Potri.008G128800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G128800.3 pacid=42807934 polypeptide=Potri.008G128800.3.p locus=Potri.008G128800 ID=Potri.008G128800.3.v4.1 annot-version=v4.1
ATGTGTAGTGAGTGTCATTCAAACAATAACCAGTTTTTTAATGGATACCATTGCCAAAGTTGTGGCAAGTGGTTGTGTTTCAATTGTATGAGAGGTTACC
AATCAAATGGTGATTTTGGGGAGGCTATCAAATCTTGCAAGTTCTGTAATGGGGTTACTGTGAAACGTGATGGTGGAAGGAAGAATAGTGATAAAGTACA
CCCTACTGACTCTCCTAGAGGTAGCCCTGAGCCACCATCACCTTCTTTTAGTGCCGAGCCAATCCATAGTGATCGTCTTCCCCTTTATCTTGAATCCAGG
GATTGTGGGTTTTCTCCTAATGCAATAACCACCAGGAGTATGACTTCTTTTAGTGCTCACCCGTCTCCTGTTTCTGTTCGTCGCTCATCAAGCAGGAGTG
ATGAAGAAGAAGCAGAGGATTCTGGGAAACTCCTTTACAGCCCATCGAGTGAGTACTGTCATGATATTTCAGATATAGATTCAAGTAGTGTTAGTGCTAG
GCTTGAGTTTTACAATTGTAAGACGGTGGGATCAAGTCCTTTAGACAGCCCTTCTCGGATTGATTTTAGTTCTTGCAGAGTTGGGCATACTGTACAGCAG
GGGCGAGAAGGAAGCCCATTGTCTCAAAGTGATGGTCCCTTTGATCAAGAAAATATGGCTATTTTAAGCAGGCCTGACAAAAGGACTGAGGATCCAGAGA
ATACTGATGATTGCTCTGATGACGGGTCAGTTTTGAGGGATCAATATCACAAGTCTCCAAAACCATTGGATTTTGAAAGCAATGGCCTTATCTGGTTTCC
CCCACCTCCTGAGGATGAAAATGATGAGGAGGAGAGTAATTTCTTCACATATGATGATGAAGATGATGATATTGGGGACTCTTCTGCAATTTTCTCATCG
AGTAGTAGCCTCTCTAGCACATTTCCATCAAAGGAGAAACAGAATAAGATTAACAAGGACCCTACTAAAGCTATGATACAGGGGCATTTTAGGGCTCTTG
TGGCGCAGCTTTTACAGGGAGAGGGTATCAAAGCCAGCAAGGATGAAAACAATGGAGAATGGCTTGACATAGTCACAGCAATTGCATGGCAAGCTGCAGC
TTTTGTGAAACCAGATACCAGCAGAGGAGGTAGTATGGATCCGGTTGATTATGTAAAGGTTAAATGTATAGCATCAGGAAATCCAAGAGATAGCACCCTT
GTAAAGGGAGTAGTTTGTACAAAAAATATAAAGCACAAGCGCATGACCACACAATACAAAAATCCCAGATTACTTCTTTTAGGAGGAGCACTCGAATATC
AGAGCGTTGTGAATCAGTTGGCTTCTTTTAATACATTAGTGCAACAGGAAAATGATCATCTCAAGCTGATCATGTCGAAGATAGAGGCTCTTCGCCCAAA
CGTTCTGCTGGTTGAGAAAAGCGTATCTCCATATGCCCAAGAATATCTACTGGGAAAGGAAATTTCCTTGGTGCTGAATGTAAAAAAGCCTTTACTGGAA
CGTATAGCTCGGTGCACAGGTGCTCAGATTAGTCCATCGTTTGAGAATATTTCTACAACACGACTGGGACATTGTGAACTATTCAGGGTAGAGAGAGTGA
GTGAAGAACATGAGACAAGCAATCAGTTCAACAAAAAGCCATCTAAAACATTGATGTCTTTTGAAGGGTGTCCAAGGCGTTTAGGTTGCACGGTCCTTCT
