Potri.008G052500 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.008G052500.1 pacid=42805836 polypeptide=Potri.008G052500.1.p locus=Potri.008G052500 ID=Potri.008G052500.1.v4.1 annot-version=v4.1
ATGGCGTCAAATAACCCTAGTCAACCAGAGTTCGCGACCAAAACAAGAGAAGAAGGAGAGGTCTCTTCTTCATCCAACGACGACCAAAATCCTGTTTGCG
CTTCTGCACTGTTTGCTGATGCTTTTAACACACCTGCTTCCGCAAGGACCATTCCGTTTCATCCGATGAACAAATTCACTTTATCCAATCGGGCTGGCAA
GGCCATTTTTAGCACTAATCCCGCAAGATGTGTTGATCCGAATCTCCAAACATCACAACCAAATAATAACAAGAGCTTTGAGAAGAATAGGGTGCCACAC
ATATCTGCTAATCCTGGAAAGTGTGCATCTTCAGGGGCTAATGATAACTTAGTGATAAGATTTTCAGATGATGAAAGTGGGAGTGAATCTGAAGATAGAG
ATGACAAGCCTTTAAAAACTAAACCCAATACAACTGTGGTTAATGGAAATGGAAGGCTGCCCCCCATTTTACCGGCAAAATCAAGCATGTCGCTGCAGAC
TTCTAGAAATGTGAACAGTATTCCCAAGAAATCATCAATGAGTCGCACATTTAACTCTTCAATGACAAAGATAAATGGAGTTGCCAATTCCAAGGGTGCT
GACTCCTCGTCTGTTGGGCAAGGTTCTCAAGTAAAAAATATTAATTCCATCAAAAGAAATCTAGCAAGCCAAGAGCATGGTATTGAGCAAGGTGTGGATT
TGAACAGCACTAAGGTTCAGGACTTGAGGCAGCAGATTGCACTTCGGGAAAGGGAACTGAAGCTTAAGGCAGCTGCACAAAACAAGGAATCAGGTTCAGT
TTCCGACAAGTGTATGAACATTTCCAGTAGTGTGACCAGGAAATCTAATGCAGCTTCTTCTGAAGTTGGGCAATTAGCACCAAAAGAACCAGACAGGAAA
CGCATAAAACCTGACGGATCTTATTCTAAACACTTAAATTCAGATGGCCAACAAAAGATGCTTGTAGAGAAATCTAATTTACCGTCAAAAGATCAAGCGC
TGGAGAACAGTAGTTTGCAGGATAGAAATATGGGTAATTGTAGCAAGAAAGAAAGACCGACAAAAAGAACAGAGTCGAGTGTAGTTAAATGGGAGAGGCA
AGATAGGAGGGTGGATATTTCATCAGCAAAGCTACCTGCTTCTCACATCAATGATAATTCCAGTCAGCCTGACACGAGCAGAATGCAGATGGATCCTTGT
GTGGTATTGAACCAGACCCCACTGCTTACTAATGCAAATGCTAGTACTTTATCGAAGAAGAGAAAGAGTGTTGACTTGAATCCTGTAAAGAATTGCGGAA
CCCAGCCACCAGCTTGTTTGTTAAAGACATCAACTAGTGGACAGAATTTAATAAACAGCTGTGAGCACTTGCAAGGCATATCTGGTGATAAGCTGTCCTG
TCAAGCATCTCTGAATTTAAACCCTTGGAATTGTCTGGGAACTGTAAATGTGGCAGAACATAACAGCATAGATATTCAGTTGTTAGTTGAAATGGAGGAA
TCCTTAGACAGGGAGCTGGAAGAAGAACAAGAGCACAGACACAAATGTGAAATTGAAGAAAGAAATGCTCTTAAAGCTTATCGAAAAGCACAGAGAGCTT
TGATTGAGGCTAATTCTAGATGTACAGAACTTTATCGCAAGAGGGAACTGCACTCAGCTCACTTTCGATCTCTCATTGTGAATGATTCCAGTTTGTTTTT
