Potri.008G069100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G069100.1 pacid=42807837 polypeptide=Potri.008G069100.1.p locus=Potri.008G069100 ID=Potri.008G069100.1.v4.1 annot-version=v4.1
ATGCCGACGCCGGTAACAACAGCAAGGCAATGCTTGACTGAAGAAGCAGCTCACGCTCTCGACGAGGCAGTGAACGTCGCACGCAGAAGAGGACATGGCC
AGACCACGTCGCTTCATGCCGTCTCGGCTCTGTTATCACTCCCTTCGTCACCTCTACGCGAAGCGTGTGCGCGTGCGAGGAACTCTGCGTACTCACCGCG
ACTCCAATTCAAAGCGCTCGAGCTCTGCCTCGGGGTGTCACTCGACCGAGTGCCGACGAGTCAACTCGGTGATGACTCACCTCCTGTTTCGAACTCGCTC
ATGGCTGCTATTAAGCGTTCGCAAGCGAACCAGAGGAGACAGCCGGAGAATTTCAATTTGTATCATCAAATTCAACAGCAGCAGCAATCGTCTTCTTCGA
TTTCGTGTATTAAAGTGGAGCTTCAGAACCTGATTCTATCGATTTTGGATGATCCGGTTGTGAGCCGGGTTTTCGGTGAAGCGGGATTTCGGAGCTCGGA
AATCAAGTTGGCTATTGTTAGACCGTTGCCTCAGGTTTTTAAATTCCCATCCTCACGTTTCAAAGGCCCGCCCTTGTTTTTATGCAATATATTATCAAGT
GAGGATCCGGATTCGTTGTACTCGTGTCCGGGTCGGAGCGGTGTTTTTAGTTTTCCATTTTCGGGCGGGTCGTTCCTTAATAATAACAACAGCAGTCATA
GTACTACTAATAGGGATGTTAATTGTCGGAGGATTGGTGAGGTTCTAGCGAGCAGCAGAGGGAGGAATCCTCTGCTTGTAGGTTCAGGTGCTTACGATAC
ACTTGCAATTTTTAGTGAGATAGTAGAGAAAAGAAAAGAGAATATCTTGCCTGTGGAGTTACGTGGGTTAAGTGTTATTTGCATTGAAAGCTATGTTAAC
AAGTTTATTACAAGTGAGGATTTTGATAAAAAGAGAGTGGATTTGAGGTTTGAGGAGTTGGGTCAGTTTGCGGAGAGGCATTTGGGACCTGGGTTGCTGG
TGAATTTTGGTGACTTGAAGGCATTTGTTAGTGACGACAGTGATAATAATGGTTTGGGTGATGCAGCTAGTTATGTTATTGAGAAACTGACCAAGCTGTT
GCAGTTATATGGAGGGAGGGTGTGGCTTATCGGTGCGGCGAGTTATGAGAACTATTCCAAGTTTGTTGGAAGATTTCCTTCCACTGAAAAGGATTGGGAT
TTGCAGCTCTTGCCTATCACTTCTCTCCCGACTTCTTCCATGGCTGAATCTTACCCCAGGTCAAGCTTGATGGAGTCATTTGTTCCATTTGGCGGGTTCT
TCTCCACACCTTCTGACTTGAATGGCCCTTTAAATACCCCATACCATTGTATACCCCTTTGTCATCTATGCAATGAGAAGTGCAAACAAGAAATACTTTC
TGTTTCAAAGGGAGGATTTGTCGGCTCAGTAGCAGACCATTACCAATCTAGCTTACCTTCTTGGCTGCAGATGGCTGAAATCGGCACTAACAAGGGACTG