GAGGGGCACATGTCGTGAAAAGCTAAAGAAGGTCAAGCATGTTATTCAATATGCAGTTTTTGCAGCCTATCACTTGTCCCTTGAGACTTCTTTCCTTGCC
GATGAGGGTGCAAGTCTTCCTAAGATGACAATTAGACCCTCAATTGCTATTCCTGAGAGAACAGCAGCAGATAATTCCATTTCAGTTATCCCTCCAATGA
TTTGCCATGCTGAAGTTGCTTTGTCTGCTCAGGATGATGGATCACTGGGTCTTAAACCAGAGCATGAAGGATCAGAATCATTGACGGGGAACCTTGATGC
TGGTGTCATTCATCCTTTATCTCCATGTTCCGTCACTTGTAGGTCTGGAAATGAATTCTCTATTGCATGCCATGGCGATTTAGTGTCTAATGCGGGGGGA
TTGGATGCATTTTCTGCAAGTCAATGTGAGGGTCTAAAGATGTTTGCTGTTTCCCCTGGTATCAAAAATCTTTCACAGCCAGAACTACAAGATATCATGG
CTGAAGAAGAGGGACAACTTTTGGCGACTCATGAATCAGTACAATCTGAGAAGATTGATGAAGATGAGGTTTCCAGTGAATATTTCTCAGTCACTGACAC
ATACCAGAGCATATTAGTATCATTTTCAAGTCGCTGTGTGCTGAAAGGAACTGTATGCGAACGCTCTCGGCTTCTGCGCATAAAATTCTATGGAAATTTT
GATAAGCCACTTGGAAGATATCTTCGTGATGATCTGTTTGATCAGAAATCATGCTGTAGGTCATGTAAGGAGCCAGCCGAAGCACATGTTCTGTGTTTCA
CTCATCAGCAGGGTAATCTTACAATCAATGTTAGATCCCTTTCCTCTGTAAAACTACCTGGAGATCGTGATGGTAAGATATGGATGTGGCACCGATGCCT
AAGATGTGCTCATATAGATGGAGTTCCACCAGCAACTCGTAGAGTTGTCATGTCAGATGCTGCCTGGGGACTTTCTTTTGGGAAGTTTTTGGAACTGAGC
TTTTCAAACCATGCAACTGCCAATCGTGTTGCACCCTGTGGTCATTCATTGCAGAGGGACTGTCTTAGATTCTATGGGTTTGGGAGCATGGTTGTGTTCT
TCCGGTATTCCCCTATTGATATTCTCAATGTCCATTTACCGCCATCAATGCTTGAATTCAACGGCATTGTTCAGCAAGAGTGGACAAGGAAAGAGGCAGC
AGAGCTCTTGGGAAAAATGGAAACCTTCTATGGTGAGATATTTGGTGTGCTTGACAGCATGGAACAGAGAAGTAAATATTTTGGAAGTGAATTGTCAGAT
ACCAATGAGTTACAGAATCGCATCATGGAGCTGAAAGATCAACTCGTAAAGGAAAAAAACAATTACAGTGGCATATTACAACTGGCAGTTATGGAGAGTT
TACAGTTGGATCAAACAGCCATGGACATTCTAGAACTCAATCGCTTAAGACGTACTCTTCTAATTGGTTCTCATGTTTGGTATCGGAAGCTTTATTCACT
GGACTGTCTCCTTAAGACAAACTATCTTGTCAAGGCTAAAGAAGGGGATGTGTCTTATACTGAGCTGAAAGATTTGAAGAACGATATATTTTGCAAGGAT
AGCAAGCTTGACCATGATCATGAAGAAAATATCTCTGGCTATTCAAAATCACAGGAACATGTAGGGAATGATTTTCAGTCAGAGAAGAAAGAAACAGGTC
TGTCCTTCGAACATTGTGTTCTTGAACATTCAATGTTGCCATCATGTTATCATAACACAGAGGATGAAGTGCATGCAGGTGAGGAAACCGCGAGTAAGAC
