External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14890 216 / 2e-71
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT2G01610 214 / 1e-70
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G23205 189 / 8e-61
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G70720 179 / 7e-57
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT5G62350 148 / 6e-45
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G25260 144 / 4e-43
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT3G47380 134 / 1e-39
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT4G12390 134 / 3e-39
PME1
pectin methylesterase inhibitor 1 (.1)
AT1G62770 133 / 5e-39
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
AT1G62760 132 / 2e-37
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G109300
315 / 2e-110
AT1G14890 218 / 5e-72
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128700
150 / 1e-45
AT5G62350 221 / 7e-74
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127500
141 / 4e-42
AT5G62350 220 / 2e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.003G113700
139 / 2e-41
AT4G12390 178 / 1e-56
pectin methylesterase inhibitor 1 (.1)
Potri.015G128300
138 / 4e-41
AT5G62360 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128100
137 / 1e-40
AT5G62360 221 / 1e-73
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.015G128900
127 / 9e-37
AT4G25250 145 / 4e-44
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.012G127400
125 / 5e-36
AT4G25250 150 / 7e-46
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Potri.014G129400
125 / 8e-36
AT3G62820 163 / 6e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038645
238 / 1e-79
AT2G01610 212 / 2e-69
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10027198
143 / 6e-43
AT5G62350 191 / 3e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10038914
139 / 3e-41
AT5G62350 192 / 1e-62
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031133
137 / 1e-40
AT5G62350 211 / 8e-70
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031713
134 / 2e-39
AT5G62350 202 / 4e-66
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031711
132 / 2e-38
AT5G62360 145 / 4e-44
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10032230
129 / 2e-37
AT1G62770 166 / 9e-52
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031138
126 / 4e-36
AT5G62360 172 / 3e-54
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10028910
122 / 1e-34
AT1G62760 169 / 1e-51
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
Lus10031712
120 / 4e-34
AT5G51520 149 / 2e-45
Plant invertase/pectin methylesterase inhibitor superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04043
PMEI
Plant invertase/pectin methylesterase inhibitor
Representative CDS sequence
>Potri.008G132600.1 pacid=42806304 polypeptide=Potri.008G132600.1.p locus=Potri.008G132600 ID=Potri.008G132600.1.v4.1 annot-version=v4.1
ATGCCATCAGAAACGCCAGGCCCTCTGCTACTCTTCACCTTCTTCCTAACAACCTTCCTTCTCTATTCCCAACCAATCTCAACCGTCGCTTCACCGGACA
CCGGCTTTTCTCAATCCAGCGGCACAGATTACATTCGCTCAAGCTGCGGCGCAACATTGTACCCCGAAATCTGCTACACCTCCCTCTCTCGCTACGCCAG
CGCTGTAAAACAAAGCCCCAGTCGCCTAGCTCGCGTGGCCATCGGCGTCAGCCTCTCAAGAGCACGCCGCCTGGCAGCCTACGTATCCAACCTAACCCGC
CATGAAGACTTCGGAGGCGATCACCGTGCCACGGCCGCCATTCACGACTGCTTATCCAACATGGGAGACGCCGTTGATGAGATGAGTGGGTCGTTGAAGC
AGATGCGGAAAGTTGGAGCAGCTGGGCTGTCGGCAGAGTCGTTTCAGTTTCAGATGAGTAACGTGCAGACGTGGATGAGTGCGGCTTTGACAGATGAGGA
GACGTGTACGGATGGATTTGAGGATGTAGCGGACGGGGCTGTGAAAACGGAGGTGTGTAACCGTGTTGCGGATGCCAAGAAATTTACTAGCAATGCTCTT
GCTCTGGTTAACACTTTTGCTGCCGCAGGAACGCCATGA
AA sequence
>Potri.008G132600.1 pacid=42806304 polypeptide=Potri.008G132600.1.p locus=Potri.008G132600 ID=Potri.008G132600.1.v4.1 annot-version=v4.1
MPSETPGPLLLFTFFLTTFLLYSQPISTVASPDTGFSQSSGTDYIRSSCGATLYPEICYTSLSRYASAVKQSPSRLARVAIGVSLSRARRLAAYVSNLTR
HEDFGGDHRATAAIHDCLSNMGDAVDEMSGSLKQMRKVGAAGLSAESFQFQMSNVQTWMSAALTDEETCTDGFEDVADGAVKTEVCNRVADAKKFTSNAL
ALVNTFAAAGTP
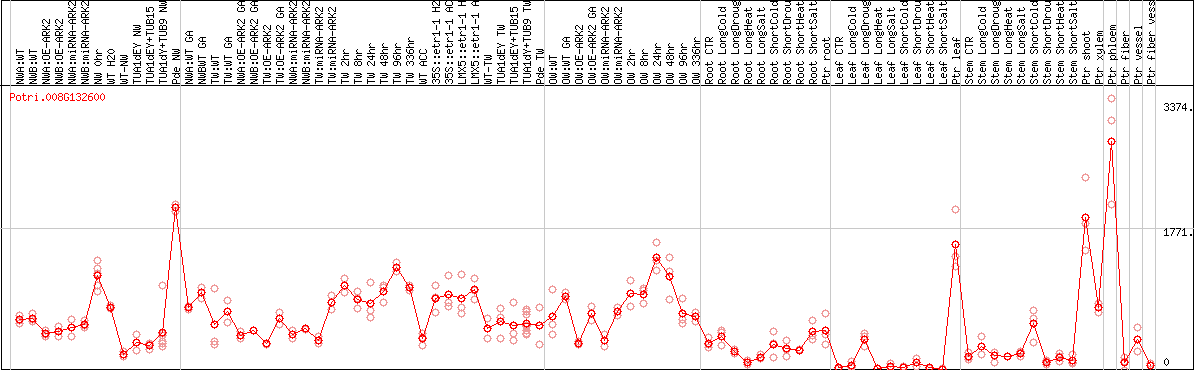
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G132600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.