External link
Symbol
Pt-PBE1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G26340 472 / 1e-170
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
AT1G13060 446 / 2e-160
PBE1
20S proteasome beta subunit E1 (.1.2)
AT4G31300 93 / 2e-22
PBA1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT5G40580 84 / 8e-19
PBB2
20S proteasome beta subunit PBB2 (.1.2.3)
AT3G27430 82 / 2e-18
PBB1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT1G53850 68 / 2e-13
PAE1, ATPAE1
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
AT1G77440 67 / 3e-13
PBC2
20S proteasome beta subunit C2 (.1.2)
AT3G14290 67 / 6e-13
PAE2
20S proteasome alpha subunit E2 (.1)
AT1G21720 65 / 2e-12
PBC1
proteasome beta subunit C1 (.1)
AT4G14800 56 / 2e-09
PBD2
20S proteasome beta subunit D2 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G058100
530 / 0
AT3G26340 463 / 6e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Potri.006G077900
91 / 8e-22
AT4G31300 416 / 2e-149
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.018G145900
88 / 2e-20
AT4G31300 405 / 3e-145
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.017G071100
82 / 5e-18
AT5G40580 497 / 1e-180
20S proteasome beta subunit PBB2 (.1.2.3)
Potri.004G066000
81 / 9e-18
AT3G27430 495 / 1e-179
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.001G162900
68 / 2e-13
AT3G14290 471 / 4e-171
20S proteasome alpha subunit E2 (.1)
Potri.003G072500
67 / 4e-13
AT3G14290 464 / 1e-168
20S proteasome alpha subunit E2 (.1)
Potri.002G080800
64 / 4e-12
AT1G21720 383 / 1e-137
proteasome beta subunit C1 (.1)
Potri.005G180500
63 / 8e-12
AT1G21720 391 / 9e-141
proteasome beta subunit C1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011369
474 / 3e-171
AT3G26340 465 / 1e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Lus10006426
467 / 1e-168
AT3G26340 462 / 7e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Lus10020180
90 / 4e-21
AT4G31300 429 / 7e-155
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10032102
86 / 3e-19
AT3G27430 473 / 6e-171
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10014581
86 / 4e-19
AT5G40580 471 / 2e-169
20S proteasome beta subunit PBB2 (.1.2.3)
Lus10026984
86 / 7e-19
AT4G31300 357 / 2e-124
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10004885
66 / 1e-12
AT1G21720 356 / 2e-127
proteasome beta subunit C1 (.1)
Lus10020599
64 / 3e-12
AT1G21720 355 / 7e-127
proteasome beta subunit C1 (.1)
Lus10037454
66 / 4e-12
AT1G53850 464 / 8e-163
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
Lus10003936
66 / 6e-12
AT1G53850 463 / 9e-162
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF00227
Proteasome
Proteasome subunit
Representative CDS sequence
>Potri.008G177000.7 pacid=42807413 polypeptide=Potri.008G177000.7.p locus=Potri.008G177000 ID=Potri.008G177000.7.v4.1 annot-version=v4.1
ATGAAGTTTGATACTAGCGGTCTCGAGTCCACTGCTTCGGTCTTTGGAACGGCCGGCAGTGAGCTTGTTGATGGGTTTTCTGCTGCTCCAGCTTTTGAGC
TACCCACTACTACAGATTTTGATGGCTTCCAGAAGGAAGCTGTACAGATGGTGAAGCCGGCCAAGGGGACAACTACGCTTGCCTTTATCTTTAAGGATGG
TGTCATTGTTGCGGCTGATTCTCGTGCTAGCATGGGAGGCTATATTTCATCACAGTCAGTTAAAAAAATCATTGAAATCAATCCCTACATGCTTGGTACA
ATGGCTGGAGGAGCTGCTGATTGTCAGTTTTGGCACAGAAATCTGGGCATTAAGTGCCGACTGCATGAATTGGCAAACAAGCGTAGAATATCAGTCACAG
GGGCATCAAAGCTTCTGGCAAACATTCTGTACTCTTACCGTGGAATGGGCTTGTCTGTTGGGACCATGATTGCTGGTTGGGATGAAACGGGTCCTGGACT
ATATTACGTGGACAGTGAAGGTGGAAGGCTGAAGGGAACAAGATTCTCTGTTGGATCTGGTTCTCCATATGCATATGGTATACTGGATAGTGGGTATCGA
TTTGATATGTCAATCGAGGAAGCTGCAGAGTTGGGCAGAAGAGCTATTTATCATGCAACTTTCCGTGATGGAGCCAGTGGTGGAGTAGCAAGCGTTTATT
ATGTGGGACCTGATGGATGGAAGAAATTATCTGGTGATGATGTCTCAGAGCTCCACTACAATTATTATCCAGTTGTGTCAACTGAAGCTTCAGAACCGGA
CCGAATGGTTGAAGCATAA
AA sequence
>Potri.008G177000.7 pacid=42807413 polypeptide=Potri.008G177000.7.p locus=Potri.008G177000 ID=Potri.008G177000.7.v4.1 annot-version=v4.1
MKFDTSGLESTASVFGTAGSELVDGFSAAPAFELPTTTDFDGFQKEAVQMVKPAKGTTTLAFIFKDGVIVAADSRASMGGYISSQSVKKIIEINPYMLGT
MAGGAADCQFWHRNLGIKCRLHELANKRRISVTGASKLLANILYSYRGMGLSVGTMIAGWDETGPGLYYVDSEGGRLKGTRFSVGSGSPYAYGILDSGYR
FDMSIEEAAELGRRAIYHATFRDGASGGVASVYYVGPDGWKKLSGDDVSELHYNYYPVVSTEASEPDRMVEA
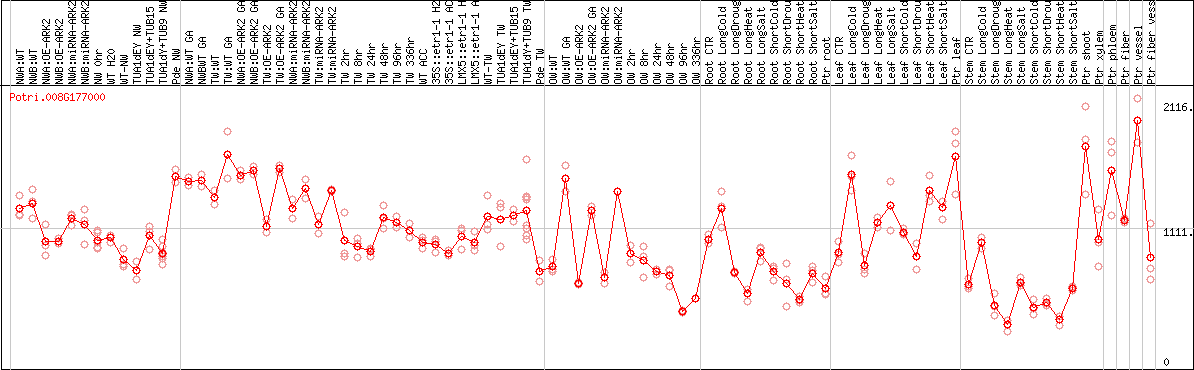
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G177000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.