External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G54770 233 / 6e-77
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G76460 148 / 3e-43
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G22910 145 / 1e-42
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT1G20880 145 / 3e-42
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G46780 137 / 2e-38
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G78260 135 / 2e-38
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT1G22330 134 / 1e-37
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G33470 131 / 4e-37
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT5G11412 107 / 2e-28
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G53680 102 / 8e-27
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G216800
409 / 4e-146
AT3G54770 216 / 4e-70
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.013G094400
155 / 4e-46
AT1G33470 257 / 3e-86
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.019G068400
154 / 1e-45
AT1G33470 246 / 7e-82
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.005G256700
143 / 3e-41
AT1G76460 335 / 5e-116
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G179100
142 / 5e-41
AT2G46780 275 / 2e-91
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G004600
141 / 1e-40
AT1G76460 377 / 9e-133
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.002G098100
140 / 2e-40
AT1G78260 164 / 5e-49
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.014G105101
139 / 3e-39
AT2G46780 252 / 4e-82
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.012G022100
127 / 1e-35
AT2G46780 161 / 1e-47
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041423
222 / 3e-72
AT3G54770 181 / 3e-56
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10036503
182 / 6e-57
AT3G54770 156 / 5e-47
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10002684
137 / 1e-38
AT2G46780 228 / 1e-72
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10030202
137 / 1e-38
AT2G46780 224 / 2e-71
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10023485
123 / 2e-34
AT3G54770 71 / 6e-15
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10039151
110 / 2e-28
AT1G33470 182 / 7e-57
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10043168
81 / 3e-17
AT3G07810 456 / 2e-157
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10032579
81 / 3e-17
AT3G07810 447 / 8e-154
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10023817
79 / 8e-17
AT3G15010 188 / 5e-56
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10043016
79 / 1e-16
AT3G56860 463 / 1e-160
UBP1-associated protein 2A (.1.2.3.4.5)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.008G044700.1 pacid=42808579 polypeptide=Potri.008G044700.1.p locus=Potri.008G044700 ID=Potri.008G044700.1.v4.1 annot-version=v4.1
ATGACAATGAGCAACGGTGTAGGGCAGTTTGGTGATACCACTCTGACCAAAGTGTTTGTTGGAGGGTTAGCGTGGGAAACACCAAAAGAGGCCCTGAGAG
AGCATTTCGAAAAGTATGGTGAGATCTTGGAGGCTGTGATTATTTCTGATAAAATCACTGGTAGATCCAAGGGCTACGGATTTGTTACGTTCAAGGAGGC
TGGAGCGGCAAATAAGGCTTGCGAGGATGCTGCACCCATCATCAATGGCCGCCGTGCCAACTGCAACCTGGCTTCCCTTGGTGCACGCCGCCCTAGGTCT
TCAACACCGGCCCCTCCTCAGCAAGGACCTAACATCATCGCTGGACCGAGGTCCACGCCAGCACCGCCAGCAAATCACGTCCAGTGGTATTATCCGGCAG
GGTCAGCTGCAGCGTCACCATTTCACCATCAGCATCACCAAGCCGTTCCTTTTTATGGGTACTCTCCTGCTTACATTGCCACAGATATCAGTTACAATCA
TAAGCTAAGCTACACCGGCGGGTCCTACATGAATGGGCATTTCTCTCAGGTGTATCCAGGGCAAGCTATGGTGGGTGCCAATACATTGATGCCAATGTAC
CCTCTTTATCACTTCCATCAATCACAGGCGGTGGGCCTACCTGCTGCCCACATCTTCCCACCAAACACGGCGGGTCCCATGACAACGGTCCCAACTATCA
TGTCCAAACCCCCATCCATTGCTCCTCCTAACGCAGGTACGGCCGGCACAGGTGAAAGCCTTAAAAGGGTTGGATAA
AA sequence
>Potri.008G044700.1 pacid=42808579 polypeptide=Potri.008G044700.1.p locus=Potri.008G044700 ID=Potri.008G044700.1.v4.1 annot-version=v4.1
MTMSNGVGQFGDTTLTKVFVGGLAWETPKEALREHFEKYGEILEAVIISDKITGRSKGYGFVTFKEAGAANKACEDAAPIINGRRANCNLASLGARRPRS
STPAPPQQGPNIIAGPRSTPAPPANHVQWYYPAGSAAASPFHHQHHQAVPFYGYSPAYIATDISYNHKLSYTGGSYMNGHFSQVYPGQAMVGANTLMPMY
PLYHFHQSQAVGLPAAHIFPPNTAGPMTTVPTIMSKPPSIAPPNAGTAGTGESLKRVG
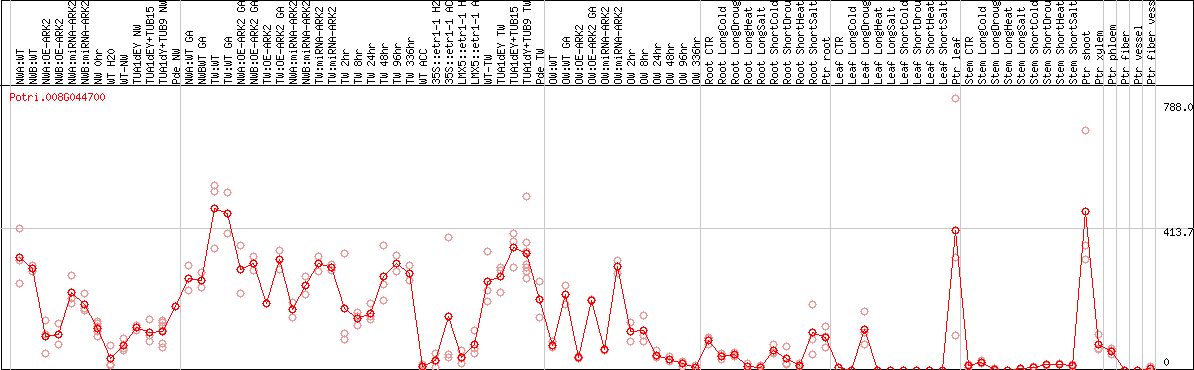
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G044700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.