TCCCTCCAGGCAGGATGAACATGTTGGCATTGGGATGGACCGTGAAAATAATGTATCTAGAAATGTGGATTTAATACCCTCATCAAGCGATCAGATGCAG
CCTGAGTATGATGGTTGTAATCAGCCAGGGTATGATTCAGTCACTGGTGCTCCCTCAAATTCACTTTATCAGCATGTAAATGGCCATAGTTTGGGATCCG
AGCCATGCAGTGAACCTGATGCTAGTACATCAGAGCCATTGCCCCGCAACAGCCTGATTGCTGCAAATGGAGTAAGCTCTCAATCCAATGATTCTAATAT
TTCAGCTGGGGAAGATGAAGAGACGTTTCCATTGGACCATGAGACTGATCAACCTATCTTCAAAATTCAGCAAAGAGATCAAAATTCTGTGGGAAGGGAA
AGCCACACAGATTGTCATCCAAACAAAGATTTTTATGTTGATGGCCCTCAGGATTCTTTGATTCTTGAAGCAAAATTAAGATCCAAACTATTTGCTCGTC
TACCAATCAGAACCTTTTCAAAGAATGGTGGTTCATCCAACATGGAGCCTGCAGATGAACCAGGGATTGAAATTGACAACAGAAGTGAAAGAACCCAGGG
AAGCAATGTCAGTATACCATTGTCAGAGACAGAAAAGGATCGAGATTATGACCTTGAAGGCAATAATAAGCCTGAAAGAAGTATATCTGAGCTTCCTGTC
CAGATTCAGAACCATGAAAAGAATTTTCATTCTGCTGCTGATTCCAAGGACGATTCCACTGGAGGTCATCAGCTGACAACTTCTGTCATATCTTCACCTC
TTTTAGTATTGAGGAGTGCATTTGCTCAAATGAAAGCTATGCACCCAATGACTTTGATAGAATCACAATGTAGAAAAAACCAACAAAATGATACCTGTGG
CGATTTCATTGTAGAGGATGGTTTTATGGACACTGAGGAAATCCAGTGTGATAACGTGATAGCAAAGTCAAAGGAGGAGATTATAAGGGGTATGTGTGGA
ACTGAAATTGGGACTTTTACCCACAATGTGGCGGTTGACCCATTTTGGCCACTATGCATGTATGAGCTTCGAGGAAAATGCAACAATGATGAATGTCCTT
GGCAACATGTTAGGGACTTCTCTGATCAAAATTTGCATCCAAATCAGCATGATGATTCTGATAGTGCTGATTGTCAGGTTGGATTAACACTGCATGAACA
AAAATGTAAAGGTGGAGCAAAACTTTCCAAGTGTCACAGTGTATTGAATCCACCAACTTATCTTGTTGGCTTGGATGTTTTAAAATCCGATTCATATAAA
TCTGTCATTGCACGGAGAAATGGGCAATGCTGGCAAATACAGTTCAGTCTTTGCTTAGCTTTATCAAGTTTCTTTCAAAAAGATTTGCTTGCTGATCAGC
TTTCTATCCGTGCTGATGATGGCCGTATCGAGGTCCATGGGAGTTGGAATAGACAGACATCATACTTTCAGAGTAGAGAGAACACAGTGAATCATCTCAA
TCAAGCATTAGCCAGCAGTTTGCAGTCCCTTGAAATGGCTCTTCTTTTTCTCTGTCAGGAGGTTTACAAGCTGGAGGGTATGAAGAAGCCTCTCTCCATG
CTGTCACGAGCAATTGAGGCTGATCCAACATCTGAAGCTCTGTGGATGATGTATCTGCTCATTTACTACAGCAACATTGAGTCAATTGGGAAGGATGACA
TGTTCTCCTATGCGGTTAAAAACAACGAAAGATCTTATGGACTTTGGCTCATGTACATCAACAGTCGTATACATCTTGATGATCGAATGGTAGCATATAA
