Potri.008G134800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G134800.1 pacid=42806299 polypeptide=Potri.008G134800.1.p locus=Potri.008G134800 ID=Potri.008G134800.1.v4.1 annot-version=v4.1
ATGTCGTGGGAGGAAGAAGTTGTGATCCGCGATGTCACCAACGCCGGTATCGTCGTAAGCGACCGTATTGGTAGAGAGGTTGCAGCTCAGATCGATCTCG
AAGAAGCTCTAGAAGCTTCTAGATACGCGAGCCATCCTTATTCCACTCATCCTCGCGAGTGGCCACCTTTAATTGAGGTGGAGGATACTCAGGAGTTGCC
TCCCGTACTCGTTGAAAGATATAATGCAGCTGGCGGAGAAGCGACTGCTTTGTGTGGAATATTTCCTGAGGTACGGAGGGCTTGGGCATCGGTGGATAAT
TCTTTGTTTCTATGGCGTTTCGATAAGTGGGATGGTCAATGTCCTGAATACAGCGAGGAACAAGCCATCTGTGCTGTCGGCCTTGCCAAATCTAAGCCTG
GTGTTTTTGTTGAAGCTATCCAGTATCTTTTGGTTTTATCTACACCAGTGGAGTTGGTTCTTGTAGGAGTATGCTGTTCCGGGTCAGGCGATGGTGCGGA
TCCATATGCTGAGGTTTCCTTACAGCCTTTGCCAGAGTATACAGTACCATCTGATGGAGTTACCATGACCTGTATTGTGTGTACTGATAGGGGACGCATC
TTCCTTTCTGGGCGCGATGGACACATTTATGAGTTGCATTACACTACTGGTTCGGGATGGCACAAGCGATGTCGTAAAGTTTGCCTGACAGCTGGATTGG
GTAGTGTTATTTCAAGGTGGGTTGTACCAAATGTGTTCAAGTTTGGAGCTGTTGATCCTATTGTTGAAATGGTTGTTGACAATGAAAGACAGATTTTGTA
TGCACGAACAGAAGAGATGAAACTTCAGGTTTATCTTCTGGAATCAAATGGAGATGGTCCGCTGAAGAAAGTGGCAGAAGAAAGAAATTTGTTCAGTCAA
AGAGATGCACATTATGGAGGCAGACCATCAGCAGGACCAAGAGTCCCAAGTAGATCAGCAAAGCCATCTATAGCTTGCATCTCTCCTTTATCCACACTGG
AATCCAAGTGGTTGCACCTTGTTGCTGTTTTATCAGATGGCAGAAGGATGTACATTTCTACTTCCCCATCTAGTGGCAATAATGGAGCTGTTGGAGGTTT
GGGTGGATTCGGTACCAACCATCAGAAGCCGAACTGTTTAAAAGTTGTAACAACTAGACCTTCTCCTCCTTTAGGGGTCAGTGGTGGACTTGCCTTTGGG
GCCATTTCTCTTGCCAATAGAACTCCAAATGAGGATCTCACACTAAAAGTTGAAACAGCATCCTACTCTGCTGGAACTCTTGTCCTCTCTGATTCATCAC
CACCAACCACGTCTTCTCTTGTCATCGTGAGCAAGGATTCAAGCTCACAGACATCTGTTTCGGGTAGCTTAGGTACAAGTTCAAGGGGTTCACGGGCTTT
