Potri.008G073500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G073500.1 pacid=42808377 polypeptide=Potri.008G073500.1.p locus=Potri.008G073500 ID=Potri.008G073500.1.v4.1 annot-version=v4.1
ATGAAGAGAAAGCGTTTACATCAGTCCAAGCATCCATTTAATGCACACCCTTTTGAGGCCTTATGCTGTGGCTCTTGGCAAAGCGTGGAGCTTATACAAA
TCAGGGATGGGGCTATGACTGTGCATTTTGTAGATGGCCATCATAGAATTGAGGAGAAGGGTCCTTTCTCAAACGTTCGAGTGAAGTCGAGGAAAGCTAC
TTCATCTGATTGCACATGCTTTCTGAGGCCTGGTATTGATGTTTGTGTGCTTTCATCTTCTGAGCGTGCGAAGAATACTGGGGAAGGAAATTCAGAACCT
GTGTGGGTTGATGCTAAGATAAGTTCTATCAAGCGGAAACCTCATGTATCTCACTGCTCATGTCAATTTTTTGTTAACCTCTATGTTAACCAAGGCCCTC
TTGGTTCAGAGAGAGCCAGACTTAGTAAAGAAACTGAAGCAGTAGGAATCAATGAAATTTCTGTTCTTCAGAAACTTGACAACGATCCATGTGAAGCAGA
CAACAATCAGCAAGAAGCTCAGTTCTATCGATGGGAATTTTGTGAGGACTGTTCTTTAGTACAAAGAAGCAAACTCTTTTTGGGGAGGTTTTCAGCTGAC
CTTACATGGCTGCTTGTAGCATCAGTTCTGAAGCAAGTTGAATTTGATGTAAGGTCAGTCCAAAATAAAATTGTCTACCAAATTTTGGGTGGTGAGAATG
AGCACTGTTCATTAAAATCTAATAATCACATAAACTGTGTAACTTTCAAAGTGAAAGACAGTATTTCAACCCCATTTGTGGTCCAATTGGTTCCCACTGA
TGCTTGTAGTGAAGCAGGCCATATTAGTGACACCAATGGGACCAAACAATCACCATGCTATGATGTTATGAGTTTGCGGCGGTCAAAGCGTCGAAATGTA
CAACCTGAACGCTTCCTTGCCTGTGATGCTCCTGCAGAGACAGAAATTGGCTGGGTAAGATCCCTGCCATACACACCACTGAAGTGGAAGGCAGAAGAAG
AAGAAGAAGAAGAAATGCACTTACCACTGGCATACCTGTTTGGGACACATGCAGGTGCAAGTTGTGCAGAAGAGCAAACATGCAACGAAGTGGGAGCTAG
TTCTCCTAAACTAGAACTCTTGGAAGGTATCCCTGTTTCCAGAACCAAGACTTACTTGAAGGAGATTAAATCAAATGTGGTTAACAGAAGAGATCACCAA
ACTGAGCCCGGGGAGGTTCGAGCAGGGATGGCTAAGAGAAGAGAATGCCAAAAGTCAACCATGGCTGACCGAATAGAACACCAAACTCGACTCGGGGATG
CTGAATCAGGCATGGCCAACAGAAAAAAGCATGGAACTCAAATTAGGGAGGTTAAATCAGGCGTGGCTAACAGAAGAGAACACCAAGATCAACTTGCTAT
TGTTCCTGTGCATACTGAAGATGTTCTGGCAACCTTTGAACAATTTGATTCCCCTGTGAAGACTCCTGAACCTTATTCACAAGCGTTTATTGAATTTCCG
ATTAGTTATTACAGAAAAAAGAGTTCTCCTGCTGCGCACAGGAAGAATGATCGTGATGAAGATTTGATGTTCGGAAATGGATGGGGTGGAAAATTTTCTA
CTAAGAAAGTTCAAAGAGCAAGATATCGATCCACACATTTAAAGCAGGATGGTTCTTGTGCGCCAATGACTTACAAGCGGACAGCCTTAAGTGCTGGTGC
TTACAATAAACTCATAAGTTCTTATATGAAGAACATTGATGCCACAATCAAGTCAAAGGAGGTACCACGCATTATTGACCAGTGGGAAGAGTTTAAAGCA
