Potri.008G079300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G079300.1 pacid=42805992 polypeptide=Potri.008G079300.1.p locus=Potri.008G079300 ID=Potri.008G079300.1.v4.1 annot-version=v4.1
ATGGACGACGTTTTTTTCTCCCTTCACTCTGCATCTCCTCGAACCAACAGCGATGATGCTTCTTCGTCCTCACCTCCGCTCCGAACACCAAAGATCTTCG
ACCGTTACTTTTCTTCCTCCTCCTCACCATCCTCCTCCTCCGACGATGATCTACAGCCCTCCAATCCCAATCCCTCCCTGGAAGCCTCCACTAAACGCCT
GGATTACATGATCCAGTTCCTCGACCGAAAACTCTCCAATAACAACTGTAATAATAGCTCTAACAACAACGAAAGTGTTTCCCACAGACACAAAACTCCG
GCTCTACCAGAATTCATTGGCAAAGGAGGGGGAACCGGGATTTTCCGTATCCCAGTTCGAGCCGCGGTTCACCCAGACCGTCCACCCAGTCTTGAAATTC
GGTCACATCCGCTGAGAGAATCTCAAACCGGTCGGTTTTTGAGAACGATTGTAACTACCGAGACGCAAGTATGGGGTGGGAGAGAAAACGGCGCCGTTCA
GGTCTGGGAATTGAAGGAAATGTACGGCGGGAGTGATGAAACGGCGCCGTTCAAGGAGTCTGTGGCGTTGAATTCGGGTTCGGGCGTTACGTGTTTGGTT
GGTGATGAAGGGAGTCGAGTTGTTTGGAGTGGACATAGAGATGGTAGAATTCGGTGTTGGAAAATGGATACGGGTCCAGGGTTGGACCGAAGTCGTGTCA
AGGAGGTCTTGTCTTGGATGGCACATCGCGGGCCAGTCATGACCATGATCTTGACTTGTTATGGGGATTTGTGGTCGGGATCAGAAGGGGGGGTTATTAA
GATATGGCCCTGGGAAGATCTGGAAAAGGCCTTTTCATTTACCGCAGAAGAGAGGCATATGGCTGCACTGTCAGTGGAGAGGTCATATATTGATATAAGG
AACCAAGTGACCATGAATGGATTCAGCAATGTATTGAATTCCGACGTTAGATACTTGCTTTCTGATAATTCCAGGGCTAAAGTCTGGAGTGCTGGGTTCC
TGTCGTTCGCATTGTGGGATGCACACACTAGGGAGTTGCTGAAAATGTTCAATATAGATGGTCAGATTGAAAGACTGGACATGTTGTCAGGACAGGACTT
AACCTTTGAAGATGATATCAAGATGAAAATTGTTGCTGGTTCAAAGAAAGAGAAAATGCAAACTTCATTCGGTTTCTTTCAGCGTTCACGTAATGCTATA
ATGGGAGCAGCTGATGCTGTTCGTCGAGTTGCTGTAAAAGGTGGATTTGGAGATGATAATAGGAGAACAGAAGCGGTAATTATAACAACAGATGGAATGA
TTTGGACTGGGTGTGCAAATGGCTCGCTTGTGCAATGGGATGGAAATGGCAATCGTTTGCAAGATTTCCAGTATCATCCTGTTGCTGTTCAGTGCTTGTG
TACATTTGGTTTGCAAATATGGGTTGGTTATGCAAGTGGCACTGTTCAGGTGTTAGACCTTGAGGGTAATCTGGTTGGAGGATGGGTGGCTCACAGCAGT