ACGGGAAACCGTATCTTCGGTACCTGTTGAAGGACGAATGCTTTTTGTTGCAGATGTTTTACCTTTACCAGATACAGCTGCCATGTTGCAGTCACTATAT
TCAGAACTTGATTGTTTTGGATTCCAAAGCGCATGTGAGCCCTGTGAAAAAGCATCTATAAAACTTTGGGCAAGAGGTGATCTTGCAATGCAACATGTAC
TTCCAAGGCGAAGAGTCATCATTTTCAGCACCATGGGAATGATTGAAGTGGTTTTCAACAGGCCCGTAGACATTCTGCGAAGGTTGTTTGAGTCCAACTC
CCCTAGATCAATCTTGGAGGATTTCTTTAATCGTTTTGGATCTGGTGAGGCAGCTGCAATGTGTTTAATGCTAGCTGCAAGGATAGTGCATTCTGAAAAT
CTTATAAGCAATCAAGTTGCTGAGAAGGCAGCTGAAACATATGAGGATCCAAGAGTTGTTGGAATGCCACAACTTGAAGGCAGCAATGTACTGTCAAACA
CTAGAACAGCAACAGGAGGTTTTAGCATGGGACAGGTTGTTCAGGAGGCTGAGCCTGTGTTTTCGGGTGCTCATGAAGGGCTCTGCTTGTGCTCTTCAAG
GTTGCTTTTACCTGTCTGGGAACTCCCTGTGTTCGTCTCAAAAGGTGATGTAGGTCCATCTGATGCTTCATTTGAAAATGGAGTGGTTGGATGTAGACTC
TCAGTTGGGGCTATGCAAATCCTAGAAAACAAGGTTCGTTCTTTAGAGAAGTTCCTTAAGTCCAGAAGGAACCAGAGAAGGGGGCTTTATGGTTGTGTAG
CAGGTTTGGGAGACTTGACTGGTTCTATATTGTATGGAGCTGGTTCAGATTCAGGTACTGGTGACAGAAGTATGGTGAGAAACTTATTTGGCACATATCC
ACAAAGTGTAGAGGCAAATGGTGGTGGGGCAACTAATAAACGTCAGCGACTACCATACAGTCCTGCAGAATTGGCTGCCATGGAGGTCAGGGCAATGGAG
TGCATCAGGCAATTGCTTCTTAGATCTGGGGAAGCTCTGTTTCTGCTACAGCTCCTTTCTCAACATCATATAACACGTGTGGTTCAGGGGCTTGATGCTA
GTATACGACAGAGTCTGGTTCAATTGACATTCCATCAACTGGTTTGTTCTGAGGAGGGTGATCGCCTTGCTACAATGCTGATTGCTGTCTTAATGGAGTA
CTATACTGGCCCTGATGGCAGGGGAACAGTAGATGATATTAGTGGGAAATTACGGGAGGGTTGTCCAAGCTACTTCAAGGAGTCTGATTACAAGTTTTTC
TTAGCTGTTGAATGTCTCGAAAGAGCTGCCGCAACTCCAGATCCTGTAGAGAAGGAGAATATTGCTAGAGAAGCCTTCAATTTCTTGAGCAAAGTTCCGG
AGTCTGCAGATTTAAGAACTGTTTGCAAACGGTTTGAGGATTTAAGGTTTTATGAAGCTGTGGTTCGCTTACCTCTGCAAAAAGCACAGGCTCTTGACCC
TGCAGGTGATGCTTTCAATGAGCAACTTGATGCAGCTACTAGAGAATATGCACTTGCTCAGCGTGAACAATGCTATGAGATAATTACCAGTGCTTTGCAT
TCTCTGAAAGGTGAGGCCTCACAAAAGGAATTTGGATCGCCAGTCAGACCTGCTTCAACTCGACCTGCACTTGATCAGGCTTCTCGGAAGAAATACGTAT