AAACATAGCTCAGATCAGAAGGAGAAAATGGAACCATCTTCGGTTAAGGATGATGGAGAAAGCTCAGAAACTGAAATGTTGTGGAGAGAGATGGAACTAT
GCCTGGCATCAGCATATATACTTGAAGATAATGAGGCTCTACTTTCCACTCGGACTACACAGAAAAATTGCCAGCACGACTTCAAACTGGATGAAGAAAT
TGGGATATTATGCCAAATATGCGGCTTTGTGAAAACTGAAATAAAATATGTTTCAGCACCATTTATGGAACACACAGGCTGGACTGCAGAAAGTAAGCCA
CAGAATGAAGAGGATTCGGAGCTTAAACCTGATGAAGATGAGGGCTCGAGTCTTTTTGGCAATCATACTTCAGGTGAAGATGTGCCTGTATCAGAAGTGA
ATGACAATGTTTGGGACTTGATCCCTGAGCTCAGACCGAAATTACACATGCACCAGAAAAAGGCTTTTGAGTTTCTATGGAAGAACACTGCTGGATCCTT
GGTACCGGCACATATGGAAAAAACATCTAAGAAGATAGGTGGTTGTGTTGTTTCTCATACTCCTGGAGCTGGAAAAACATTTCTCATCATTGCATTCCTT
GTCAGCTACTTGAAGTTGTTTCCAGGAAAACGGCCTCTGGTCCTTGCTCCAAAGACAACTCTCTATACCTGGTACAAGGAATTTATCAAGTGGGAAATCC
CTGTTCCAGTTCATCTTATCCATGGCACCAGATCTTCTAGAGCCTTCAAGCAGACTCCAGCAGCGCTTCGAGGATCAGGTCCAAGGCCCAGCCAGGATGT
TGTGCATATTTTGGATTGCCTGGAGAAAATGCAAAAGTGGCATGCACAACCAAGTGTGCTTGTCATGGGCTATACCTCATTTCTGACATTGATGAGAGAA
GACTCGAAGTACAACCACAGAAAATATATGGCTAAAGTGCTGCGAGAGAGTCCTGGAATGTTGATACTTGATGAAGGACACAACCCCAGGAGTACAAAAT
CAAGGTTGAGAAAGGTTTTGATGAAAGTGGAAACAGATCTGAGAATATTGCTTTCAGGTACTTTGTTTCAGAATAATTTTTGTGAATATTTCAACACCCT
TTCCCTGGCAAGACCAATGTTTATCAAAGAAGTTCTGAAGGCATTAGACCCGAAGTTTAAGAGGAAAAAGAAAGGGGCACAAAAGGCACGTCATTTGCTG
GAATCTCGCGCCAGAAAATTCTTCATAGATAACATTGCAAGCAAAATTAATTCGGATGAGGCTGAAGAGAAAATGCAGGGTTTGAATATGCTGAGAAACA
TGACTAATGGGTTCATTGACGTGTATGAAGGCACAGCATCTGACACCCTTCCTGGTATACAGATTTACACCATATTGATGAATCCAACAGATATTCAACA
TCAGATTTTGGTGAAACTCCATAAAATAATGGAAAAATGCCCTGGATACCCCCTGGAAGTAGAGCTTTTGATAACTCTTGCTTCAATACATCCTTCATTG
GTCAATTCTTCTGTCTGTGTTAAAAAATTTTATAATCTGGAGGAACTGATGGAACTTGAGAAGTTAAGATTTGATTGTAAGAAAGGGTCCAAGGTGATGT
TTGTGTTGAATCTTGTGTATCGTGTAGTCAAGAACGAGAAGGTTCTGATCTTTTGCCACAACATTGCACCCATTAAATTATTCCTTGAATTGTTTGAGAA
TATCTTTCGATGGCAGCAGGGTAAAGAAATTCTGGTTCTTACAGGGGAACTAGAGCTGTTTGAACGAGGAAGAGTGATGGATAAGTTTGAGGAATTGGGA
GGGCCTTCAAGGGTTCTACTTGCATCAATTACAGCTTGTGCCGAGGGAATAAGTTTGACAGCAGCTTCACGGGTAATTCTTTTGGACTCGGAGTGGAATC
