAAAGCCAGTGGTACTTCTGAGATAGAGCATCAGATAAATGCCTCTGGAGGAACCAGTG
AAAAACAGGATTTTTACGCCACAAAATGCTGTAGTCCATTCATCACTGTAGCTAATGAAAGATTTGAAATGCAGAGAGAGGATCAAGTTTTGGGAAATGT
CAACATGGGTGAGACTTGTAATATGGTTTCTGAAGGGCAACAGAGGGTCCTGGATGATGGCGATGCTTCAGTAGACGAGGTTTCTGATCTTGCTAACGTA
GCAAGTTTACATGTTTCTCCCCCCCCCATCGGACTAAGGGCGGATGTAGTTGTGGAGAATTCTTTCATTAACAATGAGGTATCTCCTCCTGTGACTTTGG
ATGATGAAGTGAAAAAAGAACTTGTGACTAAAAACAGAACTAATGGTGACCAATGCTCTTCAAGTGATGACACATTGATGAATCAGCCTACTACCTCACT
GGATGAAAGATGTGGTCATGAGCAAGAGACCCGTGCTGTACAAAATAAGACGCAGAAAGAAGCTGAAATAAAAAATGGGAGTAATGATGAAATCGTTCCG
AGTAATGTTATTAGTGTGACTAATCTTGTGCCTATAGATGAAAGTTCTGAATTTCATAGAGAGCTGCATGCCACAGTCAACTTGTGCAATGGCATGGCTT
TTGAAAATGGCAAGCAGCTGGTGTTTCAGACCACAAATGATTCTAACAAGGAACTTATCTCATGTTCTGTTGCACAAATGGAAATAAATTCATCTACTGC
TTCATCAGAATTCTCTAAACTTCATAGGCAGGCCTATGCTGAAAATGATTTGTGCAGTGGCTCAACTTTGGACACCATTGTGCCACCAGAAATTCCAACT
ACAGACATACGCACTGTGGAGGAACTTGCCTCTAATTCTGCAACCATAAATCAAGAACTTTCTGAAGCTTCAAAAGAGATTTGTGCTATTCAAGACTCGT
ATGCTTGCATGGATTTAGAACCTGAAGTAGAGAAGGAAATTCATTCCAGTGATGGGGTGACTAGAGACAGTGAGGTGCAGAAAATACACCAGGGCACAAG
TTTGATTAATGAGGACATACATGTTTCTGCTCGTGTTATTCTGGTAAATCAGCCGCCGACTCCTTCCCCAGTTATAAAATGTTCCAACATTGATGACAAA
TCATGTGTTGGAGAAAGCATGGTGACGTGCAACAAATTTTTTTCATCTCAGGAGATTGAGAGCATCGAGTCTGCTGTTGTAGACTCTAGACCAACTGCTG
GAAAGGGAAGAAAAAGAAAGGGTGAAGTGGAGCAGCTAACAGAAAACAAGTTCGATAGCAATGGCTTTATTAGGAGTCCATGCGAAGGATTGAGACCGAG
GGCTGGGAAAGATGTTGGGAAATCAGCTGAGGAGAATCCAATACCAAAGAGGTTAAAGAAACCTTCTAATGTTTCAGTTTCTCGCTCAAAAAGGAAAGAA
ATCACACAAAGATCTTACAAGTGTGACCTAGAAGGTTGCCGCATGAGGTTTGAGACAAAAGCAGAGTTACAGTTGCACAAGGGCAATCGATGCACATATG
ATGGATGTGGTAAGAAGTTTAGTTCCCACAAATATGCAATAGTTCATCAACGTGTTCATGAAGATGATAGACCCCTCAAGTGTCCATGGAAGGGTTGCAC
CATGTCATTCAAGTGGGCATGGGCTAGGATTGAGCATATACGGGTGCACACTGGTGAAAAGCCATACCAGTGCAAGGTCGATGGCTGTGGCCTTTCTTTC
AGGTTTGTATCAGACTTTAGTCGGCATAGGAGGAAAACAGGACACTATTTGAACACACCTGACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G037800.1 pacid=42808429 polypeptide=Potri.008G037800.1.p locus=Potri.008G037800 ID=Potri.008G037800.1.v4.1 annot-version=v4.1