GCCAGATTGTTCAGCTTGCTGTCCAATCTCCTGACAGAGTATTTCATGAGTACCTATACTGGACTATGATTGATTTGGGCCTGGAAAATGAGCTGCTGGA
GTATGGAGGTCCAGATTTAGTACCATTTTTGCAAAGAGCTGGTCGGGAACCTTTACAAAAGGTTCATGCTGTTTCTGCCATAACACCTGCATCTTCTCCA
ATTGGTCATTCAGGAGCACCAATTGCTTCTAATCAAGCAAAATGTTTTGATCTGTTGGCACGGTATTATGTCCTGAAGAGGCAACACATACTTGCTGCTC
ACGTGTTGTTAAGGCTGGCTGAAAGACGCTCAACTGATGCTGGAGATGCTCCTAGTCTTGAACAAAGGCGCCAATACCTAAGTAATGCGGTTCTGCAGGC
CAAGAATGCTAGTGACAGTGGTGTTGTGGTAGGTTCTACAAGGGGTGCTATGGACACTGGATTGCTTGATTTGCTTGAGGGGAAGCTTGCGGTTCTTCGG
TTTCAAATAAAAATAAAAGATGAACTGGAGGCTATAGCGTCCAGGCTACAATCTTCATCTGACATGTCTGAGGCTGTCCAAAATGGATCAGCTCATGATA
GCAATGCTGAAGCAGAGCAGGCAAAAATTGCAAGGGAAAAGGCAAAAGAGTTATCATTGGATTTAAAGAGCATAACTCAGCTCTATAATGAATATGCAGT
TCCTTTTGAACTTTGGGAGATATGTCTGGAGATGCTATACTTTGCAAATTATTCTGGTGACGCTGATAGTAGCATTGTAAGAGAAACCTGGGCTAGACTA
ATTGATCAAGCTCTTTCAAGGGGTGGTGTTGTGGAAGCTTGTTCGGTTCTGAAGAGGGTTGGTTCCTACATGTATCCTGGAGATGGAGCTCTTCTACCTT
TAGACACCTTATGTCTTCACCTTGAGAAGGCTGCACTGGAGAGACTGGAATCAGGGGTGGAAACTGTTGGAGATGAAGACATAGCAAGAGCTCTTCTTGC
TGCTTGCAAGGGTGCGATTGAACCTGTGTTGAACACTTATGATCAGTTATTATCAAATGGAGCTATTTTGCCATCTCCCAACCTTAGGCTACGCCTTCTG
CGATCAGTGCTTGTGGTAATCCGTGAATGGGCAATGTCTGTTTTTGCACAGAGGATGGGTACAAGTGCAGCTGGCGCTTCATTAATATTAGGCGGATCAT
TCTCAGTGGAACAAACTGCAGTCATCAATCAAGGGATTCGCGATAAGATCACAAGTGCAGCAAACAGGTACATGACAGAGGTGAGGAGGTTGCCCCTTCC
CCAGGGCAGGACCGAAGCTGTTTACCAAGGATTTCGAGAACTTGAAGAGTCCCTGATAAGTCCTTTTTCTTTTGACAGATTTTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G134800.1 pacid=42806299 polypeptide=Potri.008G134800.1.p locus=Potri.008G134800 ID=Potri.008G134800.1.v4.1 annot-version=v4.1
