Potri.008G077800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G077800.1 pacid=42806832 polypeptide=Potri.008G077800.1.p locus=Potri.008G077800 ID=Potri.008G077800.1.v4.1 annot-version=v4.1
ATGGCGGAAGTGGTCAAGATGGCAATCGACTGTGAAGATGAGTCAAAACCGAGCGGCCAAGATCAACCAAAGAAGACCCTGAAGAGGAAGCGAGCAACTT
TAACACCAACACAACAACAGCAACTCGTGAACTTAACAGGTGAACAGAAAGAAGTCCAGATTGAGGAATTGAAGAGAGAAATGGAAGGGCTATTTGGGTA
TTACAAAGAAACGATGAATCAGAAAATGGGTTTTGGGTTTGGTGTAGATTTGGGTGGAAGTGAGTGTATTAATGTGAATGGGATGGTGGGTCTGTTAATG
GAAGAGAGTGACATGTCGTTTTCGAAACTGGTAGAGGAAATATACGGGAAGTTAGTGAAAAAGAGTGGGAATTTGACTGTTGCAGTGGTGAAAAGTGCTG
TGTTGTTTGTTGGTCAAAGAATAACATATGGTGTGCCTAATGTAGATGCTGATGTTTTGGAGGATGAGACTCAAAGTTGTCTTTGGTGCTGGGAGACACG
AGATTTAAAGCTAATGCCGAAATCTGTCCGTGGTGCTTTGAAAATCAGGCGTATGTGCCGAGCAAAGATCCATGAGAGGATTACTGCTGTTTTCGCGATG
ATAACTGCATTACAGAAGTCAGAGACTGATGAAAATTACAAAAGTGATTTAATAAAATCATCAGGGAAGCTTGGCAAAGTTTTAAGAGAGGCAGACATTC
GTTTGTTAGTGGATGGCATGTTACAGAAGAATGGAGCAGATATGGCTGAAAAGCAAGTAAAAAGAGAAGAAAAATTGATCATCAAGCAATTGGAGAAAAA
TAAACGAGAAGAGGAAAAGGAGAAAAAAAGGATGGACCTTGAATTTCAAAAGGAAAAGAGACAAACTGAAAAAGAGCAAAAGCGACTGCAGGAGGAGGCA
GAAAAAGATGAACGGCGTCGTGAAAGGGAAGAGTTTGAAATGAAGAGACAACTGAAAAGGCAGCAAGAGGAAGCTGAGAAGGAACAGCGGCGCAAGGAGA
AGGAAGAAGCTGAGTTGAAAAGGCGAGTTGCTGTACAGAAGCAAGCTTCAATGATGGAACGCTTTCTAAAAAGAAGCAAAAGTAGTTCCCCATGTCAGAA
TGACCAGAGTTTAACTAAAGCAACCACTTCTGATTCCTCAAGCAAAAAGAGTAAAAGGATGGATGAAGCAGTTACTCAGTTAATGGACTGTGCTCCTTTG
TTAAATGATAACATTACAAGTGATGACATTCTCAAGTCACACTTGTCTTCCTGGTGTCATTTGGGTTGCTCTATCCGTTCCAATAGAAAACAGCATTGGA
GTATCCGGAGGAAGCCTAAGACTGGACTGTTTAAGGAACTTAAGCTAACTGCTATCAGGGACCCAACTCATGATGATGATTCAAGTGCAGAGAAGCTTGA
TAGTGGATGGGGAGATCAAACTTCTGATGACATATCGTGCATAGATGTTAGAAAATGCAATCGGAGGAAGCAATTGTTGCAGTTTGATAAGAGCCATAGA
CCTGCTTTTTATGGTATCTGGCCTAAGACAAGTCATGCTGTTGGACCACGCCATCCATTAAGGAGGGATCCAGACTTAGATTATGATGTTGACAGTGATG
AAGAATGGGAAGAGGAGGATCCTGGTGAAAGCCTCTCGGATTGTGATAAGGATGATGGAGAAGAAAGTTTAGAGGAAGAATATTCAAAAGCTGATGATGA
