ETR2.2 (Potri.008G164400) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ETR2.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G164400.2 pacid=42806397 polypeptide=Potri.008G164400.2.p locus=Potri.008G164400 ID=Potri.008G164400.2.v4.1 annot-version=v4.1
ATGTTAAAAGCATTAGCTCCTGGGCTGCTGTTGATTTTATCACTCCTGATATCAGCTTCTGCAAACGACAACGGATTTTCTCGATGTAATTGTGAAGATG
AGGGCAGTTTATGGATCATAGAAAGCATTCTAGAGAGCCAAAGAGTGAGTGATTTCTTGATTGCCGTGGCCTATTTTTCAATCCCTATCGAGCTGCTATA
CTTCGTTAGCTGCTCAAATGTTCCATTCAAATGGGTTCTCTTCGAGTTCATTGCTTTCATTGTTCTATGTGGATTAACCCATTTGATCAATGGCATGACG
TACGGCCCCCACACATTTCAGCTTATGCTTGCCCTCACTGTCTTCAAGATTTTAACTGCTCTGGTTTCTTGTGCCACTGCCATTACCCTTTTCACTCTCA
TTCCTTTGCTTCTCAAAGTGAAGGTCAGAGAATTCATGTTGAAGAAGAAGGCGTGGGATCTTGGTCGTGAAGTTGGGATTATAATGAAACAAAAAGAAGC
TGGGTTGCATGTTCGTATGCTTACTCAAGAGATTCGAAAATCACTGGATAGGCATACGATTTTGTATACTACCCTTGTTGAGTTATCTAAGACATTGGGG
TTGCAAAACTGTGCTGTTTGGATGCCTAATGAGATGAAAACCCTGATGGATTTGACCCATGAGCTGAATAGGGGTAATTATTTGAGTTCAGATAATCCTT
CAATTCCAATCACTGATCCTGATGTTGTTAGAATTAAACGAAGTGAAGCGGTAAACATTCTTAGGCCTGACTCAGCGCTTGCTGCTGCAAGCCATGGGGA
GTCTGGCGAGCCAGGGCCTGTGGCTGCAATTCGAATGCCAATGCTTCATGTTAGCAATTTCAAAGGGGGGACTCCTGAGATAGTTCAGGCTTGTTATGCG
ATTTTGGTATTGGTTCTCCCCGGTGGACAACCCAGATCCTGGACCAACCAGGAAGTAGAGATAATTAAGGTGGTAGCTGATCAGGTGGCCGTGGCTCTCT
CTCATGCTGCAGTTCTTGAAGAGTCCCAACTCATGAGGGAGAAGTTGGAGGAGCAAAACCGAGCTTTGCAACAGGCCAAAATGAACGCAATGATGGCAAG
CAAAGCTAGGGGTGCTTTTCAGAAGGTAATGAGTGATGGGATGAAGAGACCAATGCACTCGATCTTGGGTTTGATTTCCTTGATACAGGATGGTAATTTG
AGTGGTGAGCAACGGATTATTGTTGATGCAATGATGAGGACTAGCAATGTCCTATCAACGTTGATAAATGATGTGACTGAAATTTCAATAAAGGATAGTG
GAAGATTTTCATTGGATATGAGATCATTTGGGTTACATGCCATGATCAAAGAAGCAGCTTGCCTTGCCAAATGCTTGTGCATTTATAGGGGCTTTGGTTT
TTCAATTGAGGTTGACAAGTCCTTGCCGGATAATGTTATGGGTGATGAAAGGAGGGTTTTTCAGGTGATTTTGCATATGGTTGGGAACCTACTTGATCAC
AACAATGGAGGAGGGTTTGTAGTGCTTCGGTTTTTCTCTGAGAATGGAAGTCAAGAGAGGAATGATCAGAGATGGACCACCTGGAGACCTTGCATGTCTG
