Potri.008G050400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G050400.1 pacid=42806837 polypeptide=Potri.008G050400.1.p locus=Potri.008G050400 ID=Potri.008G050400.1.v4.1 annot-version=v4.1
ATGATGATCTCCAGGGGATTATTCGGATGGTCCCCGCCGCATATACAACCGTTGACGCCGGTCTCCGAGGTTTCCGAGCCGCCGGAGAGTCCGTCTCCGT
ACTTAGATAACAGTGCGGAGGCGGCCGCGGCTGCTGCTGCGGCGGCTCAAGCGGAAGCAGAGGAGGAGATCGAGGAGGCTGAGGAGATGGAGCCGCCGCC
AGCTGCTGTTCCTTTCTCCGGGTTGTTTGCTTGTGCTGATAGGTTAGATTGGGGGCTCATGATCGTTGGGTCGCTTGCAGCGGCTGCGCACGGGACTGCT
CTTGTTGTTTACTTGCATTACTTTGGGAAGATTATTGGGGTTTTGAGTATTAAGCCGGAGGAGCGATTTGACAGGTTTACTGATCTTGCTATGCACATTG
TTTATTTAGCCGTGGGAGTTTTTGCTGCTGGTTGGATTGAGGTTTCCTGCTGGATTTTAACTGGGGAACGTCAGACCGCGGTCATCAGGTCAAAGTATGT
TCAGGTATTGCTCAATCAGGACATGAGCTTTTTCGATACATATGGGAATAACGGGGATATTGTTAGCCAAGTATTGAGTGACGTGCTCCTCATTCAATCT
GCATTGAGTGAAAAGGTTGGAAATTATATTCATAATATGGCTACATTTTTCAGTGGTCTTGCTATTGGATTTGTCAACTGCTGGCAAATTGCACTAATAA
CATTAGCCACTGGCCCATTCATTGTTGCTGCTGGAGGTATCTCAAATATATTTCTTCATAGGCTTGCTGAGAGCATTCAAGATGCATATGCTGAAGCTGC
TAGCATTGCTGAACAGGCAGTCTCTTATAGTAGGACCTTGTATGCATTCACAAATGAAACTCTGGCAAAGTATTCATATGCAACATCACTACAAGCAACT
TTGAGATATGGTATTTTGATTAGCCTTGTTCAAGGACTTGGGCTTGGTTTTACGTATGGGCTTGCAATATGTTCTTGTGCCTTGCAACTTTGGGTTGGAA
GGTTTCTGGTTACAAGTCATAAAGCTCATGGTGGTGAAATAGTAACAGCCCTGTTTGCTATAATTTTAAGTGGCCTTGGGCTGAATCAAGCTGCAACCAA
TTTCTATTCATTTGACCAAGGACGAATTGCTGCATATAGACTCTTTGAGATGATAAGTCGGTCATCCTCCACAGTTAATCAGGATGGGAACAACCTAGTG
GCAGTGCAAGGAAATATCGAGTTTCGAAATGTTTATTTTAGCTATCTGTCTCGTCCTGAGATTCCCATCTTGAGTGGATTTTACCTCACTGTACCAGCTA
AAAAGACAGTGGCTCTTGTTGGAAGAAATGGTTCTGGAAAGAGCAGCATTATCCCACTCATGGAGCGCTTTTATGATCCTAACTTAGGAGAAGTTCTTCT
TGATGGAGAGAATATTAAAAACCTGAAACTGGAATGGCTCAGAAGCCAAATAGGGCTGGTCACCCAGGAACCTGCCTTGCTGAGTTTGAGTATAAGAGAT
AATATTGTATATGGACGAGATGCTACTTTGGATCAGATTGAGGAAGCTGCCAAAATAGCACATGCACACACATTTATCAGTTCACTTGAAAAAGGCTACG
AGACACAGGTCGGTAGGGCTGGTTTAGCATTGACTGAGGAGCAGAAAATTAAACTTTCTATTGCCAGGGCAGTACTTTTAAATCCTACAATTCTTCTGCT
TGATGAGGTTACTGGTGGACTTGATTTTGAAGCTGAGAGAGCAGTTCAAGAGGCACTAGATCTTCTCATGTTGGGACGTTCAACCATTATAATAGCTAGA
