External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G02530 97 / 1e-24
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT5G59950 94 / 6e-24
RNA-binding (RRM/RBD/RNP motifs) family protein
AT5G37720 80 / 3e-18
DIP2, ALY4
interacting with DNA-binding domain of Zn-finger PARP 1, ALWAYS EARLY 4 (.1.2)
AT1G66260 78 / 2e-17
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G23350 42 / 0.0001
PABP4, PAB4
poly(A) binding protein 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G208000
116 / 2e-32
AT5G59950 138 / 8e-40
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.001G236900
100 / 1e-26
AT5G59950 187 / 2e-59
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.016G093944
100 / 2e-26
AT5G59950 196 / 1e-62
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.017G129900
89 / 1e-21
AT1G66260 216 / 3e-69
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.004G087000
86 / 1e-20
AT5G37720 213 / 2e-68
interacting with DNA-binding domain of Zn-finger PARP 1, ALWAYS EARLY 4 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004946
101 / 1e-26
AT5G59950 209 / 3e-67
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10005445
101 / 1e-26
AT5G59950 210 / 9e-68
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10024413
99 / 1e-25
AT5G59950 219 / 1e-71
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10025329
99 / 1e-25
AT5G59950 224 / 2e-73
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10005895
95 / 3e-24
AT5G02530 185 / 8e-58
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10023417
94 / 1e-23
AT5G59950 130 / 3e-37
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10000446
83 / 3e-19
AT5G37720 211 / 1e-66
interacting with DNA-binding domain of Zn-finger PARP 1, ALWAYS EARLY 4 (.1.2)
Lus10010982
83 / 4e-19
AT5G37720 209 / 7e-66
interacting with DNA-binding domain of Zn-finger PARP 1, ALWAYS EARLY 4 (.1.2)
Lus10040299
58 / 3e-10
AT5G59950 77 / 1e-16
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10016174
45 / 6e-06
AT3G52380 275 / 3e-91
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.008G052601.2 pacid=42806110 polypeptide=Potri.008G052601.2.p locus=Potri.008G052601 ID=Potri.008G052601.2.v4.1 annot-version=v4.1
ATGATGATGTTGAGGTTTGGGGTATTATGGGAATGTACGTGTGCAGTAGCTAAGGCCGTGATGTTCTTTCATGACGATGATAAAAAAAAGGCAGCAGGGC
ACTATGTTGTCACGTGTCCTATAATCGAGGTGATTTTCTCCGAGGTTGGTGATTTGTTGAGGTGCTCACTTCATTATGATATGAGTGGGAGATCAAAGGG
AACAGCCGAAGTTGTATTTGCACTCCAAACAGATGCTCTAGCAGCTATCAAGAGATACAATAATGTTCAGCTTGATGGGAAGCCCCTGAAAATTGAGCTT
GTAGGGGATAATGTGATAACTCCTGTCCCTGTGCTTGTAACAACAACTACCAACTTGGCAAAACCAAAAATGTCATTAGAAGTGTTCAAGAAAGAATTGG
TGCGAGGGAAGGGCCATGGCAGCGGTCAAGAATTGGCCAGAGGCCACAGCCAGGGAGACACCATGTCGAGAAGCTTACTGCAGAAGCACTTGATTCTGAC
CTGGACAGACATACCACCTTGA
AA sequence
>Potri.008G052601.2 pacid=42806110 polypeptide=Potri.008G052601.2.p locus=Potri.008G052601 ID=Potri.008G052601.2.v4.1 annot-version=v4.1
MMMLRFGVLWECTCAVAKAVMFFHDDDKKKAAGHYVVTCPIIEVIFSEVGDLLRCSLHYDMSGRSKGTAEVVFALQTDALAAIKRYNNVQLDGKPLKIEL
VGDNVITPVPVLVTTTTNLAKPKMSLEVFKKELVRGKGHGSGQELARGHSQGDTMSRSLLQKHLILTWTDIPP
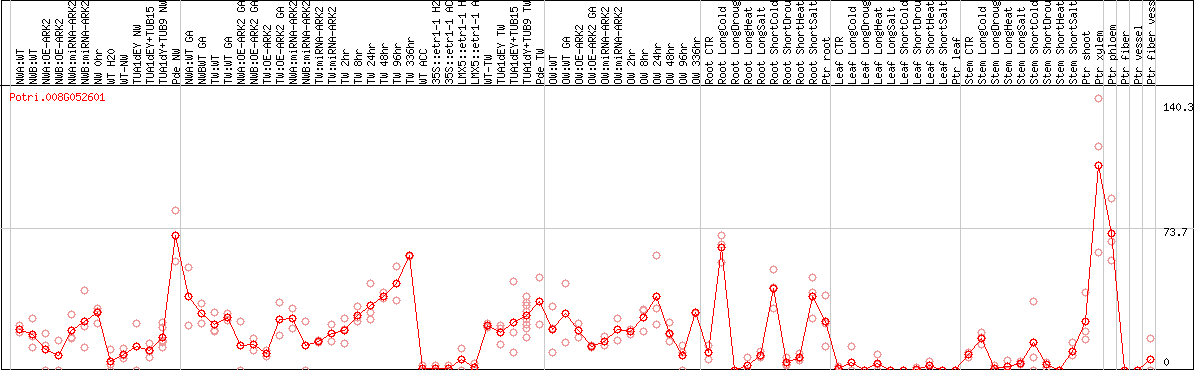
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G052601 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.