CAAGTAATAAAAATGGCTGTTGGGGGTGGCTATGTTTTCACCCTGGCTAATCATGGCGGTATACGTGGATGGAATGTCATGTCTCCAGGACCCCTTGATG
GCATACTGCGTTCAGAGTTAGCTGGAAAAGAATTTTTGTATACAAGGATAGAGAATTTAAAAATATTGGCTGGTACATGGAATGTAGCACAAGGAAGAGC
TTCGCAAGACTCACTTGTGTCCTGGTTAGGTTCTGCTGCTGGGGACATTGGCATTGTTGTTGTTGGGTTGCAAGAAGTTGAGATGGGTGCTGGTGTTCTT
GCAATGTCTGCAGCTAAGGAAACTGTTGGACTGGAGGGCAGTTCTGCTGGGCAGTGGTGGCTGGATATGATTGGCAAAACATTGGATGAAGGGTCAACAT
TTGAACGTGTTGGTTCTCGGCAGCTAGCAGGCTTGCTGATTGCTATGTGGGTTAGAAATAATCTTAAAGCTCATGTCGGGGATGTTGATGCTGCTGCAGT
TCCATGTGGCTTTGGGCGTGCAATTGGTAATAAGGGTGCTGTTGGTTTGAGGATTAGGGTATATGATCGGGTAATGTGCTTCATTAATTGTCACTTTGCA
GCACATTTAGAAGCGGTCAATCGTCGCAATGCTGATTTTGACCATGTATATCGGACAATGACCTTTGGCCGACCATCTAACTTCTTCAATGCTGCAGCCG
CTGGCACTTTATCTGCAGTCCAAAATCCCTTGCGTCCAGAAGGAATACCCGAGTTATCAGAGGCAGACATGGTCATATTTCTTGGCGATTTTAATTATCG
GCTTGATGGTATATCTTATGACGAAGCAAGGGATTTTGTGTCTCAGAGAAGCTTTGATTGGCTTAGAGAAAAGGACCAACTCCGAACAGAAATGGGAGTT
GGAAAAGTGTTCCAAGGAATGCGCGAAGCAGTTATTAGATTTCCTCCCACATACAAGTTTGAAAAGCACCAACCTGGTTTAGCAGGGTATGATTCAGGTG
AAAAGAAGCGCATCCCTGCCTGGTGTGACAGGGTACTATATCGTGATAGCCGGTCAGCCCATGTATCTGAATGCTGCTTAGACTGTCCTGTTGTTTCTTT
GATATCCCAGTACGATGCTTGCATGGATGTAACCGACAGCGATCACAAGCCTGTTCGTTGCATATTCAGTGTTGATATTGCACGAGTTGATGAGTCAGTG
AGGAGACAAGAGTTTGGAGACATTATGAAATCAAATGAGGAAATTAGATACATAATTGATGAACTAAGCAAGATTCCAGAAACTATTGTCAGCACAAACA
ACATAATCCTTCCAAACCAGGATACAACAATTTTGCGTATCACAAATAAATGTGGGGAAAATGATGCTTTGTTCGAAATTATTTGTGAAGGCCAGTCTAT
AATTGATGAGAATGGACAAGCATCAGATCATCATCCAAGAGGTTCCTATGGCTTTCCGCAATGGCTTGAGGTCACTCCAGCAGCTGGTATAATTAAACCA
GGCCATATTGCCGAGGTGTCAATTCATCTTGAAGATTTCCCCACTTTGGAAGTTTTTCTTGATGGTGTCCCTCAAAATTCATGGTGTGAAGACACCAGAG
