Potri.008G047600 [POPLAR]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Flax homologues |
No hit found |
||||||||||
| PFAM info | |||||||||||
|
Representative CDS sequence |
>Potri.008G047600.5 pacid=42806484 polypeptide=Potri.008G047600.5.p locus=Potri.008G047600 ID=Potri.008G047600.5.v4.1 annot-version=v4.1
ATGCCGGTGTCAGGAAATGAAGAGACTGGGGTTAAACCCCATGCCCAGCAGTCTAGTCTTAACATTGCAGGTGTTCCTATCAAGAAAAGGAGGTTCATCT
GGCCTCCTTCGCCTCCACCAGAAGAACAATCTGTGCCACTGTTGGGAAATGATTCTGCACAGAAAGAACCTGGCAGCACATCTAAGGAGTCAAGTCCTTC
CAATTCTAGTGTTGCAGCGAGTTCTGATTTATCTGATCCATTCAAGAATTCTGTAGCTGAAGAAAATAAGAACAGGTTGGACAGCATTGTGCAAATGAAT
GCTGAGAACTGCTCAGGAGTTAAAGTTGTAGCACAGAACCTTGCAACCCATTCTGATTCTTTGGCTAAGTTTGGCAAACAAGAGAAGCCCGTGGTGGAAG
AAAAGTCTGCCAACACATTGTTGATTTCAGCGAAAACTGAGTTGAATTTAGAATCCAATAAAGGTCCTGGACTCAATGTTGGTAAAGAGATATGCGGCCA
ACAAATATTGGAAGGAAAGTGTAAATCAGAGATGCCTATAGCTTCAGTAACTTCTCAGTTTTCTTTAGGTTTAAAGGAACACGATGTTTCATCTTTGGAA
TGTTATAGCAATGATGGTAGTCAAATTAATGAAAATGTAGGAGCTGTTTCATTGAATTTGTCTTTAAGCGAGGGTGAGACTGGTGTTCTGCACAAAATGG
ACAATATTCTTGCTACCGACAGCACTGATGTTTTTGCTAACAGATCGAACTGGGATTTGAATACTACTATGGATACATGGGATGGTTCTTCTAGTGATGA
ACATGCTGCTCAAGAAACTGCTGATGGGTGGAACAGAGTTGGTGTCAAATGTGATATTACTACTGGAATAGTTGGAGCTGGTATGTCTAATGGAAGGCAA
CTCCTTGATAGCAGTGAGTGCAAATCTAGTTTCCCCCAAACATTTTCTGATTGTGCCAGGGAGTATACATCTGAGGATTCTCTTCATCTTCGTCTCAGTC
CATCCTTTCCTTCTTTTAACTTAAGCCAGGAGCATTCCAGTTCTTCTGCTAATAAAGAAAGCTGCATTATTCCTAACATTAGCTTGCCTGGAAGCTTGTT
GTCAGCAGGCAATGCAACCGTGGCTAACTGTAGGGGCATAAAATCAGAACCATTTGATGGGAGCCTCAAACATGATCTTAGAGGAGCCAAAGTCAATCCA
TTTGATTTTTTTGTTAAGCGCGAGTTAGTAGAGAAGGGTAGTCTAGAAACCTCGAAGTCGTCAGCTTCTGGTTCTTTGAAATTAGTTGGTCATGGATTCA
TAAAACCAGAACCATTTCATGATGGCAAACCAGAAACACCCAGGATGGTAGGTGGGGGGTCAATCCAACCTGATAAACAGGTGCTACAAAGTCAGGATAC
TGGAGAGCAATCTCCTTGTTCAGCAAGTAAAATCGTGCTGCAAGTTCAGGATACTACAGGACAACCTTCATGTTCAACAGATAACCAGGTGAGGGAAGGT
CAAGATATCTTAGCAAAACCTACTTCTTCAACAGATTTGTTTATAAGTGGCAATGCATCAGACCGATTAGAGTATACCACATGTGTTGAAGGTGCCCTTC
TTAGGAACGCTATGCCAAAGGAAGCACCTGAAAGTGCTGGGCAGGTTTCTTCAGAAATGGTTTCTATGCCTGTAGGTCACAGTGGTGAGGAATTAGATGC
