External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G28670 276 / 3e-90
ESB1
ENHANCED SUBERIN 1, Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
AT1G07730 268 / 5e-88
Disease resistance-responsive (dirigent-like protein) family protein (.2)
AT2G39430 197 / 6e-61
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT3G55230 176 / 3e-53
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT3G24020 149 / 3e-43
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT4G13580 145 / 4e-42
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT2G28671 99 / 6e-24
unknown protein
AT2G21100 64 / 7e-12
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G42510 63 / 9e-12
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT1G65870 63 / 9e-12
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G211800
389 / 2e-136
AT2G28670 257 / 6e-83
ENHANCED SUBERIN 1, Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Potri.010G211900
211 / 1e-65
AT2G39430 249 / 3e-80
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.008G049100
201 / 6e-62
AT2G39430 251 / 6e-81
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G054100
166 / 1e-49
AT4G13580 308 / 3e-106
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G054000
156 / 4e-46
AT3G24020 352 / 5e-124
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.003G174300
155 / 7e-46
AT3G24020 315 / 2e-109
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.010G212000
103 / 4e-26
AT1G07730 103 / 1e-25
Disease resistance-responsive (dirigent-like protein) family protein (.2)
Potri.003G216300
63 / 2e-11
AT1G55210 221 / 5e-74
Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Potri.003G216400
61 / 6e-11
AT1G58170 225 / 1e-75
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006317
284 / 1e-95
AT2G28670 271 / 1e-89
ENHANCED SUBERIN 1, Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Lus10029585
181 / 2e-57
AT2G28670 166 / 6e-53
ENHANCED SUBERIN 1, Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Lus10030318
153 / 8e-45
AT3G55230 276 / 2e-93
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10011767
150 / 8e-44
AT3G24020 339 / 7e-119
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10023689
150 / 1e-43
AT3G24020 336 / 1e-117
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10003301
142 / 4e-38
AT3G55230 254 / 2e-80
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10025063
64 / 1e-11
AT2G21100 183 / 4e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10021084
62 / 2e-11
AT5G42500 164 / 2e-51
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10034482
63 / 3e-11
AT2G21100 182 / 8e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10034479
62 / 3e-11
AT2G21100 189 / 9e-62
Disease resistance-responsive (dirigent-like protein) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0650
AOC_barrel
PF03018
Dirigent
Dirigent-like protein
Representative CDS sequence
>Potri.008G049200.1 pacid=42806369 polypeptide=Potri.008G049200.1.p locus=Potri.008G049200 ID=Potri.008G049200.1.v4.1 annot-version=v4.1
ATGGAAATTCACAAAATTCTCTCTACCACAAACAAGGCCATAGCCTGTCTTCTGTTCCTCACTCTTACCATTGTATTTACTACCTCAGCTAGAACCCTGG
ATGAGCAGCCAGAGGTACCTGTTGTTCCCCCGGAGTCAGACAATGCAGTCGATGATACTGTCACCAGTGTTCCACCACTACCTACACCATTAGCACCCAC
ACAGCCTAATCCAACTGGGGCTGCAAGCACTAGTGCCACGATAAGTAATCCTGGTCATACCTTAAGCTTCTTCATGCATGACATTCTTGGTGGAACAAAC
CCCTCGGCTAGAGCCGTGACTGGCATAGTCAGCAACCCGGCAGTCTCTGGCCAAGTCCCATTTGCCAAACCTAATGGTGCAAACCTCCCAATAAATAATG
GCGTGCCCCAAAATAGCAACAATAACGGCCTTATAAACAACAACAACCTACCTTTCCTCACTGGCCTTGGTGGCACCACACAGCCTGTGGTTCAAAACAA
TGGTAATAACTTCAACAATGCATTTAACTTGCCACTGAGCAATGGAGGCAATCTTCCATCTGGTTCCACTCTTCAACAGCTCATGTTTGGAACCATCACT
GTGATCGACGACGAACTAACCGAAGGGCATGATTTACGGTCAAGTTTTGTAGGTAAGGCACAAGGGTTCTATGTGGCTAGTTCTCTGGATGGTACCAGCC
AAACTATGGCATTTACAGCTATGTTTCAGAGTGGTCATTATGCTGATAGCCTTAGCTTTTTCGGGGTTCTCAGAACAGGAGTCTCCGAGTCTCAGCTTGC
TATCATGGGTGGAACTGGAAAGTATGTCAATGCTCAGGGTTATGCTATTATTAAGATCATCCCTTCTACCAATCAGAATACGACCGATGGTGTTGAGACC
TTGCTGGAGTTCGTTGTTTATGTTAGCTACTAA
AA sequence
>Potri.008G049200.1 pacid=42806369 polypeptide=Potri.008G049200.1.p locus=Potri.008G049200 ID=Potri.008G049200.1.v4.1 annot-version=v4.1
MEIHKILSTTNKAIACLLFLTLTIVFTTSARTLDEQPEVPVVPPESDNAVDDTVTSVPPLPTPLAPTQPNPTGAASTSATISNPGHTLSFFMHDILGGTN
PSARAVTGIVSNPAVSGQVPFAKPNGANLPINNGVPQNSNNNGLINNNNLPFLTGLGGTTQPVVQNNGNNFNNAFNLPLSNGGNLPSGSTLQQLMFGTIT
VIDDELTEGHDLRSSFVGKAQGFYVASSLDGTSQTMAFTAMFQSGHYADSLSFFGVLRTGVSESQLAIMGGTGKYVNAQGYAIIKIIPSTNQNTTDGVET
LLEFVVYVSY
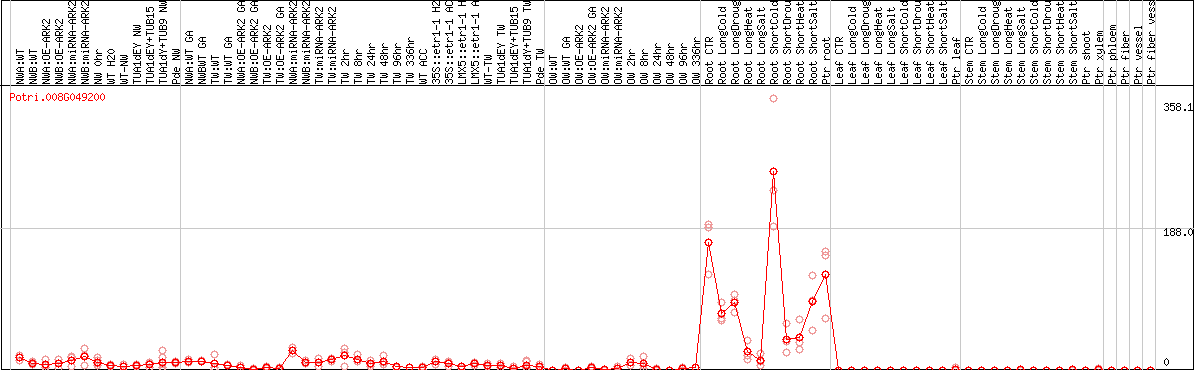
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G049200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.