ATTATTTAGTGACAACCCTTCCCATGCTTCCAATCTGTCTGACAGAATTGATTCTGCATGGACGGGTACCGATCAACTTCCAATTAAAGTTCAACCTCCG
CATGCATCTCAGGCAGAGGCAGATGGATTCCAACCTGTGTCTGTCAGGCAGCCAAATTTGTTTGATAATCCTCCTTTTCGAAGGATGGTGGCACCAAAGA
GAGTTCATTCTTTTGATTCTGCATTGAGAGCCCAAGAAAGAATCCAGAAAGGGTTGCCTCCATTGCATTTGTCGACTATCAGATCATTCCATGCATCTGG
AGATTACAGGAGTATGGTGAGAGATCCTGTTTCAAATGCAATGAGGACTTATTCCCAGACATTGCCACTGGAGGCACATAAGTTGAATTTAATGCACAGT
TCCACACACTCATTTATCTCCTCCGCAGCTAATATGGCTGGAGGAGCTCGGTTATTGTTACCAGTGAGGGCCAACAGTGACTTAGTTATAGGCGTGTATG
ACAACGATCCTGCAAGTGTAGTTTCCTATGCCCTCAGTTCTAAGGAACATGAGGATTGGGTTACTGATAGGTCAAATGAGAGTGCAGGAATCTGGAGTAC
AATTAAGCACAGTAAAGAAGATTCTGCAGCTTCCTCTTTTACATCCTGGCAGTCTTTAGATTCCATGGACCTAGATTATATGAGCTATGGAAGTTATGGG
TCTGAAGATCCTTTTTCAACCTTGGGCACCTTGTTTATGGATTCAAAAAAATCTCCACACTTGACAATTTCTTATGAAGATGCTTCTTCAATTGCTGAAG
GCAAAGTGAGGTTTTCTGTTACTTGTTATTTTGCGAAGCAGTTTGACTTTCTTAGGAAGAAATGCTGCCCTAGTGATGTTGATTTTGTTCGTTCACTGAG
TCGCTGCCAGAAGTGGAGTGCACAAGGAGGAAAAAGCAATGTATACTTTGCCAAGTCATTGGATGAGAGATTCATCATAAAACAAGTAAAAAAGACTGAG
TTGGAGTCCTTCGAGAAATTTGCCCCAGAATATTTCAAGTATCTGATTGATTCTCTCAACTCAAGGAGCCCCACTTGCTTGGCAAAAATTCTTGGTATTT
ATCAGGTTACTGTAAAGCATTTGAGAGGTGTCAAGGAAACAAAAATGGATTTGATGGTGATGGAAAACCTCTTTTTTAACAGAAATATTGGGAGAGTCTA
CGACCTTAAGGGCTCTTCTCGATCCCGCTACAATACTGACACATCTGGGTCTAACAAGGTGTTACTAGATACGAATCTAGTGGAAAGACTGCGTACAGAG
CCCATATTTCTTGGAAGTAAGGCGAAGAGAAGCCTTGAGAGAGCTATATGGAATGATACGTCATTTTTAGCATCTGTTGATGTTATGGACTACTCTTTGC
TCGTTGGAGTGGATGATGAGCGGAAGGAGCTGGTTTTGGGGATCATTGATTTCATGAGACAGTATACATGGGACAAGCATTTAGAGACGTGGGTTAAGTC
ATCTGGAATACTTGGCGGTCCGAAAAATGCTTCCCCAACAATTGTTTCTCCAAAACAATACAAGAAAAGATTCCGAAAAGCAATGACTTCATACTTTCTT
ACTGTACCTGATCAATGGTCTTCACGGACTGAATCACTGCATTCTTGTCATTCATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G128800.3 pacid=42807934 polypeptide=Potri.008G128800.3.p locus=Potri.008G128800 ID=Potri.008G128800.3.v4.1 annot-version=v4.1