CGTCTTAGTCTAATTAGGAATGCTGATTATATAGCTGTAATGGAGGAGGGTCAGCTTGTTGAAATGGGTACACATGACGAATTAATAACCTTAAATGGTC
TATATGCAGAGCTTCTCAAATGCGAAGAAGCAGCAAAACTTCCAAGGAGGATGCCAGTTAGAAACTACAAGGAGACTGCTGCTTTCCAGGTTGAAAAGGA
TCCTTCCACAGGCCACAGCTATCAAGAACCATCCTCCCCTAAAATAGCCAGGTCACCATCACTTCAGAGAGCTCCTGGTATATTTCGGCCTCCAGATAGC
ATGTTTAATTCACAAGAATCACCAAAAGTTCTCAGCCCACCACCAGAGAAGATGATGGAAAACGGTCTTCCCTTGGATGGAGCTGATAAGGAACCATCAA
TAAGAAGGCAGGATAGTTTTGAAATGAGACTACCTGAATTACCCAAGATTGATGTCCAGTCTGCACATAGACAGGCATCAAATGGTTCAGACCCTGAATC
CCCTGTTTCTCCACTTTTGACATCTGATCCTAAAAATGAGCGTTCCCACTCACAGACTTTCAGTCGACCCCACAGTCACTCAGATGATGTTCCAATTAAA
GTGAAGGAATCAAAGGATACAAAGCATCTGGAAGAACCATCATTTTGGAGACTAGCAGAGCTTAGTTTGGCAGAGTGGCTTTATGCTGTTTTAGGAAGCA
TTGGTGCTGCTATTTTTGGTTCTTTCAATCCACTACTTGCTTATGTTATTTCACTGATAGTGACTGCTTATTATGGACGAGACATGCAGCAGGATGTAAA
CAGGTGGTGCTTGATAATTGCCATCATGGGTATGGTGACTGTTGTTGCCAATTTCTTGCAGCACTTCTATTTTGGTATCATGGGAGAGAAAATGACTGAA
CGTGTCCGCAGAATGATGTTTTCAGCAATGCTGCGGAATGAAGTCGGATGGTTCGATGAAGAGGATAATGGTGCTGACACATTATCCATGCGCTTGGCAA
ATGATGCTACTTTTGTAAGAGCTGCATTCAGCAACCGACTTTCAATATTTATACAGGACAGTGCTGCAGTTATTGTGGCTGTTGTCATTGGGGTGCTGCT
TCAGTGGCGATTGGCTCTTGTAGCACTGGCAACCTTGCCAGTTCTCACTGTCTCTGCAATTGCACAGAAGTTGTGGCTTGCTGGATTTTCCAGGGGCATC
CAGGAGATGCACAGAAAGGCATCATTGGTCCTTGAGGATTCTGTTAGAAACATTTATACAGTTGTGGCATTCTGTGCGGGTAATAAAGTAATGGAGCTCT
ACAGATTGCAACTGCAGAAAATATTTAAGCAGAGTTTTTTCCTTGGAATGGCCATTGGTTTTGGATTTGGTTTTTCTCAGTTTCTTCTTTTCGCCTGCAA
TGCCCTTCTTCTCTGGTACACTGCATACTCTGTAAAGAATCACAATGTGAATCTTCATACGGCTCTCAAGGAGTATATGGTTTTTTCTTTTGCAACATTT
GCACTGGTGGAGCCTTTTGGATTGGCTCCCTACATTCTTAAGCGGCGAAAATCTCTTATTTCAGTATTTGAAATTATAGATCGAGAACCTAAGATTGACC
CAGATGATAACTCAGCACTGAAGCCTCCTAATGTTTATGGAAGCATTGAGTTGAAGAATGTGGATTTCTGTTATCCAACTCGTCCAGAAATGTTGGTATT
GAGCAATTTCAGTCTCAAGGTGAATGGAGGACAAACTGTAGCTGTGGTGGGAGTTTCAGGGTCTGGAAAAAGCACTATAATTTCTTTAATTGAAAGATTT
TATGATCCAGTTGCCGGTCAGGTCCTACTGGATGGCCGGGATTTAAAACTTTATAATTTGAGATGGTTGAGGAACCATCTTGGTCTTGTTCAGCAGGAAC
CGATTATCTTCTCAACAACTATAAGAGAAAATATTATATATGCTAGGCACAATGCCAGCGAGGCTGAGATGAAAGAGGCTGCAAGAATTGCTAATGCTCA
CCATTTCATCAGTAGTCTGCCTCATGGTTATGACACACATGTGGGAATGAGGGGCGTGGACCTTACCCCAGGGCAGAAGCAGAGAATTGCAATTGCTCGA
