External link
Symbol
CRC.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69180 155 / 1e-48
YABBY
CRC
CRABS CLAW, Plant-specific transcription factor YABBY family protein (.1)
AT2G26580 111 / 2e-31
YABBY
YAB5
YABBY5, plant-specific transcription factor YABBY family protein (.1.2)
AT1G08465 104 / 2e-28
YABBY
YAB2
YABBY2, Plant-specific transcription factor YABBY family protein (.1)
AT2G45190 104 / 6e-28
YABBY
FIL, YAB1, AFO
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
AT4G00180 103 / 1e-27
YABBY
YAB3
YABBY3, Plant-specific transcription factor YABBY family protein (.1.2)
AT1G23420 84 / 3e-20
YABBY
INO, YAB4
INNER NO OUTER, Plant-specific transcription factor YABBY family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G154350
234 / 2e-79
AT1G69180 155 / 2e-48
CRABS CLAW, Plant-specific transcription factor YABBY family protein (.1)
Potri.009G000100
116 / 3e-33
AT1G08465 215 / 3e-72
YABBY2, Plant-specific transcription factor YABBY family protein (.1)
Potri.018G129800
116 / 7e-33
AT2G26580 250 / 5e-86
YABBY5, plant-specific transcription factor YABBY family protein (.1.2)
Potri.001G120200
115 / 4e-32
AT2G45190 236 / 4e-79
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Potri.002G145100
113 / 1e-31
AT2G45190 265 / 2e-90
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Potri.016G067300
110 / 4e-31
AT1G08465 150 / 2e-46
YABBY2, Plant-specific transcription factor YABBY family protein (.1)
Potri.003G112800
109 / 4e-30
AT2G45190 243 / 1e-81
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Potri.006G067800
108 / 4e-30
AT2G26580 253 / 5e-87
YABBY5, plant-specific transcription factor YABBY family protein (.1.2)
Potri.014G066700
108 / 7e-30
AT2G45190 277 / 3e-95
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036811
187 / 6e-61
AT1G69180 187 / 4e-61
CRABS CLAW, Plant-specific transcription factor YABBY family protein (.1)
Lus10000644
172 / 8e-55
AT1G69180 177 / 5e-57
CRABS CLAW, Plant-specific transcription factor YABBY family protein (.1)
Lus10024603
103 / 2e-27
AT2G45190 190 / 2e-60
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Lus10032240
100 / 2e-26
AT2G45190 193 / 1e-61
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Lus10029135
92 / 5e-23
AT1G23420 200 / 9e-65
INNER NO OUTER, Plant-specific transcription factor YABBY family protein (.1)
Lus10043264
70 / 4e-14
AT5G35410 630 / 0.0
SNF1-RELATED PROTEIN KINASE 3.11, CBL-INTERACTING PROTEIN KINASE 24, SALT OVERLY SENSITIVE 2, Protein kinase superfamily protein (.1)
Lus10036496
68 / 4e-14
AT2G45190 183 / 2e-58
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Lus10019407
68 / 5e-14
AT2G26580 228 / 8e-77
YABBY5, plant-specific transcription factor YABBY family protein (.1.2)
Lus10010361
67 / 1e-13
AT2G45190 171 / 4e-53
YABBY1, FILAMENTOUS FLOWER, ABNORMAL FLORAL ORGANS, Plant-specific transcription factor YABBY family protein (.1)
Lus10008374
65 / 4e-13
AT1G08465 84 / 9e-21
YABBY2, Plant-specific transcription factor YABBY family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0114
HMG-box
PF04690
YABBY
YABBY protein
Representative CDS sequence
>Potri.008G097800.1 pacid=42808350 polypeptide=Potri.008G097800.1.p locus=Potri.008G097800 ID=Potri.008G097800.1.v4.1 annot-version=v4.1
ATGAACCTAGAAGAGAAAGCAAGCACAGAGGTTCCATCATCTGAGCATCTCTGCTATGTCCGTTGCAACTTCTGCAACACTGTTCTTGCGGTCGGGATTC
CATGCAAGAGGCTGCTGGACACTGTGACTGTCAAATGTGGTCATTGCAATAATCTCTCCTTTCTCAGCACCAGACCTCCCAACCAAGGGCAATGTCTTGA
TCAGTACCACCGACTGAGTCTACAGGGAGTTTCTAGTAATGAAAAGTTCTTATTCAATGAGAAGCAAGGGTTTTGCACTGATATCAGAAAGGGCGAATCC
TCATCCTCCTCGACCTCTAGCGAACAACCGGTGCCCACAGTACCCTTCGTAGTAAAGCCGCCTGAGAAGAAACACCGACTTCCATCTGCTTACAATAGGT
TTATGAAGGAGGAGATAAAGCGCATCAAAGCAGCCGATCCTGAGATACCACACAGAGAAGCTTTTAGCACAGCAGCTAAAAATTGGGCTAGGGCTAGGTA
CCTTCCAAAGTCGGGTGCTGGTTCTGGGAGCCCTAATATTTAA
AA sequence
>Potri.008G097800.1 pacid=42808350 polypeptide=Potri.008G097800.1.p locus=Potri.008G097800 ID=Potri.008G097800.1.v4.1 annot-version=v4.1
MNLEEKASTEVPSSEHLCYVRCNFCNTVLAVGIPCKRLLDTVTVKCGHCNNLSFLSTRPPNQGQCLDQYHRLSLQGVSSNEKFLFNEKQGFCTDIRKGES
SSSSTSSEQPVPTVPFVVKPPEKKHRLPSAYNRFMKEEIKRIKAADPEIPHREAFSTAAKNWARARYLPKSGAGSGSPNI
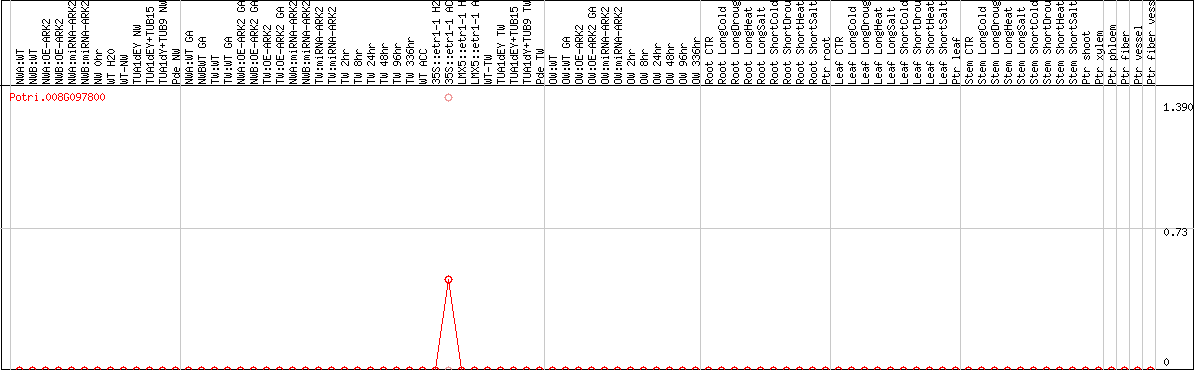
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G097800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.