External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G62280 117 / 4e-32
SLAH1
SLAC1 homologue 1 (.1)
AT1G62262 115 / 2e-31
SLAH4
SLAC1 homologue 4 (.1)
AT4G27970 78 / 2e-17
SLAH2
SLAC1 homologue 2 (.1)
AT5G24030 69 / 4e-14
SLAH3
SLAC1 homologue 3 (.1)
AT1G12480 50 / 9e-08
SLAC1, RCD3, CDI3, OZS1
SLOW ANION CHANNEL-ASSOCIATED 1, RADICAL-INDUCED CELL DEATH 3, OZONE-SENSITIVE 1, CARBON DIOXIDE INSENSITIVE 3, C4-dicarboxylate transporter/malic acid transport protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G026700
69 / 7e-15
AT5G24030 367 / 3e-124
SLAC1 homologue 3 (.1)
Potri.001G114300
53 / 1e-08
AT1G12480 432 / 4e-149
SLOW ANION CHANNEL-ASSOCIATED 1, RADICAL-INDUCED CELL DEATH 3, OZONE-SENSITIVE 1, CARBON DIOXIDE INSENSITIVE 3, C4-dicarboxylate transporter/malic acid transport protein (.1)
Potri.002G227800
50 / 6e-08
AT1G62280 120 / 7e-31
SLAC1 homologue 1 (.1)
Potri.010G170050
0 / 1
AT1G62280 136 / 7e-39
SLAC1 homologue 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024306
130 / 5e-37
AT1G62280 349 / 7e-119
SLAC1 homologue 1 (.1)
Lus10015133
110 / 4e-29
AT1G62280 387 / 6e-133
SLAC1 homologue 1 (.1)
Lus10031545
87 / 4e-22
AT1G62280 160 / 3e-48
SLAC1 homologue 1 (.1)
Lus10039312
70 / 1e-14
AT5G24030 670 / 0.0
SLAC1 homologue 3 (.1)
Lus10027553
70 / 1e-14
AT5G24030 669 / 0.0
SLAC1 homologue 3 (.1)
Lus10006678
52 / 3e-08
AT1G12480 734 / 0.0
SLOW ANION CHANNEL-ASSOCIATED 1, RADICAL-INDUCED CELL DEATH 3, OZONE-SENSITIVE 1, CARBON DIOXIDE INSENSITIVE 3, C4-dicarboxylate transporter/malic acid transport protein (.1)
Lus10007023
49 / 2e-07
AT1G12480 720 / 0.0
SLOW ANION CHANNEL-ASSOCIATED 1, RADICAL-INDUCED CELL DEATH 3, OZONE-SENSITIVE 1, CARBON DIOXIDE INSENSITIVE 3, C4-dicarboxylate transporter/malic acid transport protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03595
SLAC1
Voltage-dependent anion channel
Representative CDS sequence
>Potri.008G084901.1 pacid=42807618 polypeptide=Potri.008G084901.1.p locus=Potri.008G084901 ID=Potri.008G084901.1.v4.1 annot-version=v4.1
ATGCTCAACTTGGTTGGTGCTCAAGCTGCAGCTAATATGGGGTGGAAAGAGTGCTGTTTTCTTGTTTTCACTTGGCACGGTCCGTTACTTGGTGCTGTTT
CCGGTGGTGGTAGGCTTCCAGCTATGTCAAGGCCAGTTTTTTTCTTGTTCATTGCAGCACCAAGTAGGGAAAGCTTGGCTTGGGAATCCATAAATGGAGC
TTTTGATACTGCATCAAAGATGCTCTTCTTTCTCACCCTCTTCCTGCTCACTTCTCTGATCTGTAGACCTAATCTCTTCAAGAGATCCATGAGAAGGTTC
AGTGTTGTGTGGTGGGCTTACTCGTTTCCATTGACAGTCGTTGCTAGCTCTCGCTTCAAGGGATTATGCAGAAGAAGTGAAAGGGAGGATCGCAAATGCT
ATAATGCTTCTCCTTTCCGGGCTCGCCATCTTGGCCTCGCTTAG
AA sequence
>Potri.008G084901.1 pacid=42807618 polypeptide=Potri.008G084901.1.p locus=Potri.008G084901 ID=Potri.008G084901.1.v4.1 annot-version=v4.1
MLNLVGAQAAANMGWKECCFLVFTWHGPLLGAVSGGGRLPAMSRPVFFLFIAAPSRESLAWESINGAFDTASKMLFFLTLFLLTSLICRPNLFKRSMRRF
SVVWWAYSFPLTVVASSRFKGLCRRSEREDRKCYNASPFRARHLGLA
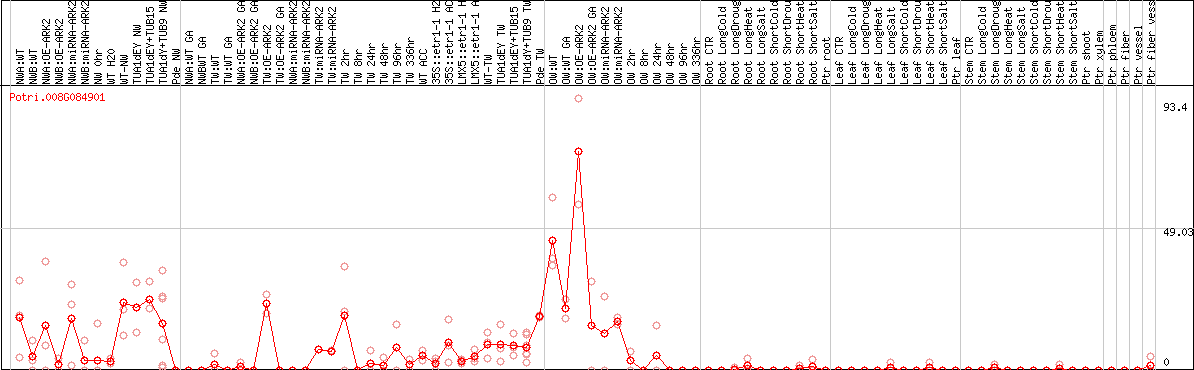
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G084901 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.