TTCTGTTAAGATAGATACTGCAATTACTATGGACAGAAATGGTGATGCCCCGGAGCAGTGTGAACTGAAAATTACAGAGGAAGTCCCTGCAGGTTCACAT
GGAAATGGTGAGGCTTCTGTAACCGATGAGGAAAAGATTAACTTGTCTGGTGATATGATAGAAGAAGATTCTTATGGCTCTGGCTATGAGTCCGATGGTA
ATACTATGTCTATGGATATAGATGAAGAGCAGCGAGAACACAAGTATGAAGATGGCGAGGTCCAAGACCCGCATCTGCAAGCTGCGGAAGAGTGTCAGAA
ATGTGAGGAAAAAGATGTTAGTCATGGTAATTCTGAACATGAAAAGGCAAACTCTGGACTTGCTGGTGATGATCATTATATTTCATCCCTTGTTGAGGAA
AATGATTCTAAAATAGAACTTTCTGAAAACAATGAAGTTACTGTGAAAGAATGCATCACCAGAACCATTGAAGATGCTGATAATGCTTCTGTGAAAGAAT
CACCAACAGTTGAGATGTCAACTTGTGGGGCAGAGCAAGAGAGGGAAACCACAATCATTCAAAGAAAATCACTTGACTTATCTGGAAAGAAAGATTGTCC
TGTGGGCCAGGGGACAGAACTGTCATCTGGACAAGACATTACTGCAGGTCAAGGGGTTTTAGTTTCTGTTGAGCAAGGTTCAGACGAGAACATTAAAACA
AATTACATGGAAAAGAATGAGTTGCCTGAGTTAGAGGCATCTTTAAATGGTGGTGATATGGCTAAGGATGTCAGCAGCAGCCGGAGTAGGATTATAAACT
TGCCTAGAGCTTCTAATTCATCATCTCCTGGCAAGACAAGATCTATATCAGGCAGGCCTTTTTCATCATATCAAGAAAGATTACCTGACGGGCCGCTTGA
GGGTGGAAAACTACATCCCCAAGGGAGGGATGAAATATATATTGATGGTCCTCGCAGATTCTCAAGAGATAGACACCAAGAGCATTTTCCCAGAAACTCT
AGAATGAATTTTGTCCGTGGTAGAGGGAGGATTTCCAGCCGGGTTGACACTCTTCGTGGTGACAGGGATTCTGAACGTAACTATGCTTCAGAATTTTACA
ATGGTTCATCAGATTTTGCTGTTCGCAGGCATAAATATGCTTCTGCTGCTGCTGAGGCTGATTCTGAATCTATTAATTATAATATCGCACCAGATGGTTC
ATTTGTAGGTACTGCACGGGGAGGGAGGAAGCTACTGGATGATGAGACACCAGTTTTCCGCAATGTACCCTCAAGGAGGCGGTCTCCTGAAGGGAGAGAT
GTGCCTGCTGCACGAGGCATTCAAATGGTGCATAGAGTTCCTCGAAACATTGGCGAAGAGGGTTCTGAAGTTATTGGAGCAAGGCACACTGAGAATATGA
GGGGATTCCCTGATGATGGTACAGAGCAGGCATTTAGGCGACCCCAACCTTCATACGAAGGGTTAGATGGTCATTTTGTCCAAGGGACTAGGAACTATTC
ATCTGTTCATAGAAGGGCTCTTCCCCAATTTCGTTCAAAATCTCCAATTAGATCTCGTTCTCCTGGTCCATGGTCATCTGCCAGAAGAAGATCTCCAGAT
GGGTTTGGTGGGACTTCAGAATTGTCTAATCGAAGATCACCAATTTACAGTATGGGGAGGATCAGATCACCTGACCATCCTGGTTTTCCCAGGGAAATGG
TTGTTAGGAGGCACGGTTCTCCACCATTCTTATCTCGACCTCCTGATACCAGGGAAACAGATCCTGGTCATTCAAGGTCTATTATCTCTAATAGGGGCCA
AACAGGAAGGGTTTTTCTCAGAAACAGTAGGAGGTTTGGTATTACAGATCCTCGAGAAAGGGCGGACAGTGATGAGTTCTTTGGAGGGCCCATACACTCT