GTGGTGTTGAAGAATGCACCCATCTTGTTATTGGATGAAGCCAGCTCATCCATCGAATCTGAATCAAGTCGAGTGGTACAAGAGGCACTAGATACTCTTA
TTATGGGAAACAAAACTACCATTTTGATAGCTCATAGAACTGCAATGATGAGGCATGTTGACAACATTGTGGTTCTTAATGGAGGACGAATTGTCGAGGA
AGGGGCCCATGATTCTTTGATGGCAAAGAATGGTTTGTATGTCCGTCTGATGCAACCCCACTTTGGAAAGGGTTTGCGACAACATCGGTTGATTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G050400.1 pacid=42806837 polypeptide=Potri.008G050400.1.p locus=Potri.008G050400 ID=Potri.008G050400.1.v4.1 annot-version=v4.1
MMISRGLFGWSPPHIQPLTPVSEVSEPPESPSPYLDNSAEAAAAAAAAAQAEAEEEIEEAEEMEPPPAAVPFSGLFACADRLDWGLMIVGSLAAAAHGTA
LVVYLHYFGKIIGVLSIKPEERFDRFTDLAMHIVYLAVGVFAAGWIEVSCWILTGERQTAVIRSKYVQVLLNQDMSFFDTYGNNGDIVSQVLSDVLLIQS
ALSEKVGNYIHNMATFFSGLAIGFVNCWQIALITLATGPFIVAAGGISNIFLHRLAESIQDAYAEAASIAEQAVSYSRTLYAFTNETLAKYSYATSLQAT
LRYGILISLVQGLGLGFTYGLAICSCALQLWVGRFLVTSHKAHGGEIVTALFAIILSGLGLNQAATNFYSFDQGRIAAYRLFEMISRSSSTVNQDGNNLV
AVQGNIEFRNVYFSYLSRPEIPILSGFYLTVPAKKTVALVGRNGSGKSSIIPLMERFYDPNLGEVLLDGENIKNLKLEWLRSQIGLVTQEPALLSLSIRD
NIVYGRDATLDQIEEAAKIAHAHTFISSLEKGYETQVGRAGLALTEEQKIKLSIARAVLLNPTILLLDEVTGGLDFEAERAVQEALDLLMLGRSTIIIAR
RLSLIRNADYIAVMEEGQLVEMGTHDELITLNGLYAELLKCEEAAKLPRRMPVRNYKETAAFQVEKDPSTGHSYQEPSSPKIARSPSLQRAPGIFRPPDS
MFNSQESPKVLSPPPEKMMENGLPLDGADKEPSIRRQDSFEMRLPELPKIDVQSAHRQASNGSDPESPVSPLLTSDPKNERSHSQTFSRPHSHSDDVPIK
VKESKDTKHLEEPSFWRLAELSLAEWLYAVLGSIGAAIFGSFNPLLAYVISLIVTAYYGRDMQQDVNRWCLIIAIMGMVTVVANFLQHFYFGIMGEKMTE
RVRRMMFSAMLRNEVGWFDEEDNGADTLSMRLANDATFVRAAFSNRLSIFIQDSAAVIVAVVIGVLLQWRLALVALATLPVLTVSAIAQKLWLAGFSRGI
QEMHRKASLVLEDSVRNIYTVVAFCAGNKVMELYRLQLQKIFKQSFFLGMAIGFGFGFSQFLLFACNALLLWYTAYSVKNHNVNLHTALKEYMVFSFATF
ALVEPFGLAPYILKRRKSLISVFEIIDREPKIDPDDNSALKPPNVYGSIELKNVDFCYPTRPEMLVLSNFSLKVNGGQTVAVVGVSGSGKSTIISLIERF
YDPVAGQVLLDGRDLKLYNLRWLRNHLGLVQQEPIIFSTTIRENIIYARHNASEAEMKEAARIANAHHFISSLPHGYDTHVGMRGVDLTPGQKQRIAIAR
VVLKNAPILLLDEASSSIESESSRVVQEALDTLIMGNKTTILIAHRTAMMRHVDNIVVLNGGRIVEEGAHDSLMAKNGLYVRLMQPHFGKGLRQHRLI
|
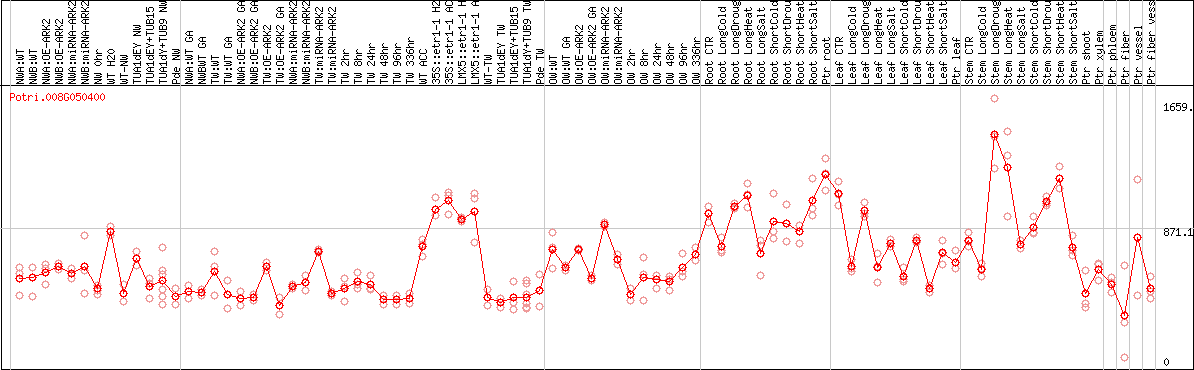
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G050400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
