Potri.008G177900 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.008G177900.1 pacid=42806615 polypeptide=Potri.008G177900.1.p locus=Potri.008G177900 ID=Potri.008G177900.1.v4.1 annot-version=v4.1
ATGAAACTGACACCAAGAGAGGTGGATAAGCTAGGACTCCACAATGCTGGATTTCTCGCACAGAAACGCCTTGCTCGAGGCCGCAAGCTTAATTATACCG
AGGCCGTCGCTCTTATTGCCTCTCAGATTTTGGAATTTGTTCGCGATGGTGACAAGAGTGTAGCTGAATTGATGGACATTGGGAAACAGATACTTGGAAG
GAGACAAGTTCTTCCAGCAGTTCCACATCTTCTCGATACTGTACAGGTTGAAGGGACATTTCCTGACGGGACCAAGTTAATCACTGTACATAATGCGATT
GCTAGTGAAAATGGGAATCTGGAGCTAGCTTTACAGGGTTCTTTTCTTCCAGTTCCTTCTCTCGACAAATTTCCTGCAATAGAAGATAATGAAATTCCTG
GTGCAATAATCTTTGGAGACGGGAATGTCATAATTAATTCTGGAAGGAAAGCAGTAACCCTCAAAGTCATTAACACTGGAGACAGGCCAATTCAGGTAGG
CAGCCACTATCATTTTATCGAGACAAATCGGAGTTTACTTTTTGATCGAAGAAAAGCACATGGCATGCGCTTGAACATCCCAGCTGGGACAGCCATACGA
TTTGAGCCAGGGGAAAGTAAAAGTGTAGTGCTTGTAAGCATTGGAGGTAAACAGGTGATCAAAGGTGGAAATGGCATTGTTGATGGGCCTGTTGATCATG
AAAACTGGACCAATATCATGGGAAACATAAGGAGGAGAGAATTTGGGAATCGAGAAGAAGAAAATGCTAGTGAAGGTGTAATTGGAGAAGGTTCTGCATT
CAACAATACAATTTCTCGCGAGGCTTATGCTAACATGTATGGCCCTACTGCTGGTGACAAAATTCGTCTTGGTGATACCAACCTGTATGCTGAAATTGAA
AGAGATTTTGCTTTTTATGGTGATGAATGTGTCTTTGGAGGTGGAAAAGTTATAAGGGATGGAATGGGTCAGTCATGCGGACATCAACCAGCTGACTCAC
TTGATACCGTTATAACAAATGCTGTGGTCATTGATTATAGTGGAATTTACAAGGCAGATATTGGTATTAAGGATTATCTTATTCATGCTATTGGAAAATC
AGGAAATCCAGATGTCATGAATGTTCCCTCTGATATGACCATTGGGGTTAACACCGAGGTTATTGCGGGGGAAGGAATGATTGTAACTGCAGGGGGCATA
GACTGTCATGTGCACTTCATATGCCCTCAACTTGCATTTGAGTCGATATCAAGTGGCATTACAACACTTGTGGGAGGTGGAACAGGACCTGCTGATGGGA
CACGTGCAACGACATGTACTCCTGCTCCTTCACATATGAAATTAATGCTGCAGTCTACTGATGACCTGCCTCTAAACTTTGGCTTCACTGGAAAAGGTAA
TGCTGCCAAACCTGAGGAGCTACATAAAATCATCAGGGCTGGAGCAATGGGACTGAAGCTTCACGAGGACTGGGGAACTACTCCTGCTGCAATAGACAAT
TGTTTGACTGTTGCAGATGAATATGACGTTCAGGCCAATATCCATACTGATACCTTGAATGAATCTGGATTCGTGGAAGATACTATTGCTGCATTTAAAG
GAAGAACTATCCATACTTATCATAGTGAAGGTGCAGGTGGTGGTCATGCTCCAGATATTATCAAAGTATGTGGTGTAAAAAATGTCCTGCCCTCATCAAC