TGCTGCTCTCACAGCGCTTTGTCGCCAGGCATCTGCTTTTGATAAGGGCAATATGTATGCTAGTGCATGCATCTTAGATTTGTTCTTACAGATGATGGAT
TGTTTGTGCATGTCTGGGAATGTTGGCAAGGCTATTCAGAAAATCCAAGGACTCTTCCCTGTGGCAGCTAACTCAGATGAGCCTCACTTTCTGTTGCTCT
CTGATATCCTCGCATGCTTAACAAATTCTGACAAATATATCTTCTGGGTTTGTTGTGTGTACTTAGTTATTTACAGGAAGTTACCTGATGCTATTGTACA
ATGTTTTGAATGTGACAAAGAACTCCTTGCAATTGAATGGCCTTATGTTCAGTTACCAAATGAAGAGAAGCAGAGGGCTGTTAAGCTAGTTGAGATGGCA
GTGGATTCTGTTGAAATGTCTGTCAACAGTGAATCACTTGAAAGTGACAAAAATGGCAGAATGGCCCAACAGTTTGCTCTCAGCCATATTAGGTGTACAC
TAGTTTTTGATGGCTTAGCTTGCTGTCAGAATTTGTTGGGCAAGTATACAAAGTTGTACCCGTCCTGTGTAGAACTTGTTCTCCTGTCAGCAAGGCTTAA
AAAAAATGGCTTGGGCAGTGTGAGCTTTGAAGGGTTTGAGGAAGCCATTAGTAATTGGCCAAAAGAAGTCCCTGGAATTCATTGTATCTGGAATCAGTAT
ATTGAGTGTGCCCTCCAAGAAGAAGGTCCTGATTTTGCAAAAGAACTCACTGTCCGCTGGTTTAACTCCGTTTCAAAAGTTCAATATCCTCAAAATGAGA
TTTTGGATGCGGTGGATGGTAATAGCTCACTTGGATCACTGGAATCAGCTTCAGCATCAAATCTAGACTTTCTGATACCCAACTCTAATCAGATGGATAT
GATGTTTGGATTAATTAATCTATCTCTTGCTAAATTATTGCACAAAGACCACGTTGAAGCTCATGTTGCCATTGACAGAGCACTGAAGGCTGCACCTCCA
GAGTACATCAAACATTGCTTATCAGAGCATGCTGTGTTCTTGCTAAACCATGAGCCAAAATTAAGGAAGGATGCTCCTGTCAGTGAAAAACTGAAAATTT
TGAATGGTTATTTGAATGATACCCAAGCTCTCCCAGTTTGCGAACCATTATCTAGACGATTCATTGACAACATTGAGAAGCCAAAAGTTCAGCAGCTAAT
CAGTAGCATATTGAGTCCGGTTTCATCTGACTTTTCTTTGGTGAATCTTGTTCTTGAAGTGTGGTATGGCCCCTCTCTTTTACCCCCAAAGTCAAACCAG
CCAAAAGAATTGGTGGATTTTGTTGAAGCCATCTTAGAGATGGTGCCATCCAACTACCCAATAGCTCTTTCTGTTTGCAAGCTCTTGTGCAGAGGCTACA
GCTATATTAATGTTACTTCTGACAGTGTTCTATACTGGGCGTGCTCAATCTTGGTCGATGCAATCTTTCATGCTATTCCAGTGCCACCGGAATTTGTGTG
GGTTGAAGCTGCAGGTATTTTGGGTGATATTTCTGGTGTCAAGTTAATTTCTGACAGGTTTTACAAGAAAGCTTTATCTGCACATCCATTCTCTATGAAG
CTATGGAGTTGCTATTATAATCTATCAAAGAGTAGAGGATATGTAAGCACTGTTATCCAAAAAGCTAGAGAGAGGGGTATTGAAGTTGGTTGA
|
|||||||||||||||
|
AA sequence
|
>Potri.008G052500.1 pacid=42805836 polypeptide=Potri.008G052500.1.p locus=Potri.008G052500 ID=Potri.008G052500.1.v4.1 annot-version=v4.1