MCSECHSNNNQFFNGYHCQSCGKWLCFNCMRGYQSNGDFGEAIKSCKFCNGVTVKRDGGRKNSDKVHPTDSPRGSPEPPSPSFSAEPIHSDRLPLYLESR
DCGFSPNAITTRSMTSFSAHPSPVSVRRSSSRSDEEEAEDSGKLLYSPSSEYCHDISDIDSSSVSARLEFYNCKTVGSSPLDSPSRIDFSSCRVGHTVQQ
GREGSPLSQSDGPFDQENMAILSRPDKRTEDPENTDDCSDDGSVLRDQYHKSPKPLDFESNGLIWFPPPPEDENDEEESNFFTYDDEDDDIGDSSAIFSS
SSSLSSTFPSKEKQNKINKDPTKAMIQGHFRALVAQLLQGEGIKASKDENNGEWLDIVTAIAWQAAAFVKPDTSRGGSMDPVDYVKVKCIASGNPRDSTL
VKGVVCTKNIKHKRMTTQYKNPRLLLLGGALEYQSVVNQLASFNTLVQQENDHLKLIMSKIEALRPNVLLVEKSVSPYAQEYLLGKEISLVLNVKKPLLE
RIARCTGAQISPSFENISTTRLGHCELFRVERVSEEHETSNQFNKKPSKTLMSFEGCPRRLGCTVLLRGTCREKLKKVKHVIQYAVFAAYHLSLETSFLA
DEGASLPKMTIRPSIAIPERTAADNSISVIPPMICHAEVALSAQDDGSLGLKPEHEGSESLTGNLDAGVIHPLSPCSVTCRSGNEFSIACHGDLVSNAGG
LDAFSASQCEGLKMFAVSPGIKNLSQPELQDIMAEEEGQLLATHESVQSEKIDEDEVSSEYFSVTDTYQSILVSFSSRCVLKGTVCERSRLLRIKFYGNF
DKPLGRYLRDDLFDQKSCCRSCKEPAEAHVLCFTHQQGNLTINVRSLSSVKLPGDRDGKIWMWHRCLRCAHIDGVPPATRRVVMSDAAWGLSFGKFLELS
FSNHATANRVAPCGHSLQRDCLRFYGFGSMVVFFRYSPIDILNVHLPPSMLEFNGIVQQEWTRKEAAELLGKMETFYGEIFGVLDSMEQRSKYFGSELSD
TNELQNRIMELKDQLVKEKNNYSGILQLAVMESLQLDQTAMDILELNRLRRTLLIGSHVWYRKLYSLDCLLKTNYLVKAKEGDVSYTELKDLKNDIFCKD
SKLDHDHEENISGYSKSQEHVGNDFQSEKKETGLSFEHCVLEHSMLPSCYHNTEDEVHAGEETASKTLFSDNPSHASNLSDRIDSAWTGTDQLPIKVQPP
HASQAEADGFQPVSVRQPNLFDNPPFRRMVAPKRVHSFDSALRAQERIQKGLPPLHLSTIRSFHASGDYRSMVRDPVSNAMRTYSQTLPLEAHKLNLMHS
STHSFISSAANMAGGARLLLPVRANSDLVIGVYDNDPASVVSYALSSKEHEDWVTDRSNESAGIWSTIKHSKEDSAASSFTSWQSLDSMDLDYMSYGSYG
SEDPFSTLGTLFMDSKKSPHLTISYEDASSIAEGKVRFSVTCYFAKQFDFLRKKCCPSDVDFVRSLSRCQKWSAQGGKSNVYFAKSLDERFIIKQVKKTE
LESFEKFAPEYFKYLIDSLNSRSPTCLAKILGIYQVTVKHLRGVKETKMDLMVMENLFFNRNIGRVYDLKGSSRSRYNTDTSGSNKVLLDTNLVERLRTE
PIFLGSKAKRSLERAIWNDTSFLASVDVMDYSLLVGVDDERKELVLGIIDFMRQYTWDKHLETWVKSSGILGGPKNASPTIVSPKQYKKRFRKAMTSYFL
TVPDQWSSRTESLHSCHS
|
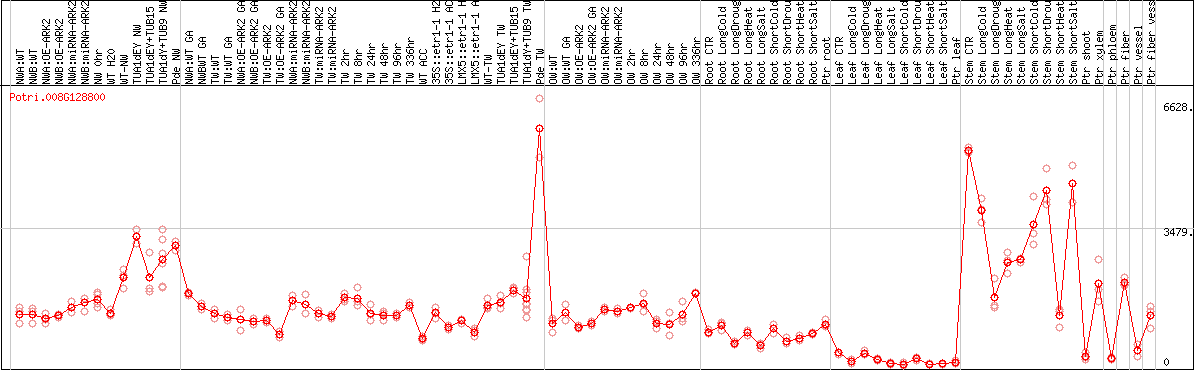
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G128800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