CTTCCAAGACAAAGCAGGCCATTGCACGAGCTTTTCGGCCTGGCCAACAGAAGATGGTTTATGTGTATCAGCTTCTGGCAACAGGCACAGTTGAAGAGGA
CAAGTATCGTAGGACAGCATGGAAGGAATGGGTCTCACGCATGATATTTAGTGAGGAATTTGTGGAGGACCCTTCTCGTTGGCAAGCAGAAAAGATTGAG
GACGATGTATTACGGGAGATAGTAGAGGAGGACCGGGTTAAGTCATTCCATATGATAATGAAGAACGAGAAGGCTTCAACAAGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G073500.1 pacid=42808377 polypeptide=Potri.008G073500.1.p locus=Potri.008G073500 ID=Potri.008G073500.1.v4.1 annot-version=v4.1
MKRKRLHQSKHPFNAHPFEALCCGSWQSVELIQIRDGAMTVHFVDGHHRIEEKGPFSNVRVKSRKATSSDCTCFLRPGIDVCVLSSSERAKNTGEGNSEP
VWVDAKISSIKRKPHVSHCSCQFFVNLYVNQGPLGSERARLSKETEAVGINEISVLQKLDNDPCEADNNQQEAQFYRWEFCEDCSLVQRSKLFLGRFSAD
LTWLLVASVLKQVEFDVRSVQNKIVYQILGGENEHCSLKSNNHINCVTFKVKDSISTPFVVQLVPTDACSEAGHISDTNGTKQSPCYDVMSLRRSKRRNV
QPERFLACDAPAETEIGWVRSLPYTPLKWKAEEEEEEEMHLPLAYLFGTHAGASCAEEQTCNEVGASSPKLELLEGIPVSRTKTYLKEIKSNVVNRRDHQ
TEPGEVRAGMAKRRECQKSTMADRIEHQTRLGDAESGMANRKKHGTQIREVKSGVANRREHQDQLAIVPVHTEDVLATFEQFDSPVKTPEPYSQAFIEFP
ISYYRKKSSPAAHRKNDRDEDLMFGNGWGGKFSTKKVQRARYRSTHLKQDGSCAPMTYKRTALSAGAYNKLISSYMKNIDATIKSKEVPRIIDQWEEFKA
KHSSDQKEKMEPSSVKDDGESSETEMLWREMELCLASAYILEDNEALLSTRTTQKNCQHDFKLDEEIGILCQICGFVKTEIKYVSAPFMEHTGWTAESKP
QNEEDSELKPDEDEGSSLFGNHTSGEDVPVSEVNDNVWDLIPELRPKLHMHQKKAFEFLWKNTAGSLVPAHMEKTSKKIGGCVVSHTPGAGKTFLIIAFL
VSYLKLFPGKRPLVLAPKTTLYTWYKEFIKWEIPVPVHLIHGTRSSRAFKQTPAALRGSGPRPSQDVVHILDCLEKMQKWHAQPSVLVMGYTSFLTLMRE
DSKYNHRKYMAKVLRESPGMLILDEGHNPRSTKSRLRKVLMKVETDLRILLSGTLFQNNFCEYFNTLSLARPMFIKEVLKALDPKFKRKKKGAQKARHLL
ESRARKFFIDNIASKINSDEAEEKMQGLNMLRNMTNGFIDVYEGTASDTLPGIQIYTILMNPTDIQHQILVKLHKIMEKCPGYPLEVELLITLASIHPSL
VNSSVCVKKFYNLEELMELEKLRFDCKKGSKVMFVLNLVYRVVKNEKVLIFCHNIAPIKLFLELFENIFRWQQGKEILVLTGELELFERGRVMDKFEELG
GPSRVLLASITACAEGISLTAASRVILLDSEWNPSKTKQAIARAFRPGQQKMVYVYQLLATGTVEEDKYRRTAWKEWVSRMIFSEEFVEDPSRWQAEKIE
DDVLREIVEEDRVKSFHMIMKNEKASTS
|
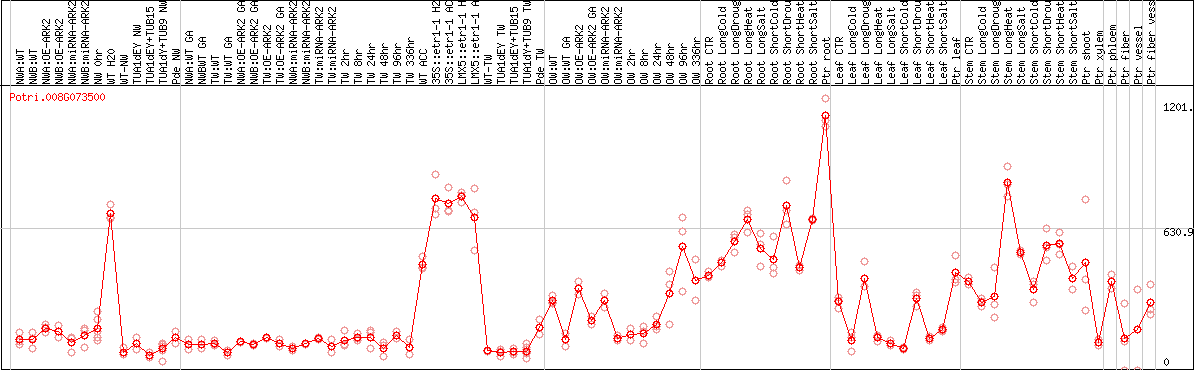
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G073500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