GATGCGAAGACCAGAGATGATGGAACGGTATTGAGTGCCAAAGTTGCAGGTCTGCAAAGGAAATGGGACAATATATGTCAGCGTCTTCATCACACCCAGC
CACCTGGGTTAAACACTCATCTCCCTCAGTTTCCAACTGTTGCGGGCTTTCAGCTGGTTGAAGACAAAAAGGAAAATGCTGAAAATCCTAGAAGCAAAAA
TACAAGTGCACTGCCAAATGGAAGCAGATGTGTGAATGTAAATTCATGCATTCCCTCTGACATACAAAAGACACCGAGAAAGCAATTAGGTTTTCCTCTT
CCTATTGTTTCTGAGGCTAAGAGTGATTGTATTCTATCCAAGCAACGGGAAAAACCTTCAAAAGAAGAAGATCTTGAGTCTGGTGGCTTCAGTTCTCCAC
AGAATTTCTCCAATTCGAGCATAGTTGATGGCAGTCAAGCATCTCCAACATCCATGACTTCTGTGACCACAGATTTGGGCTTGAGAATAAGTTCCGTCCC
TACTAGTAATGAACTGAAGAAAACTGTAAATCAAAACCACATGGAGCTTCCACAGGACCGATCAGGTTCCTTTTCAGCAAATGTTGATGTTGTGCATGGG
AGTATGTCTGACCATTGGGCTCCATCATCATCATCATCTTCTTCTCCTGACTACGGCGGGCAGTTTGATCTCAGCAATGCCAAGATGCTCTTTAGAGCTG
TCGTTGAAAGGGTTGGCTGGCAAGATGAAGCTATACGCGTTATTAGCCAAACAATAGCTCGCTGCAAGGCAAGAAATGAGAAACGACAGGGGGCAAGTCT
CAGAGGGGATATATGGTTCAGTTTCTGTGGACCTGATAGGCGTGGGAAAAAGAAAATTGCTTCTGCCCTTGCAGAAATCATATATGGAAGCAGGGAGAAC
TTTATTTCTGCAGATTTAAGTGCCCAAGATGGGATGATCCATACCCACATGCTCTTTGACCACCCAGAGGTGAATGGCTACACTGCAAAGTTAAGGGGGA
AGACTGTGGTTGATTTTGTTGCCGGGGAGCTGTGCAAGAAACCCTTGTCTATTGTGTTTCTTGAAAATATAGACAAGGCTGATGTGCAGGCTCAAAAGAG
CTTGTCACATGCTATTCAGACTGGTAAATTTGCAGATTCTCATGGGAGAGAAATTGGAATCAGCAATGCAATATTCGTGACAACTTCAACATTGACGGAG
GATAAAGTTTGCTCTTCTAGTAATGAGTTCTTCACCTATTCCGAGGAGAGAATATCAAGAGTCAGGGACTGGCCAGTGAAGATATTAATTGAACAAGCTC
TTGATGACGAAGTAGGCAAAATGGTTGCTCCATTCACACTGAGAAAAGGCGTCTCTGGTTCTATCTTTTTGAATAAAAGAAAGCTGGTTGGGGCAAATCA
AAATCTAGACAGACAAGAAATCAAAGAGATGGTGAAACGAGCTCATAAGACGTCAGCTAGGAATCTAGATCTGAACCTTCCTGCAGAAGAGAATGATGTG
CTAGACACTGACGATGGAAGCTCTGATAATGATCATGCATCTGACAACTCCAAGGCTTGGTTGCAAGATTTTCTTGAAAAAATTGACGCAAGAGTGTTTT
TCAAGCCTTTTGATTTTGATGCTCTTGCTGAAAGAATATTGAATGAACTTAATGGGTGCTTCCACAAGATTGTAGGTTCGGAATGCTTGCTAGATATTGA