ATGGGGATGTATATATCAGATTTGAAATTGCAATAAACAATAGTGGCTCCGAATCAGAGGGCTCAGCTTCAATGTTACAACATAGTGGTAAGAGGTTTGC
CAGCGATGGAGTTGAGGAGGGCTTGAGCTTCAGCATTTGCAAAAAGCTGGTCCATTTGATGCAAGGTAAGATCTGGATGATGCCCAACTCTCAAGGTTTT
GCTGAAAGCATGGGATTTGTTCTTCGGTTTCAACTTCGACCATCCATTGCAGTGGCCATCTCTGAATCTGGAGAATCTTCAGAGAACCCACATTCCAACT
CTTTTTTCAAAGGCTTGCAAGTTCTATTAGCTGATGCTGATGATTTGAACAGAGCCGTGACCCGGAAGCTGCTTGAGAGGCTAGGTTGCAATGTTGCTAC
TGTTGCATCTGGGTTTGAATGCCTTAGTGCTCTTGGACCAGCTGCATCTTTCCAAGTAGTTCTCTTGGATCTTCAGATGCCTGAGTTGGACGGATATGAA
GTTGCAGTGAGAATCCGGAAGTTTAGAAGTCGAAGTTGGCCTTTGATTATTGCCATGACAGCAAGTTCTGATGACGATGTGTGGGACAAATGCCTGCAAA
TCGGAATCAATGGTGTCATTCAGAAACCAGTTGTTCTAAAAGGAATTTCCTATGAGCTTCGAAGAGTCCTGGCAAACAAAGTTGTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.008G164400.2 pacid=42806397 polypeptide=Potri.008G164400.2.p locus=Potri.008G164400 ID=Potri.008G164400.2.v4.1 annot-version=v4.1
MLKALAPGLLLILSLLISASANDNGFSRCNCEDEGSLWIIESILESQRVSDFLIAVAYFSIPIELLYFVSCSNVPFKWVLFEFIAFIVLCGLTHLINGMT
YGPHTFQLMLALTVFKILTALVSCATAITLFTLIPLLLKVKVREFMLKKKAWDLGREVGIIMKQKEAGLHVRMLTQEIRKSLDRHTILYTTLVELSKTLG
LQNCAVWMPNEMKTLMDLTHELNRGNYLSSDNPSIPITDPDVVRIKRSEAVNILRPDSALAAASHGESGEPGPVAAIRMPMLHVSNFKGGTPEIVQACYA
ILVLVLPGGQPRSWTNQEVEIIKVVADQVAVALSHAAVLEESQLMREKLEEQNRALQQAKMNAMMASKARGAFQKVMSDGMKRPMHSILGLISLIQDGNL
SGEQRIIVDAMMRTSNVLSTLINDVTEISIKDSGRFSLDMRSFGLHAMIKEAACLAKCLCIYRGFGFSIEVDKSLPDNVMGDERRVFQVILHMVGNLLDH
NNGGGFVVLRFFSENGSQERNDQRWTTWRPCMSDGDVYIRFEIAINNSGSESEGSASMLQHSGKRFASDGVEEGLSFSICKKLVHLMQGKIWMMPNSQGF
AESMGFVLRFQLRPSIAVAISESGESSENPHSNSFFKGLQVLLADADDLNRAVTRKLLERLGCNVATVASGFECLSALGPAASFQVVLLDLQMPELDGYE
VAVRIRKFRSRSWPLIIAMTASSDDDVWDKCLQIGINGVIQKPVVLKGISYELRRVLANKVV
|
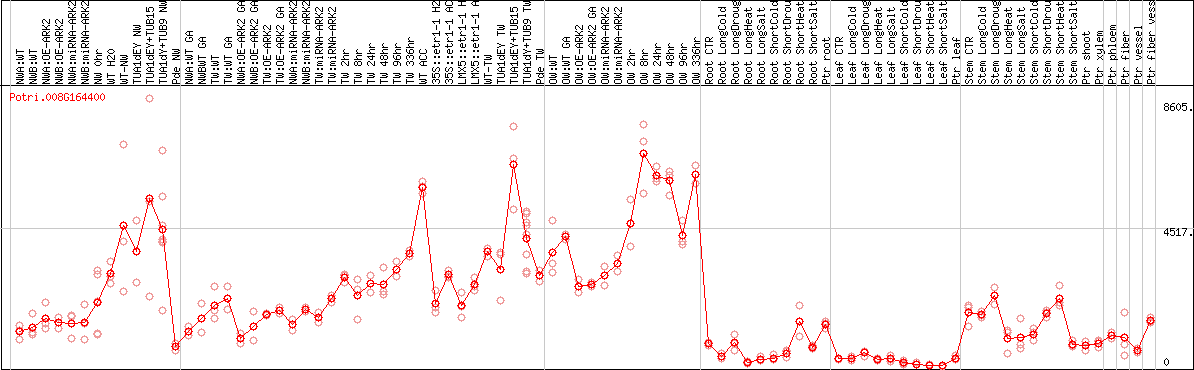
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G164400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