MASNNPSQPEFATKTREEGEVSSSSNDDQNPVCASALFADAFNTPASARTIPFHPMNKFTLSNRAGKAIFSTNPARCVDPNLQTSQPNNNKSFEKNRVPH
ISANPGKCASSGANDNLVIRFSDDESGSESEDRDDKPLKTKPNTTVVNGNGRLPPILPAKSSMSLQTSRNVNSIPKKSSMSRTFNSSMTKINGVANSKGA
DSSSVGQGSQVKNINSIKRNLASQEHGIEQGVDLNSTKVQDLRQQIALRERELKLKAAAQNKESGSVSDKCMNISSSVTRKSNAASSEVGQLAPKEPDRK
RIKPDGSYSKHLNSDGQQKMLVEKSNLPSKDQALENSSLQDRNMGNCSKKERPTKRTESSVVKWERQDRRVDISSAKLPASHINDNSSQPDTSRMQMDPC
VVLNQTPLLTNANASTLSKKRKSVDLNPVKNCGTQPPACLLKTSTSGQNLINSCEHLQGISGDKLSCQASLNLNPWNCLGTVNVAEHNSIDIQLLVEMEE
SLDRELEEEQEHRHKCEIEERNALKAYRKAQRALIEANSRCTELYRKRELHSAHFRSLIVNDSSLFFPSRQDEHVGIGMDRENNVSRNVDLIPSSSDQMQ
PEYDGCNQPGYDSVTGAPSNSLYQHVNGHSLGSEPCSEPDASTSEPLPRNSLIAANGVSSQSNDSNISAGEDEETFPLDHETDQPIFKIQQRDQNSVGRE
SHTDCHPNKDFYVDGPQDSLILEAKLRSKLFARLPIRTFSKNGGSSNMEPADEPGIEIDNRSERTQGSNVSIPLSETEKDRDYDLEGNNKPERSISELPV
QIQNHEKNFHSAADSKDDSTGGHQLTTSVISSPLLVLRSAFAQMKAMHPMTLIESQCRKNQQNDTCGDFIVEDGFMDTEEIQCDNVIAKSKEEIIRGMCG
TEIGTFTHNVAVDPFWPLCMYELRGKCNNDECPWQHVRDFSDQNLHPNQHDDSDSADCQVGLTLHEQKCKGGAKLSKCHSVLNPPTYLVGLDVLKSDSYK
SVIARRNGQCWQIQFSLCLALSSFFQKDLLADQLSIRADDGRIEVHGSWNRQTSYFQSRENTVNHLNQALASSLQSLEMALLFLCQEVYKLEGMKKPLSM
LSRAIEADPTSEALWMMYLLIYYSNIESIGKDDMFSYAVKNNERSYGLWLMYINSRIHLDDRMVAYNAALTALCRQASAFDKGNMYASACILDLFLQMMD
CLCMSGNVGKAIQKIQGLFPVAANSDEPHFLLLSDILACLTNSDKYIFWVCCVYLVIYRKLPDAIVQCFECDKELLAIEWPYVQLPNEEKQRAVKLVEMA
VDSVEMSVNSESLESDKNGRMAQQFALSHIRCTLVFDGLACCQNLLGKYTKLYPSCVELVLLSARLKKNGLGSVSFEGFEEAISNWPKEVPGIHCIWNQY
IECALQEEGPDFAKELTVRWFNSVSKVQYPQNEILDAVDGNSSLGSLESASASNLDFLIPNSNQMDMMFGLINLSLAKLLHKDHVEAHVAIDRALKAAPP
EYIKHCLSEHAVFLLNHEPKLRKDAPVSEKLKILNGYLNDTQALPVCEPLSRRFIDNIEKPKVQQLISSILSPVSSDFSLVNLVLEVWYGPSLLPPKSNQ
PKELVDFVEAILEMVPSNYPIALSVCKLLCRGYSYINVTSDSVLYWACSILVDAIFHAIPVPPEFVWVEAAGILGDISGVKLISDRFYKKALSAHPFSMK
LWSCYYNLSKSRGYVSTVIQKARERGIEVG
|
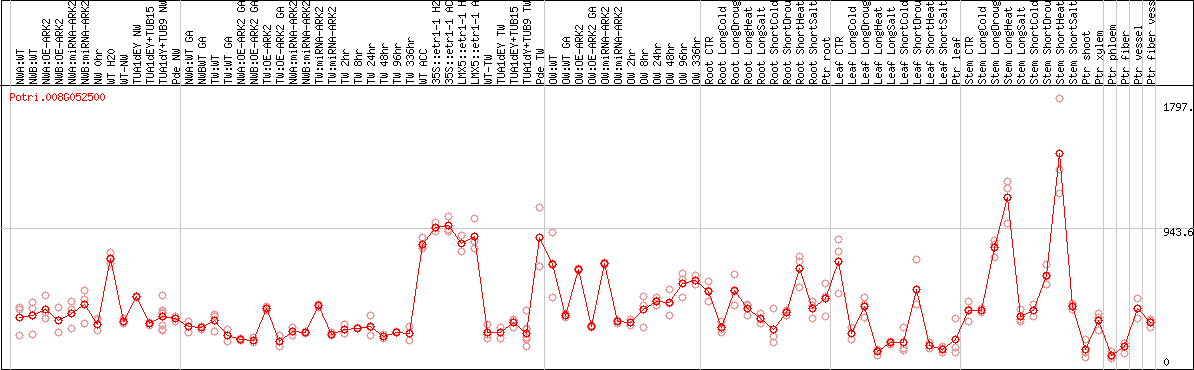
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G052500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