TCCAAAAGTCACGGAGCAGTTACTCGCAGCTGCGTATTTATCTGATCGGAAGAGAGTGGTTGAGGATTGGGTGGAGCAAGTTCTTGGTTGGGGATTTGTT
GAAGTTTTGAGAAGGTACAAACTGAAAGCAAATTCCATTGTGAAACTTGTTGCCTGCAAGGGCTTGTTTGTAGAAGAGCGCATGTCAGGAGATCACCTTC
CAACAAAAATTATAATCAGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G069100.1 pacid=42807837 polypeptide=Potri.008G069100.1.p locus=Potri.008G069100 ID=Potri.008G069100.1.v4.1 annot-version=v4.1
MPTPVTTARQCLTEEAAHALDEAVNVARRRGHGQTTSLHAVSALLSLPSSPLREACARARNSAYSPRLQFKALELCLGVSLDRVPTSQLGDDSPPVSNSL
MAAIKRSQANQRRQPENFNLYHQIQQQQQSSSSISCIKVELQNLILSILDDPVVSRVFGEAGFRSSEIKLAIVRPLPQVFKFPSSRFKGPPLFLCNILSS
EDPDSLYSCPGRSGVFSFPFSGGSFLNNNNSSHSTTNRDVNCRRIGEVLASSRGRNPLLVGSGAYDTLAIFSEIVEKRKENILPVELRGLSVICIESYVN
KFITSEDFDKKRVDLRFEELGQFAERHLGPGLLVNFGDLKAFVSDDSDNNGLGDAASYVIEKLTKLLQLYGGRVWLIGAASYENYSKFVGRFPSTEKDWD
LQLLPITSLPTSSMAESYPRSSLMESFVPFGGFFSTPSDLNGPLNTPYHCIPLCHLCNEKCKQEILSVSKGGFVGSVADHYQSSLPSWLQMAEIGTNKGL
DAKTRDDGTVLSAKVAGLQRKWDNICQRLHHTQPPGLNTHLPQFPTVAGFQLVEDKKENAENPRSKNTSALPNGSRCVNVNSCIPSDIQKTPRKQLGFPL
PIVSEAKSDCILSKQREKPSKEEDLESGGFSSPQNFSNSSIVDGSQASPTSMTSVTTDLGLRISSVPTSNELKKTVNQNHMELPQDRSGSFSANVDVVHG
SMSDHWAPSSSSSSSPDYGGQFDLSNAKMLFRAVVERVGWQDEAIRVISQTIARCKARNEKRQGASLRGDIWFSFCGPDRRGKKKIASALAEIIYGSREN
FISADLSAQDGMIHTHMLFDHPEVNGYTAKLRGKTVVDFVAGELCKKPLSIVFLENIDKADVQAQKSLSHAIQTGKFADSHGREIGISNAIFVTTSTLTE
DKVCSSSNEFFTYSEERISRVRDWPVKILIEQALDDEVGKMVAPFTLRKGVSGSIFLNKRKLVGANQNLDRQEIKEMVKRAHKTSARNLDLNLPAEENDV
LDTDDGSSDNDHASDNSKAWLQDFLEKIDARVFFKPFDFDALAERILNELNGCFHKIVGSECLLDIDPKVTEQLLAAAYLSDRKRVVEDWVEQVLGWGFV
EVLRRYKLKANSIVKLVACKGLFVEERMSGDHLPTKIIIS
|
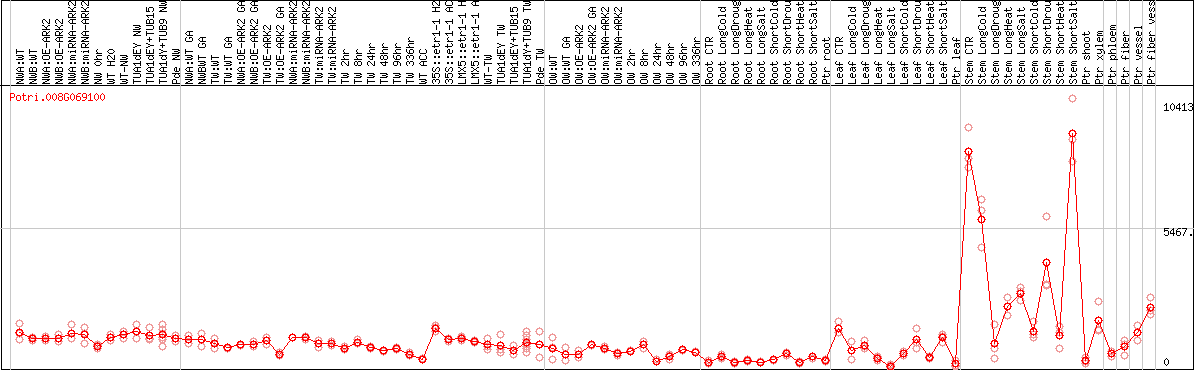
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G069100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
