External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G23740 408 / 9e-143
AOR
alkenal/one oxidoreductase, Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
AT4G21580 91 / 9e-21
oxidoreductase, zinc-binding dehydrogenase family protein (.1.2.3)
AT3G15090 75 / 6e-15
GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
AT4G13010 72 / 3e-14
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
AT1G49670 56 / 2e-08
NQR
ARP protein (REF) (.1), ARP protein (REF) (.2)
AT3G45770 51 / 4e-07
Polyketide synthase, enoylreductase family protein (.1.2)
AT3G56460 49 / 3e-06
GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G136100
426 / 5e-150
AT1G23740 504 / 9e-180
alkenal/one oxidoreductase, Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.001G342800
92 / 3e-21
AT4G13010 513 / 0.0
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.001G342900
87 / 2e-19
AT4G13010 484 / 2e-173
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.001G391100
80 / 7e-17
AT4G13010 469 / 8e-168
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.004G036100
78 / 3e-16
AT4G21580 525 / 0.0
oxidoreductase, zinc-binding dehydrogenase family protein (.1.2.3)
Potri.001G452600
76 / 3e-15
AT4G13010 469 / 1e-167
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.001G343000
73 / 1e-14
AT4G13010 369 / 2e-129
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Potri.001G373200
72 / 4e-14
AT3G15090 515 / 0.0
GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Potri.019G065700
62 / 1e-10
AT3G56460 525 / 0.0
GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030615
419 / 7e-147
AT1G23740 519 / 0.0
alkenal/one oxidoreductase, Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10030873
408 / 2e-142
AT1G23740 519 / 0.0
alkenal/one oxidoreductase, Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10003159
91 / 5e-20
AT4G13010 449 / 4e-158
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10002347
88 / 1e-19
AT4G13010 468 / 8e-167
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10003158
89 / 2e-19
AT4G13010 473 / 4e-168
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10038206
82 / 1e-17
AT4G13010 451 / 3e-160
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10025898
82 / 2e-17
AT4G13010 446 / 1e-158
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10002346
80 / 2e-17
AT4G13010 265 / 3e-89
Oxidoreductase, zinc-binding dehydrogenase family protein (.1)
Lus10003495
81 / 4e-17
AT3G15090 528 / 0.0
GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Lus10011233
77 / 2e-15
AT4G21580 502 / 1e-180
oxidoreductase, zinc-binding dehydrogenase family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0296
GroES
PF08240
ADH_N
Alcohol dehydrogenase GroES-like domain
CL0063
NADP_Rossmann
PF13602
ADH_zinc_N_2
Zinc-binding dehydrogenase
Representative CDS sequence
>Potri.008G161600.1 pacid=42808639 polypeptide=Potri.008G161600.1.p locus=Potri.008G161600 ID=Potri.008G161600.1.v4.1 annot-version=v4.1
ATGGAGATCCCATCGAGTACTTCCATTCCGTCTAAAATGAAGGCTTGGGTCTATGGTGAATATGGGAACGTTTCCAATGTCTTAAAATTAGACTCGAACG
TTACAGTTCCTCAAGTGAAAGAAGATCAGGTTCTGATCAAGGTTGTTGCTGCATCCATTAATCCAGTTGATGCTAAAAGGATGCTTGGCATGTTCAAAGT
CAGTGACTCTCCTGTGCCGACTGTTCCGGGCTATGATGTGGCTGGTGTGGTGGTTAAGGTTGGAAGCCAAGTTAAAAGGCTGAAGGTAGGAGATGAAGTA
TACGGAGACATCAATGAGAAGGCTCTGGACCACCCCAAACGATTTGGCTCTTTAGCCGAGTACACTGCTGTTGAAGAGAATCTAGTGGCTCTTAAGCCCA
AAAATCTGAGTTTTGCTGAAGCAGCCAGCCTGCCCCTTGTCATTGAAACCGCTCATGAAGGTCTTGAAAGAACTGGGTTCTCTGCGGGCAAATCTATCCT
TGTTTTAGGAGGTGCTGGTGGAGTTGGTACCCAAATAATTCAGCTAGCAAAGCATGTTTTTGGTGCATCTACAGTAGCAGCCACCTCTAGCACATCAAAA
TTGGAGTTGCTGAAAAGCTTAGGAGCAGATTTGGCTATTGATTACACCAAGGAAAACTTTGAGGACCTGCCAGAGAAGTTTGATGTAGTATATGATGCAG
TTGGGCAGTGTGACAGGGCAGTCAAGGCAGTGAAAGAAGATGGAAGTGTTGTGACAATAGTTGGTCCTATAACTCCCCCGGCTTTGATATTCGTGCTCAC
TTCTAATGGCTCTGTCTTGGACAAACTGAAGCCTTACTTGGAGAGTGGAAAGGTGAAGCCGGTGCTAGATCCCAAGGGTCCATTCCCATTTTCTCAAACT
GCAGAGGCATTTTCCTATCTTGAAACCAGCAGAGCTGTTGGAAAAGTGGTCATTTATCCGATCCCATGA
AA sequence
>Potri.008G161600.1 pacid=42808639 polypeptide=Potri.008G161600.1.p locus=Potri.008G161600 ID=Potri.008G161600.1.v4.1 annot-version=v4.1
MEIPSSTSIPSKMKAWVYGEYGNVSNVLKLDSNVTVPQVKEDQVLIKVVAASINPVDAKRMLGMFKVSDSPVPTVPGYDVAGVVVKVGSQVKRLKVGDEV
YGDINEKALDHPKRFGSLAEYTAVEENLVALKPKNLSFAEAASLPLVIETAHEGLERTGFSAGKSILVLGGAGGVGTQIIQLAKHVFGASTVAATSSTSK
LELLKSLGADLAIDYTKENFEDLPEKFDVVYDAVGQCDRAVKAVKEDGSVVTIVGPITPPALIFVLTSNGSVLDKLKPYLESGKVKPVLDPKGPFPFSQT
AEAFSYLETSRAVGKVVIYPIP
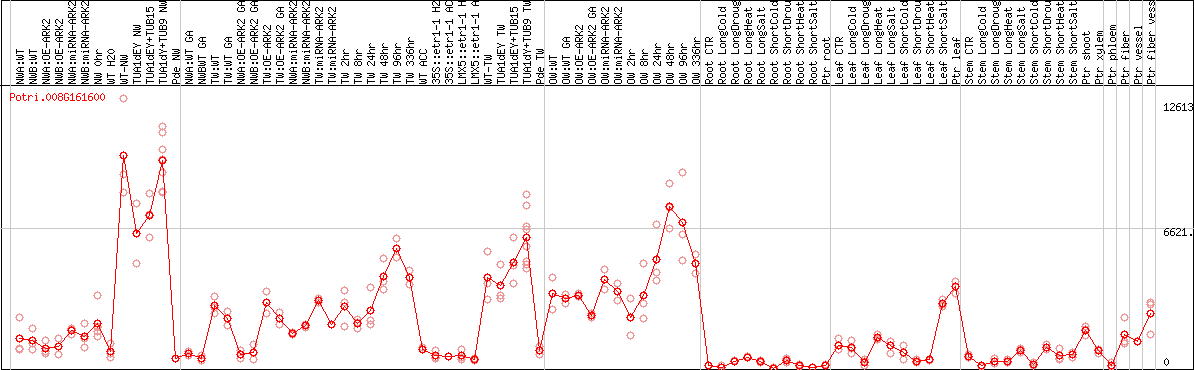
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G161600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.