External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G23770 66 / 3e-12
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
AT3G01840 49 / 2e-06
Protein kinase superfamily protein (.1)
AT1G51940 45 / 2e-05
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
AT2G17120 45 / 3e-05
LYM2
lysm domain GPI-anchored protein 2 precursor (.1)
AT1G77630 43 / 0.0001
LYM3
lysin-motif \(LysM\) domain protein 3, Peptidoglycan-binding LysM domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G160600
375 / 7e-128
AT2G33580 217 / 8e-62
Protein kinase superfamily protein (.1)
Potri.010G078700
342 / 8e-115
AT2G23770 288 / 5e-89
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.005G128501
152 / 9e-46
AT2G23770 71 / 4e-14
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.007G032300
152 / 1e-42
AT2G33580 247 / 6e-73
Protein kinase superfamily protein (.1)
Potri.001G332800
72 / 3e-14
AT3G01840 471 / 4e-158
Protein kinase superfamily protein (.1)
Potri.005G259600
72 / 4e-14
AT2G33580 637 / 0.0
Protein kinase superfamily protein (.1)
Potri.005G128200
66 / 2e-12
AT2G23770 514 / 4e-176
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.005G128300
63 / 3e-11
AT2G23770 394 / 2e-129
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.014G040000
59 / 7e-10
AT2G23770 345 / 1e-110
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021889
176 / 7e-52
AT2G23770 256 / 2e-77
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10016793
127 / 3e-33
AT3G21630 222 / 9e-64
LYSM DOMAIN RECEPTOR-LIKE KINASE 1, chitin elicitor receptor kinase 1 (.1)
Lus10022487
123 / 9e-32
AT5G66631 707 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10039406
77 / 8e-16
AT2G33580 229 / 4e-68
Protein kinase superfamily protein (.1)
Lus10016299
74 / 9e-15
AT2G23770 368 / 3e-119
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10022345
73 / 2e-14
AT3G01840 429 / 1e-141
Protein kinase superfamily protein (.1)
Lus10008105
72 / 5e-14
AT1G16120 419 / 3e-137
wall associated kinase-like 1 (.1)
Lus10041171
69 / 3e-13
AT2G33580 224 / 1e-64
Protein kinase superfamily protein (.1)
Lus10004504
68 / 9e-13
AT1G16130 411 / 5e-134
wall associated kinase-like 2 (.1)
Lus10008586
62 / 1e-10
AT2G33580 594 / 0.0
Protein kinase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0187
LysM
PF01476
LysM
LysM domain
Representative CDS sequence
>Potri.008G160501.1 pacid=42808222 polypeptide=Potri.008G160501.1.p locus=Potri.008G160501 ID=Potri.008G160501.1.v4.1 annot-version=v4.1
ATGAGACCCGCATCCCGTTTAGTCTCTTCTCTCTTCTTCTTTCTCTCTTACAGTAATATCCTCCACCACTTGCAAGCCCAGCCAAGCACTCAAGGGTTCA
CCTGCCCAGCCAACCAGAGGTCCTTTCCATGTCAAACCTATGCCTTCTACCGAGCCTCAGCTCCTAACTTCCTTGACCTTGCCTCAATCGGTGACCTTTT
CTCGGTTAGCCGCCTTATGATATCAAAACCAAGTAACATCTCCTCTCCAACCTCACCTCTCATCCCCAATCAACCCCTCTTTGTCCCTTTATCATGTTCT
TGCAACACCATCAATATTAGCACTAGCATTTCCTCTGCAAATATCACATATACCATCAAGGAAGGCAACACTTTCTACATTGTCTCGACTAAATACTTCC
AAAACCTTACGACCTACCAGTCAGTTGAGCTTTTCAACCCTACACTTATCCCCGAACTACTCGACATAGGAGTAGAGGTGATCTTTCCAATATTTTGCAA
GTGTCCTCATCAAACCCAGTTGCAAAACAAGGTGAATTATCTGGTATCTTATGTGTTTCAGCCTTCTGATAACTTATCTTCAGTTGCTTCAACATTTGGA
GTTGAAACACAATCTATTGTTGATGTTAATGGCAATAACATCCAGCCTTATGATACCATATTCGTTCCAGTCTATCAACTTCCACAACTGGCACAACCTA
CAGATAGAGCATTGAAAGTTAACAGAATTGGAACCTGA
AA sequence
>Potri.008G160501.1 pacid=42808222 polypeptide=Potri.008G160501.1.p locus=Potri.008G160501 ID=Potri.008G160501.1.v4.1 annot-version=v4.1
MRPASRLVSSLFFFLSYSNILHHLQAQPSTQGFTCPANQRSFPCQTYAFYRASAPNFLDLASIGDLFSVSRLMISKPSNISSPTSPLIPNQPLFVPLSCS
CNTINISTSISSANITYTIKEGNTFYIVSTKYFQNLTTYQSVELFNPTLIPELLDIGVEVIFPIFCKCPHQTQLQNKVNYLVSYVFQPSDNLSSVASTFG
VETQSIVDVNGNNIQPYDTIFVPVYQLPQLAQPTDRALKVNRIGT
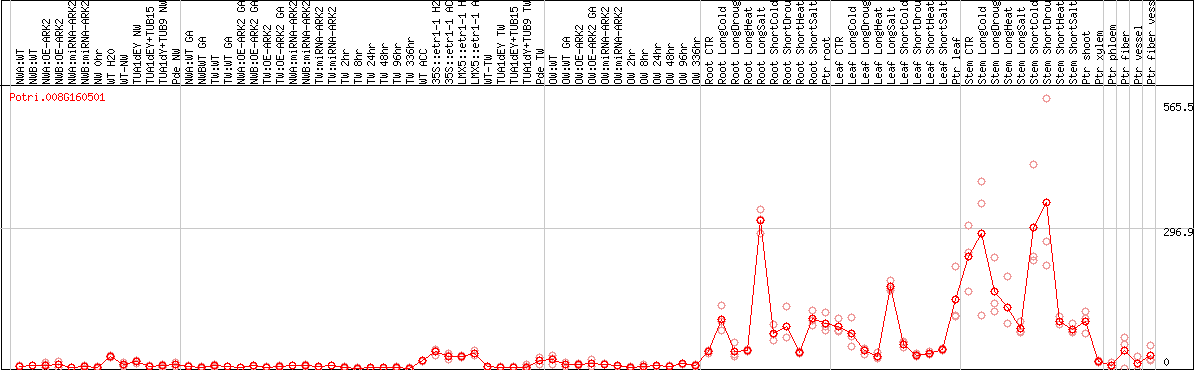
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G160501 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.