External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69400 334 / 1e-114
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT1G49910 216 / 3e-68
BUB3.2
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.2, Transducin/WD40 repeat-like superfamily protein (.1)
AT3G19590 208 / 7e-65
BUB3.1
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.1, Transducin/WD40 repeat-like superfamily protein (.1)
AT1G80670 102 / 9e-25
RAE1
RNA export factor 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G08390 52 / 3e-07
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G23430 49 / 3e-06
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT1G61210 45 / 7e-05
DWA3
DWD hypersensitive to ABA 3, Transducin/WD40 repeat-like superfamily protein (.1.2)
AT3G49660 43 / 0.0001
AtWDR5a
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
AT1G11160 44 / 0.0002
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G15550 42 / 0.0006
Transducin/WD40 repeat-like superfamily protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G162700
556 / 0
AT1G69400 350 / 7e-121
Transducin/WD40 repeat-like superfamily protein (.1.2)
Potri.001G295100
211 / 6e-66
AT3G19590 593 / 0.0
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.009G089200
207 / 2e-64
AT3G19590 602 / 0.0
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.003G180000
93 / 3e-21
AT1G80670 590 / 0.0
RNA export factor 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.001G048100
93 / 3e-21
AT1G80670 583 / 0.0
RNA export factor 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.010G255400
49 / 2e-06
AT5G08390 927 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.008G002300
48 / 6e-06
AT5G08390 1006 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.001G306500
46 / 1e-05
AT5G64730 489 / 2e-176
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.005G148500
45 / 3e-05
AT3G49660 503 / 0.0
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014343
389 / 6e-136
AT1G69400 313 / 6e-106
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10026051
388 / 6e-136
AT1G69400 318 / 9e-109
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10003954
202 / 2e-62
AT3G19590 618 / 0.0
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10023814
196 / 3e-60
AT3G19590 610 / 0.0
BUB \(BUDDING UNINHIBITED BY BENZYMIDAZOL\) 3.1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10026095
103 / 7e-25
AT1G80670 632 / 0.0
RNA export factor 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10002323
102 / 3e-24
AT1G80670 608 / 0.0
RNA export factor 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10030869
49 / 2e-06
AT1G10580 939 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10030620
49 / 3e-06
AT1G10580 926 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10041007
49 / 4e-06
AT5G23430 721 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10017157
48 / 7e-06
AT5G66240 541 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2.3)
PFAM info
Representative CDS sequence
>Potri.008G092100.2 pacid=42807320 polypeptide=Potri.008G092100.2.p locus=Potri.008G092100 ID=Potri.008G092100.2.v4.1 annot-version=v4.1
ATGAAAGGCGCGTGTTTAAATTTCGAAAACCCCATCGAAGATGCAGTTTCCAGGATCCGATTCGCCCCTCAATCCAATAATCTCCTCATTTCCTCTTGGG
ATTCCAATCTTCGATTATACGATGTGGATAGCTCTCTGCTTAGATTAGAAGCTCCTGCTCCCTCACAAGCTGCTCTTCTCGACTGTTGTTTCCAGAGTGA
GTCCGTTGCTTTTACGGCTGCGTCCGATGGTTCCATTACAAGGTATGACTTGCACTCAGGAACTATTGATGCTATTGGAAGTCATCAGGACATGGCAACA
TGTGTGGGATACTCTATTGAAACATGCCAAGTGATTTCAGCTGGATTAGACAAGAAGGTAATGTCTTGGGATATGCGCTTGGCAAATCCTCTTGCTCTCT
TCCGAAATCTGGGTGCAGAAATTGATTCCATTTCTATCTCCGGTTTTGATTTGATGGTAGCTGTTGGAGCAGCAGTAAATATATATGACTTGCGGAACTA
TGAAAGAGCGGTGGATTTAAAAGAACTATCTATGGATGTAGGGATAAGTTGTGTTGCTTCAGTTCCATTTACCAGAGGATATGCAATTGGGTTGATAGAT
GGACGTGTAGCGCTGGAGATTTCAAATCCATTAAATTCGAATTCCACTGGATACGCTTTTCGGTGTCACCCTACAACAAAGGATGGAACGGCACATCTTG
TGTCAGTTAATGACATAGTTTTCAATCCACACATTGGTGGTACTTTTGTTACTGGCGATAATGAAGGCTATGTTACTGCATGGGATGCTAAAAGTAAAAG
AAGATTACACGAGTTTCCTAGATATCCAAATAGTGTGGCATCCTTATCATACAACCATGTGGGGCAACTTCTAGCTGTTGTATCAAGCTACACATATCAA
GAAGCTAATGAAAGTTCAGGTTCTCTTGAGCGTGAAAGGACAAGTCAGGCCTCGTGCAAGTCTTAA
AA sequence
>Potri.008G092100.2 pacid=42807320 polypeptide=Potri.008G092100.2.p locus=Potri.008G092100 ID=Potri.008G092100.2.v4.1 annot-version=v4.1
MKGACLNFENPIEDAVSRIRFAPQSNNLLISSWDSNLRLYDVDSSLLRLEAPAPSQAALLDCCFQSESVAFTAASDGSITRYDLHSGTIDAIGSHQDMAT
CVGYSIETCQVISAGLDKKVMSWDMRLANPLALFRNLGAEIDSISISGFDLMVAVGAAVNIYDLRNYERAVDLKELSMDVGISCVASVPFTRGYAIGLID
GRVALEISNPLNSNSTGYAFRCHPTTKDGTAHLVSVNDIVFNPHIGGTFVTGDNEGYVTAWDAKSKRRLHEFPRYPNSVASLSYNHVGQLLAVVSSYTYQ
EANESSGSLERERTSQASCKS
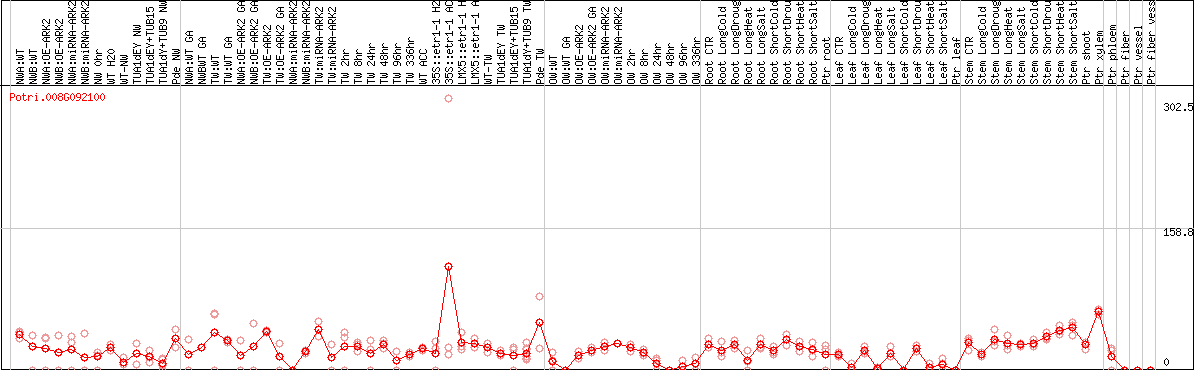
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G092100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.