MGNVEIPKWLKGLPLAPEFRPTDTEFADPVAYISKIEKEASAFGICKIIPPLPKPSKRYVFSNLNRSLSKCPELGDDVDLSNVCSSSNGGLRDGGNDGEN
RAVFTTRQQELGQCVKKAKGMVKENLQSGVHKQVWQSGEAYTLEQFESKSKAFARSLLGMLKEVNPLVIEALFWKAASEKPIYVEYANDVPGSGFGEPES
HSRYFPRRRRKRASYQSYRRSRESPVCSTNDMDDVKNSHNDEVKGVSIKNVPSLCLETTPRSSMASLTSFAEDNLMYSKQKSVTATNDMEGTAGWKLSNS
PWNLQVIARSPGSLTRFMPDDIPGVTSPMVYIGMLFSWFAWHVEDHELHSMNFLHTGSPKTWYAVPGDYVFSFEEVIRTEAYGGNIDRLAALSLLGEKTT
LLSPKAIISSGIPCCRLVQNPGEFVVTFPRAYHVGFSHGFNCGEAANFGTPQWLKVAKEAAVRRAAMNYLPMLSHQQLLYLLTMSFVSRVPRSLLPGARS
SRLRDRQREERELSVKEAFLEDMLKENDILSAFLEKNSTCHAVIWNPDLLPCASKESHLLNITSTITTTPKQNASHNNFDVNRNCNENDLFKEMSLYMET
LDDLYMEEDDLSCDFQVDSGTLACVACGILGFPFMSVLQPHEKASIELMPGEEPRVTRIDNVQPSLDSDSTGKGSVSDDHGPVKDYSVPLKDLPMPTGWN
TSHKFLRPRIFCLEHGVQIEELLQSKGGANLLIICHSDYQKIKAHAYAIAEEIESPFNYNEVPLEAALKEDLNLINLAIDDEDHHECGEDWTSKLGINLR
YCVKIRKNSPSKKVQHALALGGLFSDRSLTDFLNIKWQSRRSRSRIKLNQPFHPKPCKIIEPDKDEMSGNKSDGLTVKKEEKLVQYTRRKYKVKIDYSTN
GLEGCSRRCFAEEVSGASGDDPDKYTEQTTVIYPCNIGITRSGSAGFGFSPIEDPEMLHEVKVLEAAGGLTLNSAPSQDACSVLTATVAVKSVGGQIEDQ
LLKESKNARNICNVKASGTSEIEHQINASGGTSEKQDFYATKCCSPFITVANERFEMQREDQVLGNVNMGETCNMVSEGQQRVLDDGDASVDEVSDLANV
ASLHVSPPPIGLRADVVVENSFINNEVSPPVTLDDEVKKELVTKNRTNGDQCSSSDDTLMNQPTTSLDERCGHEQETRAVQNKTQKEAEIKNGSNDEIVP
SNVISVTNLVPIDESSEFHRELHATVNLCNGMAFENGKQLVFQTTNDSNKELISCSVAQMEINSSTASSEFSKLHRQAYAENDLCSGSTLDTIVPPEIPT
TDIRTVEELASNSATINQELSEASKEICAIQDSYACMDLEPEVEKEIHSSDGVTRDSEVQKIHQGTSLINEDIHVSARVILVNQPPTPSPVIKCSNIDDK
SCVGESMVTCNKFFSSQEIESIESAVVDSRPTAGKGRKRKGEVEQLTENKFDSNGFIRSPCEGLRPRAGKDVGKSAEENPIPKRLKKPSNVSVSRSKRKE
ITQRSYKCDLEGCRMRFETKAELQLHKGNRCTYDGCGKKFSSHKYAIVHQRVHEDDRPLKCPWKGCTMSFKWAWARIEHIRVHTGEKPYQCKVDGCGLSF
RFVSDFSRHRRKTGHYLNTPD
|
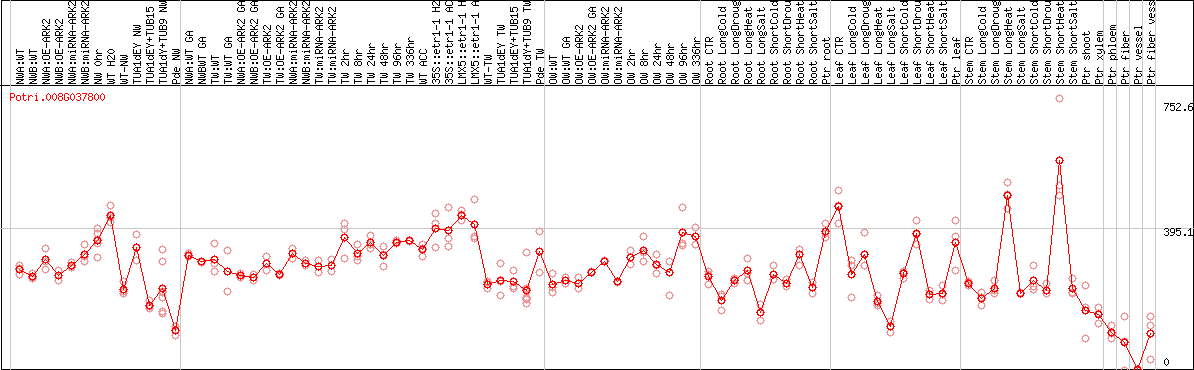
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G037800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