ACAAGGAAGCTATATTGGTGGTGAAAGTTCGTGGAACTTGCAACACAAATGAGACAAGAAACCACCGGATACGTGTGCGCCACTGCTGTTCATCGCAAAC
AGCACAACTTGACCCAAGACCGAATGGCTCGGAGCAAATCCAAGGAAATCTTCTTCACCGGGCTGATTACCAACATCTTAGCAGTTCCTATGATGTGGTC
AGCCACCTGCGTAACTTGCGTAGTCCATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G079300.1 pacid=42805992 polypeptide=Potri.008G079300.1.p locus=Potri.008G079300 ID=Potri.008G079300.1.v4.1 annot-version=v4.1
MDDVFFSLHSASPRTNSDDASSSSPPLRTPKIFDRYFSSSSSPSSSSDDDLQPSNPNPSLEASTKRLDYMIQFLDRKLSNNNCNNSSNNNESVSHRHKTP
ALPEFIGKGGGTGIFRIPVRAAVHPDRPPSLEIRSHPLRESQTGRFLRTIVTTETQVWGGRENGAVQVWELKEMYGGSDETAPFKESVALNSGSGVTCLV
GDEGSRVVWSGHRDGRIRCWKMDTGPGLDRSRVKEVLSWMAHRGPVMTMILTCYGDLWSGSEGGVIKIWPWEDLEKAFSFTAEERHMAALSVERSYIDIR
NQVTMNGFSNVLNSDVRYLLSDNSRAKVWSAGFLSFALWDAHTRELLKMFNIDGQIERLDMLSGQDLTFEDDIKMKIVAGSKKEKMQTSFGFFQRSRNAI
MGAADAVRRVAVKGGFGDDNRRTEAVIITTDGMIWTGCANGSLVQWDGNGNRLQDFQYHPVAVQCLCTFGLQIWVGYASGTVQVLDLEGNLVGGWVAHSS
QVIKMAVGGGYVFTLANHGGIRGWNVMSPGPLDGILRSELAGKEFLYTRIENLKILAGTWNVAQGRASQDSLVSWLGSAAGDIGIVVVGLQEVEMGAGVL
AMSAAKETVGLEGSSAGQWWLDMIGKTLDEGSTFERVGSRQLAGLLIAMWVRNNLKAHVGDVDAAAVPCGFGRAIGNKGAVGLRIRVYDRVMCFINCHFA
AHLEAVNRRNADFDHVYRTMTFGRPSNFFNAAAAGTLSAVQNPLRPEGIPELSEADMVIFLGDFNYRLDGISYDEARDFVSQRSFDWLREKDQLRTEMGV
GKVFQGMREAVIRFPPTYKFEKHQPGLAGYDSGEKKRIPAWCDRVLYRDSRSAHVSECCLDCPVVSLISQYDACMDVTDSDHKPVRCIFSVDIARVDESV
RRQEFGDIMKSNEEIRYIIDELSKIPETIVSTNNIILPNQDTTILRITNKCGENDALFEIICEGQSIIDENGQASDHHPRGSYGFPQWLEVTPAAGIIKP
GHIAEVSIHLEDFPTLEVFLDGVPQNSWCEDTRDKEAILVVKVRGTCNTNETRNHRIRVRHCCSSQTAQLDPRPNGSEQIQGNLLHRADYQHLSSSYDVV
SHLRNLRSP
|
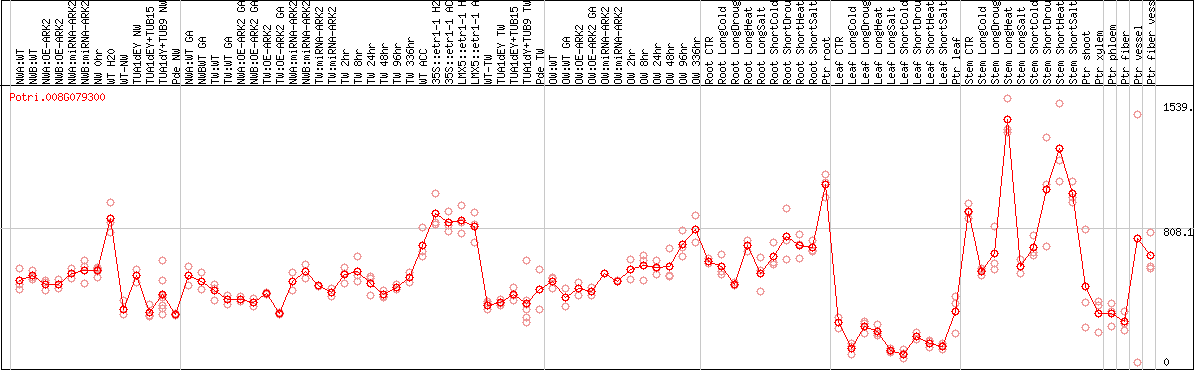
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G079300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