AGAAGAGAGTGAAGATGGTTTTTTTGTACCTGATGGGTATCTCTCAGAAAATGAGGGAGTACAACCTCACAGAATGGATGCTGATCCTTCAGTCGAGGAG
GCCAGAAGCTCGCCTAGTTGCAAGCAAGACTTAGAGAGCGAGGAGTTCTGTACATTGCTGAAACAGCAGAAATGTCTGAACAGTTTAACTGATAATGCCC
TTCGGAAAAATCATCCCATGATTGTATTGAATATAATGCATGAGAAGGATGCGTTGTTAGTGGCTGATGACCTAAGTGACATTTCTAAAGTGGAAAAAAT
GTGTTTGCAAGCACTTAGCATGCGAGCATTCCCTGGAGGACCACAAATGGAGATGTTCTTGGATGTGTCATCTGAAAATCATGATGCTTGCTTGTTAAAT
GCAAAGGCCAGTGCTACACGTATCCCAGCTGTGATAACTTTACAAGACTCGGATATGCCTATAGTTGTGTCTGTTATTCAGTCATGCTCGCAAAGTATGA
ATAAAGTGGTTGAATCACTGCAACAGAAATTCCCCACTGTCTCAAAGTTGCAATTAAGAAACAAAGTTCGTGAGATATCAGATTTTGTGGATAATCGTTG
GCAGGTCAAGAAAGAAGTTTTGGATGGTTTTGGCATTATAAGTTCGCCAGAAAAGAGCAGAGGAAGGAAGCATAACATTTCTACATTTTTCTCCAAGAGG
TGCCTGCCTCCTGCTGGTAAAAGCACGAATCCTAATGAGTCCTCACCGCCTATGCTGAAACATGGTTCTGTTGCTGAGAGCCAACAAATTTGCACATACA
GCCATCCATAG
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G077800.1 pacid=42806832 polypeptide=Potri.008G077800.1.p locus=Potri.008G077800 ID=Potri.008G077800.1.v4.1 annot-version=v4.1
MAEVVKMAIDCEDESKPSGQDQPKKTLKRKRATLTPTQQQQLVNLTGEQKEVQIEELKREMEGLFGYYKETMNQKMGFGFGVDLGGSECINVNGMVGLLM
EESDMSFSKLVEEIYGKLVKKSGNLTVAVVKSAVLFVGQRITYGVPNVDADVLEDETQSCLWCWETRDLKLMPKSVRGALKIRRMCRAKIHERITAVFAM
ITALQKSETDENYKSDLIKSSGKLGKVLREADIRLLVDGMLQKNGADMAEKQVKREEKLIIKQLEKNKREEEKEKKRMDLEFQKEKRQTEKEQKRLQEEA
EKDERRREREEFEMKRQLKRQQEEAEKEQRRKEKEEAELKRRVAVQKQASMMERFLKRSKSSSPCQNDQSLTKATTSDSSSKKSKRMDEAVTQLMDCAPL
LNDNITSDDILKSHLSSWCHLGCSIRSNRKQHWSIRRKPKTGLFKELKLTAIRDPTHDDDSSAEKLDSGWGDQTSDDISCIDVRKCNRRKQLLQFDKSHR
PAFYGIWPKTSHAVGPRHPLRRDPDLDYDVDSDEEWEEEDPGESLSDCDKDDGEESLEEEYSKADDEEESEDGFFVPDGYLSENEGVQPHRMDADPSVEE
ARSSPSCKQDLESEEFCTLLKQQKCLNSLTDNALRKNHPMIVLNIMHEKDALLVADDLSDISKVEKMCLQALSMRAFPGGPQMEMFLDVSSENHDACLLN
AKASATRIPAVITLQDSDMPIVVSVIQSCSQSMNKVVESLQQKFPTVSKLQLRNKVREISDFVDNRWQVKKEVLDGFGIISSPEKSRGRKHNISTFFSKR
CLPPAGKSTNPNESSPPMLKHGSVAESQQICTYSHP
|
DESeq2's median of ratios [POPLAR]
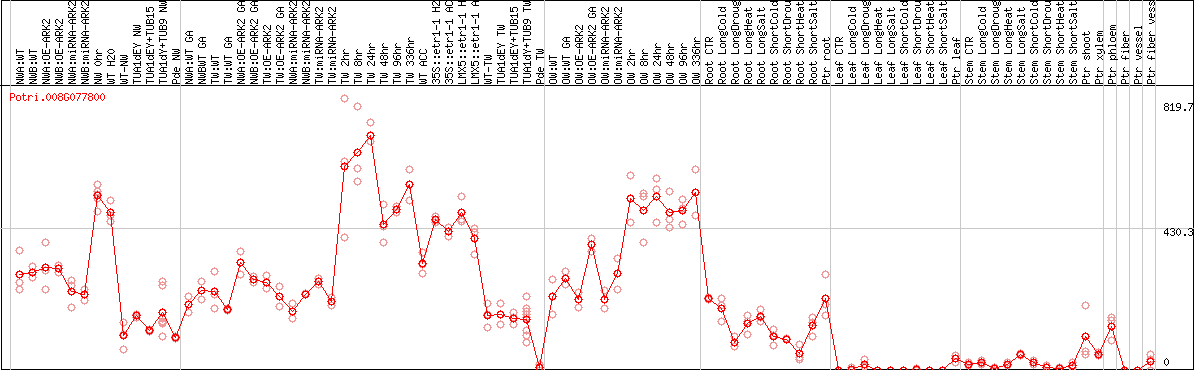
Coexpressed genes
Potri.008G077800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
