External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G75270 340 / 2e-120
DHAR2
dehydroascorbate reductase 2 (.1)
AT1G19570 320 / 2e-112
DHAR5, DHAR1, ATDHAR1
DEHYDROASCORBATE REDUCTASE 5, dehydroascorbate reductase (.1.2)
AT5G16710 297 / 1e-102
DHAR3
dehydroascorbate reductase 1 (.1)
AT1G19550 188 / 2e-61
Glutathione S-transferase family protein (.1)
AT5G02790 50 / 2e-07
GSTL3
Glutathione transferase L3, Glutathione S-transferase family protein (.1)
AT5G42150 42 / 9e-05
Glutathione S-transferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G125100
291 / 3e-100
AT5G16710 389 / 4e-138
dehydroascorbate reductase 1 (.1)
Potri.010G211600
286 / 7e-100
AT1G75270 253 / 5e-87
dehydroascorbate reductase 2 (.1)
Potri.004G080400
56 / 1e-09
AT1G78380 231 / 5e-77
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G114000
49 / 6e-07
AT1G17180 265 / 2e-90
glutathione S-transferase TAU 25 (.1)
Potri.011G113125
47 / 1e-06
AT1G78380 259 / 2e-88
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G113000
47 / 2e-06
AT1G78380 259 / 3e-88
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.004G080700
46 / 4e-06
AT1G78380 217 / 6e-72
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.011G113400
43 / 4e-05
AT1G17170 231 / 6e-78
Arabidopsis thaliana Glutathione S-transferase \(class tau\) 24, glutathione S-transferase TAU 24 (.1)
Potri.010G035500
40 / 0.0004
AT1G10370 301 / 2e-104
GLUTATHIONE S-TRANSFERASE U17, GLUTATHIONE S-TRANSFERASE 30B, GLUTATHIONE S-TRANSFERASE 30, EARLY-RESPONSIVE TO DEHYDRATION 9, GLUTATHIONE S-TRANSFERASE TAU 17, Glutathione S-transferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023441
334 / 5e-118
AT1G75270 327 / 3e-115
dehydroascorbate reductase 2 (.1)
Lus10040320
331 / 1e-112
AT1G75270 320 / 4e-108
dehydroascorbate reductase 2 (.1)
Lus10023442
313 / 9e-110
AT1G75270 317 / 5e-111
dehydroascorbate reductase 2 (.1)
Lus10009135
294 / 5e-100
AT5G16710 372 / 5e-130
dehydroascorbate reductase 1 (.1)
Lus10028510
280 / 2e-93
AT5G16710 352 / 2e-120
dehydroascorbate reductase 1 (.1)
Lus10019895
59 / 3e-10
AT5G12130 399 / 6e-135
PIGMENT DEFECTIVE 149, TELLURITE RESISTANCE C, integral membrane TerC family protein (.1)
Lus10030020
48 / 1e-06
AT1G17180 238 / 1e-79
glutathione S-transferase TAU 25 (.1)
Lus10007897
45 / 9e-06
AT2G29420 187 / 1e-59
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10020519
45 / 1e-05
AT2G02390 283 / 2e-97
GLUTATHIONE S-TRANSFERASE 18, glutathione S-transferase zeta 1 (.1.2.3)
Lus10005367
45 / 1e-05
AT2G02390 284 / 4e-98
GLUTATHIONE S-TRANSFERASE 18, glutathione S-transferase zeta 1 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0497
GST_C
PF13410
GST_C_2
Glutathione S-transferase, C-terminal domain
CL0172
Thioredoxin
PF13417
GST_N_3
Glutathione S-transferase, N-terminal domain
Representative CDS sequence
>Potri.008G049300.2 pacid=42806565 polypeptide=Potri.008G049300.2.p locus=Potri.008G049300 ID=Potri.008G049300.2.v4.1 annot-version=v4.1
ATGGCTTTAGAGATCTGTGTCAAAGCTGCTGTTGGTGCCCCTAATATTCTCGGAGATTGCCCATTTTGCCAGAGAGTTCTGTTGAGTCTGGAAGAGAAGA
AAATTCCATACAAGTCTCATCTCATCAATCTGGGTGATAAACCTCAGTGGTTTTTGGAGATAAGCCCGGAAGGAAAGGTCCCAGTGGTGAAAATTGATGA
CAAATGGGTCGCAGATTCTGACGTGATTGTTGGTATCTTAGAGGAGAAAAACCCTGAACCCCCTCTTGCCACTCCTCCGGAATTTGCCTCTGTGGGTTCG
AAGATATTCCCATCTTTTGTTAAGTTCTTGAAGAGCAAGGATCCCAATGATGGAACTGAGCAGGCTTTGCTTGAGGAATTGAAGGCATTGGATGGGCATC
TCAAGGGTCCTTTCATTGCTGGGGAAAAGATCACAGCAGTGGATTTGAGCTTGGCTCCGAAGCTTTACCATCTTGAGGTTGCTCTTGGCCATTTCAAGAA
CTGGACTATCCCTGACAATTTGACACATGTCCTCAACTACATTAAGTTGTTGTTCTCCCGTGAATCTTTCAAGAAGACAAGAGCTGCCGAGGAACATGTC
ATTGCTGGATGGGAGCCAAAGGTCAACGCATGA
AA sequence
>Potri.008G049300.2 pacid=42806565 polypeptide=Potri.008G049300.2.p locus=Potri.008G049300 ID=Potri.008G049300.2.v4.1 annot-version=v4.1
MALEICVKAAVGAPNILGDCPFCQRVLLSLEEKKIPYKSHLINLGDKPQWFLEISPEGKVPVVKIDDKWVADSDVIVGILEEKNPEPPLATPPEFASVGS
KIFPSFVKFLKSKDPNDGTEQALLEELKALDGHLKGPFIAGEKITAVDLSLAPKLYHLEVALGHFKNWTIPDNLTHVLNYIKLLFSRESFKKTRAAEEHV
IAGWEPKVNA
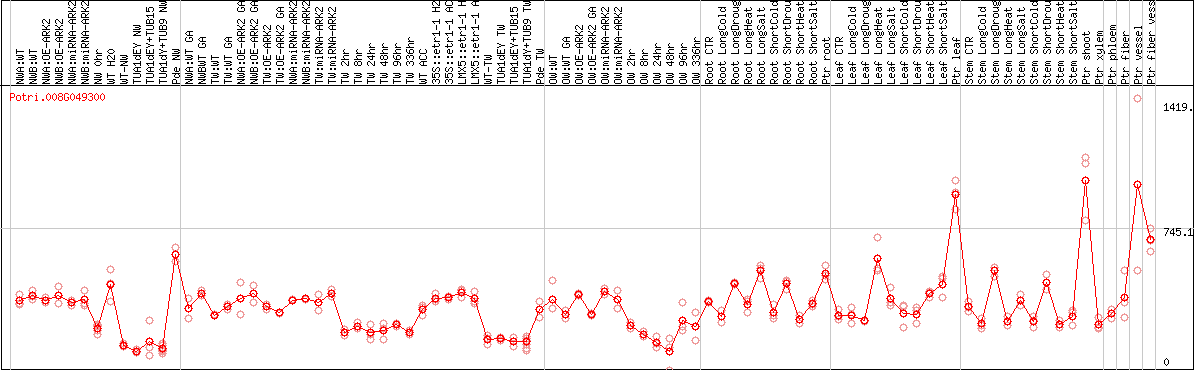
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G049300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.