Potri.008G109100 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G109100.6 pacid=42807673 polypeptide=Potri.008G109100.6.p locus=Potri.008G109100 ID=Potri.008G109100.6.v4.1 annot-version=v4.1
ATGAAAGGGAAGGGAGTATCGAGAGATTATTCTCGACGTAGAATTCGATCTGGAAAGAGAACGACAAGGAGTCTTGATAATATAGTTGTGATTGATGTCG
ATAGTGATGATGAGTTTGATAATGTTATTATCATTGATGTCCCGGAGTCTTTACAGCAGAAATTAAGAGGTTCTAATGTTGTTAGAGAAGGAAGAAGTTT
TCCGTGCATAATAAGCATTGATGATGATGATGATACAGTGGATGATCATGAAATTAATGCGCAAGGTGATGGTAACTTGGATAGTGATGGCACTTCTAGT
CACAGCAGTCCTGCCTCTGATTGTATAGGGAAATCTGTTTATAGGGATGCTGATGGTTGTCGAGTTGCTGAGGAGAATAGGCCTGTGTTCAAGTTGAGAA
AGTGCAATAGAACTTATCCTGAGAAAGCTCCCTCCAGAAAACGTTATGGTTTAGATTCAGATTCCGAGAGTGATTCGTCTGAAGATAGCACCTCTGACTG
TGAAGTCATGGAAGGATCTTTTGGAGAGGTTCGAGAACAGTGGGAAAAGGCTTCCCTGAAGAGAAAATCCAAGTCCTGTAAAGGTCTGGATGACCAGGCT
AGTCCATGCAGTTCCCACGGTGATGTTCATCCAAATGCTGAAGTAGAAAACAGAACTAAACAAAATCCAGATCCTTCTGTTTGCTCTAGTTCAAAAAATG
TAAATTTTGAGAAAGTGAACACTTGTGCCTCCACTTCCACTAGAGATGGAGTTCTGGGTGGTTGTTCTTCCAGTGCCAAGATGGAAAATCCTTTTGCAAA
TTATAACCAGAAAGGTGAAAGCTTTTCAAGGCCACAGAAATCAAGAACAAATGAGAATATTCATTTTCATTGGAAGAGCGATGATCTTTGTGGAGGAGAG
AGATCCATGGATGATAGTAGTACTTCCTATAACAAGTTTCGAACTTTAAATGGCCTTGGCACTAGGTTTCCTCCAGGGCCTTCTTCATGGTCCAATCAAG
AAAAGGATGATAAACAGTATCATCATAGAAGAGCATGTTTTCAGGATATGGAACAAAACACTGCGACAGGGCATTCTTTTCCAAATGACCAAAGTGGACC
TAACTTGCACTCAGATGATGGGAAAGCTAGTGTTTTGAATGAAGATGCTTCTTTACCTGATGGTCACTTTCTCGGTGAGAAACATGATGTTATTAACAGT
CAAGTTGATTCAAAGGAGGAATATAAAGAATTTACTCAGGTGCCATCTAGTTGCAAGATTCTTTCAAATGAGGCACAATGCAGAGAGAAGTTTGTGTCTT
ATGCTCGATCATCTGAGGACAAAGTGGTTGAAAATGTTATAGCTTCTTCATGTACTACACAAGAAGTCAGTGATGAGAAATCTGGTCATCAGAAAATGGT
TGAAAGAGCTGCCAGGGAAAAGTCTTCACAATGTCACGACAGACTAGGTAGGCCAGGAACATCAAATTCTGCAGAAGGAAAGGAGGTGTGTACTGATTTT
