External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G11260 176 / 9e-57
HD
WOX5B, WOX5
WUSCHEL related homeobox 5B, WUSCHEL related homeobox 5 (.1)
AT5G05770 127 / 3e-38
HD
WOX5A, WOX7
WUSCHEL related homeobox 5A, WUSCHEL related homeobox 7 (.1)
AT3G18010 100 / 2e-25
HD
WOX1
WUSCHEL related homeobox 1 (.1)
AT5G59340 96 / 3e-24
HD
WOX2
WUSCHEL related homeobox 2 (.1)
AT2G28610 96 / 3e-24
HD
PRS1, PRS, WOX3
WUSCHEL RELATED HOMEOBOX 3, PRESSED FLOWER 1, PRESSED FLOWER, Homeodomain-like superfamily protein (.1)
AT1G46480 90 / 5e-22
HD
WOX4
WUSCHEL related homeobox 4 (.1)
AT2G17950 89 / 2e-21
HD
WUS1, PGA6, WUS
WUSCHEL 1, WUSCHEL, Homeodomain-like superfamily protein (.1)
AT2G01500 85 / 3e-20
HD
WOX6, PFS2, HOS9
WUSCHEL RELATED HOMEOBOX 6, PRETTY FEW SEEDS 2, HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENE 9, Homeodomain-like superfamily protein (.1)
AT5G45980 72 / 5e-15
HD
WOX9B, STPL, WOX8
WUSCHEL related homeobox 9B, STIMPY-LIKE, WUSCHEL related homeobox 8 (.1)
AT2G33880 67 / 3e-13
HD
WOX9A, STIP, WOX9, HB-3
WUSCHEL related homeobox 9A, WUSCHEL-RELATED HOMEOBOX 9, STIMPY, homeobox-3 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028271
204 / 1e-66
AT3G11260 177 / 4e-56
WUSCHEL related homeobox 5B, WUSCHEL related homeobox 5 (.1)
Lus10040219
201 / 7e-66
AT3G11260 179 / 4e-57
WUSCHEL related homeobox 5B, WUSCHEL related homeobox 5 (.1)
Lus10018011
101 / 2e-25
AT3G18010 164 / 9e-47
WUSCHEL related homeobox 1 (.1)
Lus10042007
101 / 2e-25
AT3G18010 174 / 2e-50
WUSCHEL related homeobox 1 (.1)
Lus10038480
94 / 3e-24
AT2G28610 130 / 1e-37
WUSCHEL RELATED HOMEOBOX 3, PRESSED FLOWER 1, PRESSED FLOWER, Homeodomain-like superfamily protein (.1)
Lus10024808
90 / 5e-22
AT1G46480 195 / 7e-62
WUSCHEL related homeobox 4 (.1)
Lus10041909
91 / 9e-22
AT2G17950 145 / 4e-41
WUSCHEL 1, WUSCHEL, Homeodomain-like superfamily protein (.1)
Lus10023332
85 / 2e-21
AT2G28610 109 / 9e-31
WUSCHEL RELATED HOMEOBOX 3, PRESSED FLOWER 1, PRESSED FLOWER, Homeodomain-like superfamily protein (.1)
Lus10016569
88 / 3e-21
AT5G59340 121 / 6e-33
WUSCHEL related homeobox 2 (.1)
Lus10028457
89 / 4e-21
AT2G17950 146 / 3e-41
WUSCHEL 1, WUSCHEL, Homeodomain-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00046
Homeodomain
Homeodomain
Representative CDS sequence
>Potri.008G065400.1 pacid=42807798 polypeptide=Potri.008G065400.1.p locus=Potri.008G065400 ID=Potri.008G065400.1.v4.1 annot-version=v4.1
ATGGAAGAGAGAATGTCAGGCTTTTGTACCACAAAAGCAGGGCGTGGTGGGAGCAGTGGCAATAACTATGCCTCAGGAACTAAATGCGGGCGTTGGAATC
CTACTATCGAACAAGGTAAACTTCTAACTGACTTGTTCAGGTCTGGTGTTCGAACCCCAAGTACTGATGAGATCCAAAACATCTCCACTCGGCTTAGTTT
TTATGGCAAGATCGAGAGCAAGAACGTTTTCTACTGGTTTCAAAATCATAAAGCTAGGGAAAGACAGAAGAGGCGCAGAGTTTCTGTCGATGAGAAGGAT
GTCATGATTCGTCGAGATGACAAATTTTCTTCTGCGCGATATTTTACTGAAATAGGTCAGGTGAACGAACGAGAACAAGTGATTGAGACTCTTCAACTTT
TCCCTTTAAAATCCTTTGATGAAGTAGAGTCAGAGAAGTTCAGGTTGCAGGCAAATGAATGCAATGAAGCTGCAGCTGCGTTTTCTTATAAATTTGGTAC
AGAAATGGATCGTCCACAACTAGATCTGCGATTAAGCTTTCTGTAA
AA sequence
>Potri.008G065400.1 pacid=42807798 polypeptide=Potri.008G065400.1.p locus=Potri.008G065400 ID=Potri.008G065400.1.v4.1 annot-version=v4.1
MEERMSGFCTTKAGRGGSSGNNYASGTKCGRWNPTIEQGKLLTDLFRSGVRTPSTDEIQNISTRLSFYGKIESKNVFYWFQNHKARERQKRRRVSVDEKD
VMIRRDDKFSSARYFTEIGQVNEREQVIETLQLFPLKSFDEVESEKFRLQANECNEAAAAFSYKFGTEMDRPQLDLRLSFL
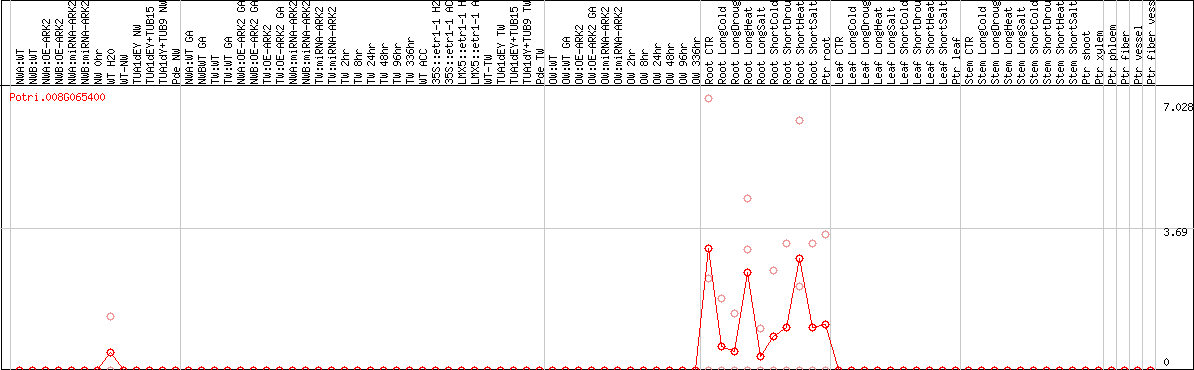
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G065400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.