MSWEEEVVIRDVTNAGIVVSDRIGREVAAQIDLEEALEASRYASHPYSTHPREWPPLIEVEDTQELPPVLVERYNAAGGEATALCGIFPEVRRAWASVDN
SLFLWRFDKWDGQCPEYSEEQAICAVGLAKSKPGVFVEAIQYLLVLSTPVELVLVGVCCSGSGDGADPYAEVSLQPLPEYTVPSDGVTMTCIVCTDRGRI
FLSGRDGHIYELHYTTGSGWHKRCRKVCLTAGLGSVISRWVVPNVFKFGAVDPIVEMVVDNERQILYARTEEMKLQVYLLESNGDGPLKKVAEERNLFSQ
RDAHYGGRPSAGPRVPSRSAKPSIACISPLSTLESKWLHLVAVLSDGRRMYISTSPSSGNNGAVGGLGGFGTNHQKPNCLKVVTTRPSPPLGVSGGLAFG
AISLANRTPNEDLTLKVETASYSAGTLVLSDSSPPTTSSLVIVSKDSSSQTSVSGSLGTSSRGSRALRETVSSVPVEGRMLFVADVLPLPDTAAMLQSLY
SELDCFGFQSACEPCEKASIKLWARGDLAMQHVLPRRRVIIFSTMGMIEVVFNRPVDILRRLFESNSPRSILEDFFNRFGSGEAAAMCLMLAARIVHSEN
LISNQVAEKAAETYEDPRVVGMPQLEGSNVLSNTRTATGGFSMGQVVQEAEPVFSGAHEGLCLCSSRLLLPVWELPVFVSKGDVGPSDASFENGVVGCRL
SVGAMQILENKVRSLEKFLKSRRNQRRGLYGCVAGLGDLTGSILYGAGSDSGTGDRSMVRNLFGTYPQSVEANGGGATNKRQRLPYSPAELAAMEVRAME
CIRQLLLRSGEALFLLQLLSQHHITRVVQGLDASIRQSLVQLTFHQLVCSEEGDRLATMLIAVLMEYYTGPDGRGTVDDISGKLREGCPSYFKESDYKFF
LAVECLERAAATPDPVEKENIAREAFNFLSKVPESADLRTVCKRFEDLRFYEAVVRLPLQKAQALDPAGDAFNEQLDAATREYALAQREQCYEIITSALH
SLKGEASQKEFGSPVRPASTRPALDQASRKKYVCQIVQLAVQSPDRVFHEYLYWTMIDLGLENELLEYGGPDLVPFLQRAGREPLQKVHAVSAITPASSP
IGHSGAPIASNQAKCFDLLARYYVLKRQHILAAHVLLRLAERRSTDAGDAPSLEQRRQYLSNAVLQAKNASDSGVVVGSTRGAMDTGLLDLLEGKLAVLR
FQIKIKDELEAIASRLQSSSDMSEAVQNGSAHDSNAEAEQAKIAREKAKELSLDLKSITQLYNEYAVPFELWEICLEMLYFANYSGDADSSIVRETWARL
IDQALSRGGVVEACSVLKRVGSYMYPGDGALLPLDTLCLHLEKAALERLESGVETVGDEDIARALLAACKGAIEPVLNTYDQLLSNGAILPSPNLRLRLL
RSVLVVIREWAMSVFAQRMGTSAAGASLILGGSFSVEQTAVINQGIRDKITSAANRYMTEVRRLPLPQGRTEAVYQGFRELEESLISPFSFDRF
|
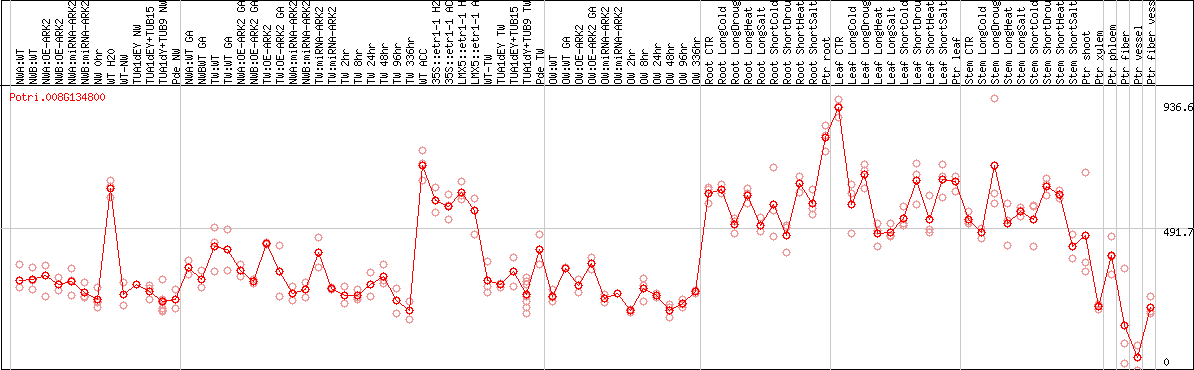
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G134800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