GCATCCAGCAGTCAACTTCATCACGAAAGGGATCCGTTATGTGCTTTACCTGGTGCTCGTTTTCCTTATGCTGTAAAGGATATAATCAATGACCGAGAAA
AGCTCAAGGAGACTGAAGAATACAAACAAGCAATGGAAGAAGAATGGGCTGCTAGACAGCAGCAGCTACAGATTCAGGCAGAAGAGGCTCAGAGATTGCG
GAAGAGAAGAAAAGCTGAAACTTTGCGTATTTTGGACATGGAGAGAAGGCAAAAACAGCGTGTGGAAGAAATGAGAGAGACACAAAAGAAGGATGAAGAG
AATTTGAACATCAAGGAGAGATTTCGTGTTGAAGTAAGGAAGGAGCTTTATAGATTGGAGGTTACCTGCATTAACATGGCATCCTTACTTCGTGGCTTGG
GAATCCATGTGGAAGGTGGTTTCCAACCCTTGCCAAATCAGGTGCATGCAGCTTATAAGCGAGCCTTGCTAAAATTTCATCCTGACCGCGCGTCAAAAAC
TGACATTCGTCGGCAAGTTGAGGCTGAGGAAAAATTTAAGCTTATTTCCCGCATGAAGGAGAAATTTTTGTCCACTTCATGTTACTAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.008G109100.6 pacid=42807673 polypeptide=Potri.008G109100.6.p locus=Potri.008G109100 ID=Potri.008G109100.6.v4.1 annot-version=v4.1
MKGKGVSRDYSRRRIRSGKRTTRSLDNIVVIDVDSDDEFDNVIIIDVPESLQQKLRGSNVVREGRSFPCIISIDDDDDTVDDHEINAQGDGNLDSDGTSS
HSSPASDCIGKSVYRDADGCRVAEENRPVFKLRKCNRTYPEKAPSRKRYGLDSDSESDSSEDSTSDCEVMEGSFGEVREQWEKASLKRKSKSCKGLDDQA
SPCSSHGDVHPNAEVENRTKQNPDPSVCSSSKNVNFEKVNTCASTSTRDGVLGGCSSSAKMENPFANYNQKGESFSRPQKSRTNENIHFHWKSDDLCGGE
RSMDDSSTSYNKFRTLNGLGTRFPPGPSSWSNQEKDDKQYHHRRACFQDMEQNTATGHSFPNDQSGPNLHSDDGKASVLNEDASLPDGHFLGEKHDVINS
QVDSKEEYKEFTQVPSSCKILSNEAQCREKFVSYARSSEDKVVENVIASSCTTQEVSDEKSGHQKMVERAAREKSSQCHDRLGRPGTSNSAEGKEVCTDF
ASSSQLHHERDPLCALPGARFPYAVKDIINDREKLKETEEYKQAMEEEWAARQQQLQIQAEEAQRLRKRRKAETLRILDMERRQKQRVEEMRETQKKDEE
NLNIKERFRVEVRKELYRLEVTCINMASLLRGLGIHVEGGFQPLPNQVHAAYKRALLKFHPDRASKTDIRRQVEAEEKFKLISRMKEKFLSTSCY
|
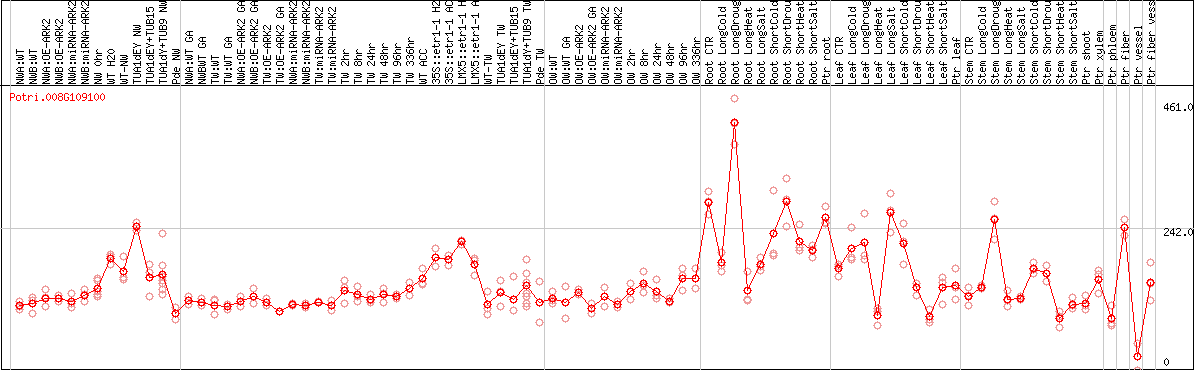
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G109100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
