External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12760 415 / 2e-148
unknown protein
AT1G15860 109 / 5e-29
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
AT3G28970 96 / 3e-23
AAR3
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G112200
112 / 4e-30
AT1G15860 267 / 4e-91
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Potri.003G180300
110 / 2e-29
AT1G15860 348 / 6e-123
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Potri.001G047900
102 / 5e-26
AT1G15860 328 / 3e-115
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Potri.004G122100
97 / 2e-23
AT3G28970 296 / 3e-100
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Potri.017G084300
91 / 8e-22
AT3G28970 269 / 1e-90
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000875
441 / 4e-159
AT3G12760 383 / 3e-136
unknown protein
Lus10009599
437 / 2e-157
AT3G12760 380 / 3e-135
unknown protein
Lus10013742
113 / 2e-29
AT1G15860 367 / 5e-129
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10039198
110 / 2e-29
AT1G15860 369 / 2e-131
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10019598
101 / 8e-26
AT1G15860 285 / 4e-98
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10040019
98 / 2e-24
AT1G15860 278 / 2e-95
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10030685
90 / 9e-21
AT3G28970 255 / 4e-84
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Lus10005235
89 / 2e-20
AT3G28970 258 / 1e-84
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Lus10026216
75 / 1e-16
AT1G15860 252 / 3e-86
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0220
EF_hand
PF03556
Cullin_binding
Cullin binding
CL0214
UBA
PF14555
UBA_4
UBA-like domain
Representative CDS sequence
>Potri.008G083800.3 pacid=42807080 polypeptide=Potri.008G083800.3.p locus=Potri.008G083800 ID=Potri.008G083800.3.v4.1 annot-version=v4.1
ATGCACAAGTTAAATAGAGGCAACCGTGAGAAAGTCCAGCAGTTCATGAGTATAACTGGGACAAGCGAGAAGGTTGCTGTTCAGGCTTTGAAGGCTAGTG
ATTGGCATCTAGAAGGAGCATTTGATGCGTTCTACAGCCAGCCTCAGTCTAGAACATATACTGATTCTAGACATTTGGAAGAGCTTTACAACAGATATAA
AGATCCTTATGTTGATATGGTACTGGTTGATGGTATCACCATCCTTTGCAATGATCTTCAGGTGGATCCTCAAGATATTGTTATGTTAGTTGTTTCATGG
CACATGAAGGCTGCCACCATGTGTGAATTCTCCAAGCAGGAATTCATTGGTGGATTACAATCACTGGGGGTCGATTCTTTGGACAAGTTTCGTGAAAAAA
TACCATATATGCGATCTGAGCTGATGGACGAACAGAAGTTTCGTGAAATATACAACTTTGCTTTTGGCTGGGCAAAAGAAAAGGGACAAAAATCGTTGGC
ATTGGATACAGCAATTGGTATGTGGCAACTGCTGTTTGCTGAAAAACAGTGGCCTTTGGTGGATCACTGGTGCCAGTTCTTACAGGCTCAGCATAATAAA
GCAATCTCGAGGGACACTTGGTCTCAACTTCTGGAGTTTGCCAGGACAGTAGACCCCACATTATCAAATTATGATGCTGAAGGTGCATGGCCCTATCTGA
TTGATGAATTTGTTGAATATTTGAATGAGAATGGCATAATGCAAAAGGGTCGGTCAACTGAATGGAGCCAAAAACGATGA
AA sequence
>Potri.008G083800.3 pacid=42807080 polypeptide=Potri.008G083800.3.p locus=Potri.008G083800 ID=Potri.008G083800.3.v4.1 annot-version=v4.1
MHKLNRGNREKVQQFMSITGTSEKVAVQALKASDWHLEGAFDAFYSQPQSRTYTDSRHLEELYNRYKDPYVDMVLVDGITILCNDLQVDPQDIVMLVVSW
HMKAATMCEFSKQEFIGGLQSLGVDSLDKFREKIPYMRSELMDEQKFREIYNFAFGWAKEKGQKSLALDTAIGMWQLLFAEKQWPLVDHWCQFLQAQHNK
AISRDTWSQLLEFARTVDPTLSNYDAEGAWPYLIDEFVEYLNENGIMQKGRSTEWSQKR
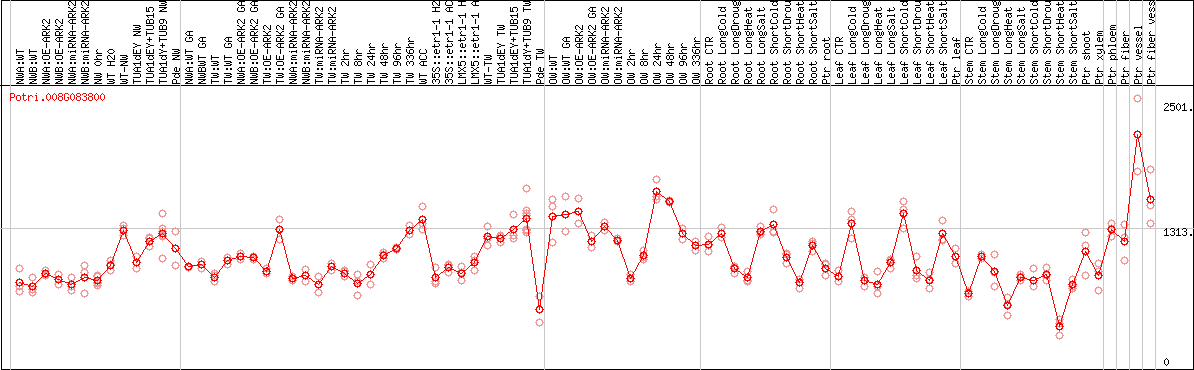
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G083800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.