GAACCCTACACGGCCATATACTTCCAATACTATAGATGAGCATCTAGATATGCTGATGGTCTGCCATCACCTTGATAAAAACATTCCAGAAGATGTAGCT
TTTGCCGAATCAAGAATAAGGGCTGAAACAATTGCTGCAGAAGACATATTACATGATATGGGAGCGATTAGCATTATATCTTCCGATTCACAGGCCATGG
GCCGCATTGGAGAGGTCATAAGTAGAACATGGCAAACTGCTCACAAGATGAAATCACAAAGGGGATTGATTGGGCCTGGTGGATCAGACAATGACAACTT
TCGCATCAGAAGATACATTGCAAAGTACACAATAAACCCTGCAATAGCTAACGGGCTTGCTAAGTTTGTTGGTTCAGTTGAGGTAGGGAAGTTAGCTGAT
CTTGTTTTATGGAAGCCGTCTTTCTTTGGGGCAAAACCAGAAATGGTGATTAAAGGTGGTGCTATTGCATGGGCTAACATGGGGGATGCAAATGCAAGCA
TCCCTACACCTGAACCTGTGATTTCAAGGCCAATGTTTGGAGCATTTGGTAAGGCTGGGAGCACTCACTCCATTGCTTTTGTCAGCAAGGAAGCTGCAGA
CAATGGGATCAAGGCAGAGTACGAACTCGACAAGAGAGTGGAAGCTGTGGGTGGTGTTAGGAAACTCACCAAACTCGACATGAAACTCAATGATGCCCTT
CCAGATATTACAGTGGATCCAGAAACATACACTGTAACTGCAGATGGCGAGGTTCTTACATGCCCCGCTGCCACCACAGTTCCCCTTTCACGTAATTATT
TTCTATTTTAG
|
||||||||||||||||||||
|
AA sequence
|
>Potri.008G177900.1 pacid=42806615 polypeptide=Potri.008G177900.1.p locus=Potri.008G177900 ID=Potri.008G177900.1.v4.1 annot-version=v4.1
MKLTPREVDKLGLHNAGFLAQKRLARGRKLNYTEAVALIASQILEFVRDGDKSVAELMDIGKQILGRRQVLPAVPHLLDTVQVEGTFPDGTKLITVHNAI
ASENGNLELALQGSFLPVPSLDKFPAIEDNEIPGAIIFGDGNVIINSGRKAVTLKVINTGDRPIQVGSHYHFIETNRSLLFDRRKAHGMRLNIPAGTAIR
FEPGESKSVVLVSIGGKQVIKGGNGIVDGPVDHENWTNIMGNIRRREFGNREEENASEGVIGEGSAFNNTISREAYANMYGPTAGDKIRLGDTNLYAEIE
RDFAFYGDECVFGGGKVIRDGMGQSCGHQPADSLDTVITNAVVIDYSGIYKADIGIKDYLIHAIGKSGNPDVMNVPSDMTIGVNTEVIAGEGMIVTAGGI
DCHVHFICPQLAFESISSGITTLVGGGTGPADGTRATTCTPAPSHMKLMLQSTDDLPLNFGFTGKGNAAKPEELHKIIRAGAMGLKLHEDWGTTPAAIDN
CLTVADEYDVQANIHTDTLNESGFVEDTIAAFKGRTIHTYHSEGAGGGHAPDIIKVCGVKNVLPSSTNPTRPYTSNTIDEHLDMLMVCHHLDKNIPEDVA
FAESRIRAETIAAEDILHDMGAISIISSDSQAMGRIGEVISRTWQTAHKMKSQRGLIGPGGSDNDNFRIRRYIAKYTINPAIANGLAKFVGSVEVGKLAD
LVLWKPSFFGAKPEMVIKGGAIAWANMGDANASIPTPEPVISRPMFGAFGKAGSTHSIAFVSKEAADNGIKAEYELDKRVEAVGGVRKLTKLDMKLNDAL
PDITVDPETYTVTADGEVLTCPAATTVPLSRNYFLF
|
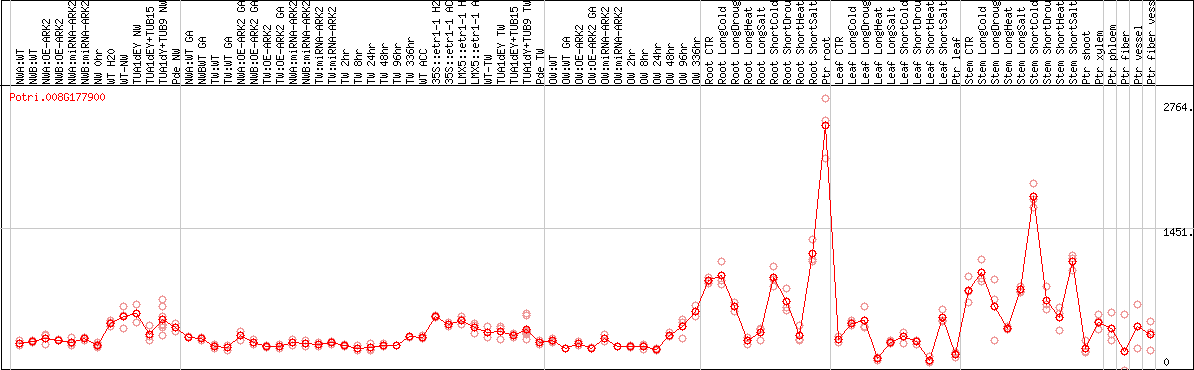
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.008G177900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
