External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G39650 293 / 4e-99
Protein of unknown function (DUF506) (.1)
AT3G07350 130 / 1e-35
Protein of unknown function (DUF506) (.1)
AT4G14620 127 / 3e-34
Protein of unknown function (DUF506) (.1)
AT2G38820 125 / 1e-33
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
AT3G22970 120 / 4e-32
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
AT4G32480 119 / 1e-31
Protein of unknown function (DUF506) (.1)
AT3G54550 117 / 8e-31
Protein of unknown function (DUF506) (.1)
AT3G25240 116 / 1e-30
Protein of unknown function (DUF506) (.1)
AT2G20670 109 / 9e-28
Protein of unknown function (DUF506) (.1)
AT1G62420 102 / 2e-25
Protein of unknown function (DUF506) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G203700
481 / 4e-173
AT2G39650 287 / 9e-97
Protein of unknown function (DUF506) (.1)
Potri.005G216900
135 / 9e-37
AT3G22970 320 / 2e-107
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.002G046300
134 / 2e-36
AT3G22970 322 / 5e-108
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.008G159800
125 / 6e-33
AT3G22970 350 / 3e-119
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.019G099600
117 / 1e-30
AT4G32480 290 / 4e-98
Protein of unknown function (DUF506) (.1)
Potri.007G109700
115 / 1e-30
AT2G38820 147 / 1e-42
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.005G058200
115 / 2e-30
AT2G38820 154 / 3e-45
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.010G080600
116 / 8e-30
AT3G22970 371 / 3e-127
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Potri.004G044300
113 / 5e-29
AT4G32480 260 / 3e-86
Protein of unknown function (DUF506) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002695
406 / 3e-143
AT2G39650 313 / 4e-107
Protein of unknown function (DUF506) (.1)
Lus10023412
360 / 2e-125
AT2G39650 273 / 2e-91
Protein of unknown function (DUF506) (.1)
Lus10040293
350 / 4e-121
AT2G39650 270 / 5e-90
Protein of unknown function (DUF506) (.1)
Lus10000956
328 / 1e-113
AT2G39650 266 / 1e-89
Protein of unknown function (DUF506) (.1)
Lus10014718
132 / 9e-36
AT3G22970 327 / 3e-110
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Lus10003087
130 / 6e-35
AT3G22970 318 / 7e-107
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Lus10039397
129 / 8e-34
AT3G22970 362 / 2e-122
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Lus10018754
127 / 1e-33
AT2G38820 319 / 6e-108
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Lus10006634
127 / 2e-33
AT3G22970 367 / 2e-125
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
Lus10024840
125 / 4e-33
AT3G22970 324 / 4e-109
Protein of unknown function (DUF506) (.1), Protein of unknown function (DUF506) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04720
PDDEXK_6
PDDEXK-like family of unknown function
Representative CDS sequence
>Potri.008G056100.1 pacid=42807109 polypeptide=Potri.008G056100.1.p locus=Potri.008G056100 ID=Potri.008G056100.1.v4.1 annot-version=v4.1
ATGATGTGTGCGGCGACGGCAACAGGGGATTGGTGGGCGAGGGGGATTGGTGGGCAGATCGGGGGAGCTTTTAGCCACGAGAGCGAACATGATTTGGCTC
TCATGGTTAGCGATTTTCTCGAGAACGGTTGTAGCTCCGGTGCCGATTCCTGGTGTAGCAGCGATAGCGATTCTGGTCTCTCTGATCTTCACCATCTCGC
CGACAAGATTTCGTTTTACAAGCGCACAGTAGCTCAATATGAGAGTGACTTGTTGTTGATAGTTCATTCTCTTGTGCAATCGATCAAGGACACAGACCTT
CATCGTGTCAAGTCAGGTCCGTGTAATGCAAGCTGCATCAACTTTTCTTTGGTAAAACTGTTGAGGCTTTCGGGATATGATGCTGCTGTTTGTGTGTCCA
AGTGGCAGGGTAGTGGCAAGGTGCCGGGAGGGGATCATGAATATATTGATGTGGTCAACTGCATCAATGCTGGAAGCTCTGAGCGTGTGATCATTGATGT
TGACTTCCGAAGCCACTTCGAAATAGCTAGAGCTGTTGACACTTATGATAGAATATTGAAGTCACTTCCAGCCATTTATGTAGGCTCCTTGACCAGGTTA
AAACGGTATCTTCAAGTCATGGCCGAGGCAGCCAGATCTTCTCTCAAGCAAAATTCGATGCCTCTTCCACCTTGGAGGTCTCTGGCTTATTTGCAAGCGA
AATGGTACTCACCATATCAGAGACAGTTTGGTCCAGATAATCAAAATTTTAGTAGTGTCGATTCTTCATACCATAAGCAGTGTGGTGGACATTTAAAGAG
ACTGCAGTCCTCGCTGCAATTTGAAACAGAAGGAGAACGACTCATGAAGCCTATTAACAGTGATAATAACCGGGGGATGAAGTTTGAGAGGCGGAAACAC
TCCTTATTCAGGGCTCTTTGA
AA sequence
>Potri.008G056100.1 pacid=42807109 polypeptide=Potri.008G056100.1.p locus=Potri.008G056100 ID=Potri.008G056100.1.v4.1 annot-version=v4.1
MMCAATATGDWWARGIGGQIGGAFSHESEHDLALMVSDFLENGCSSGADSWCSSDSDSGLSDLHHLADKISFYKRTVAQYESDLLLIVHSLVQSIKDTDL
HRVKSGPCNASCINFSLVKLLRLSGYDAAVCVSKWQGSGKVPGGDHEYIDVVNCINAGSSERVIIDVDFRSHFEIARAVDTYDRILKSLPAIYVGSLTRL
KRYLQVMAEAARSSLKQNSMPLPPWRSLAYLQAKWYSPYQRQFGPDNQNFSSVDSSYHKQCGGHLKRLQSSLQFETEGERLMKPINSDNNRGMKFERRKH
SLFRAL
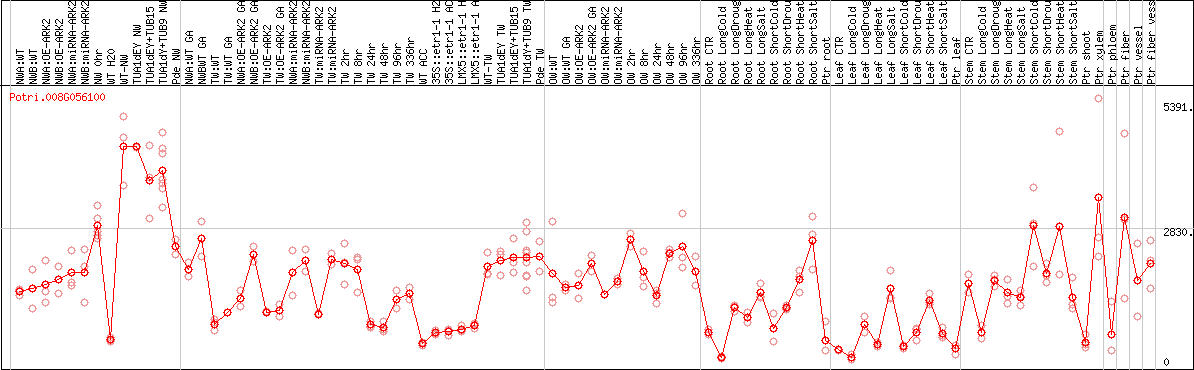
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G056100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.