GGTCGATTTCATGACCTTGGTGGTGATGGGAATGTGGAAGATAGAAGAAGATTTAGCGAGAGGCGGGGACCTGTCCGTTCATTTAAGCCTCCTTTTAATG
GTGCTGGTAGTGAAAATTTTCATCTTAATCCAGAGGATGGCCCCCGACCTTTTAGGTTCTTTCCGGAAGATAATCCAGAGTTCCATGAAAGAACTAATTT
GAGGGAAAGGGAGTTTGATGGACGGATTAGGAACCGTCCTGGAAATGCACCCAGAAGACCCCGAGGCATTGAAGAGCAGGAAGGAAATTACAGGCATGGT
AGGCAGGTATTGTATGATGATGGGTTTGATATTTCACGAATGAAAAGGAAAAGGTTTTAA
|
||||||||||
|
AA sequence
|
>Potri.008G047600.5 pacid=42806484 polypeptide=Potri.008G047600.5.p locus=Potri.008G047600 ID=Potri.008G047600.5.v4.1 annot-version=v4.1
MPVSGNEETGVKPHAQQSSLNIAGVPIKKRRFIWPPSPPPEEQSVPLLGNDSAQKEPGSTSKESSPSNSSVAASSDLSDPFKNSVAEENKNRLDSIVQMN
AENCSGVKVVAQNLATHSDSLAKFGKQEKPVVEEKSANTLLISAKTELNLESNKGPGLNVGKEICGQQILEGKCKSEMPIASVTSQFSLGLKEHDVSSLE
CYSNDGSQINENVGAVSLNLSLSEGETGVLHKMDNILATDSTDVFANRSNWDLNTTMDTWDGSSSDEHAAQETADGWNRVGVKCDITTGIVGAGMSNGRQ
LLDSSECKSSFPQTFSDCAREYTSEDSLHLRLSPSFPSFNLSQEHSSSSANKESCIIPNISLPGSLLSAGNATVANCRGIKSEPFDGSLKHDLRGAKVNP
FDFFVKRELVEKGSLETSKSSASGSLKLVGHGFIKPEPFHDGKPETPRMVGGGSIQPDKQVLQSQDTGEQSPCSASKIVLQVQDTTGQPSCSTDNQVREG
QDILAKPTSSTDLFISGNASDRLEYTTCVEGALLRNAMPKEAPESAGQVSSEMVSMPVGHSGEELDASVKIDTAITMDRNGDAPEQCELKITEEVPAGSH
GNGEASVTDEEKINLSGDMIEEDSYGSGYESDGNTMSMDIDEEQREHKYEDGEVQDPHLQAAEECQKCEEKDVSHGNSEHEKANSGLAGDDHYISSLVEE
NDSKIELSENNEVTVKECITRTIEDADNASVKESPTVEMSTCGAEQERETTIIQRKSLDLSGKKDCPVGQGTELSSGQDITAGQGVLVSVEQGSDENIKT
NYMEKNELPELEASLNGGDMAKDVSSSRSRIINLPRASNSSSPGKTRSISGRPFSSYQERLPDGPLEGGKLHPQGRDEIYIDGPRRFSRDRHQEHFPRNS
RMNFVRGRGRISSRVDTLRGDRDSERNYASEFYNGSSDFAVRRHKYASAAAEADSESINYNIAPDGSFVGTARGGRKLLDDETPVFRNVPSRRRSPEGRD
VPAARGIQMVHRVPRNIGEEGSEVIGARHTENMRGFPDDGTEQAFRRPQPSYEGLDGHFVQGTRNYSSVHRRALPQFRSKSPIRSRSPGPWSSARRRSPD
GFGGTSELSNRRSPIYSMGRIRSPDHPGFPREMVVRRHGSPPFLSRPPDTRETDPGHSRSIISNRGQTGRVFLRNSRRFGITDPRERADSDEFFGGPIHS
GRFHDLGGDGNVEDRRRFSERRGPVRSFKPPFNGAGSENFHLNPEDGPRPFRFFPEDNPEFHERTNLREREFDGRIRNRPGNAPRRPRGIEEQEGNYRHG
RQVLYDDGFDISRMKRKRF
|
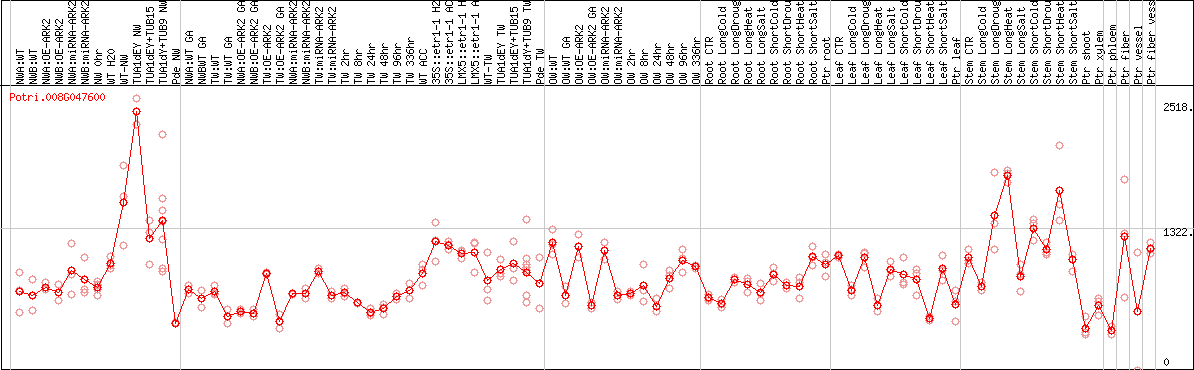
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G047600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
